

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
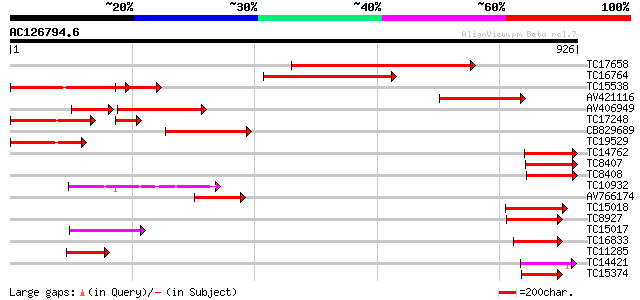
Query= AC126794.6 - phase: 0
(926 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
TC17658 homologue to UP|CLPA_PEA (P35100) ATP-dependent Clp prot... 547 e-156
TC16764 homologue to UP|CLPA_PEA (P35100) ATP-dependent Clp prot... 385 e-107
TC15538 similar to UP|CLPA_PEA (P35100) ATP-dependent Clp protea... 286 8e-78
AV421116 271 3e-73
AV406949 247 5e-66
TC17248 similar to UP|CLPA_PEA (P35100) ATP-dependent Clp protea... 187 5e-48
CB829689 187 9e-48
TC19529 similar to UP|CLPA_PEA (P35100) ATP-dependent Clp protea... 155 4e-38
TC14762 homologue to UP|CLPA_PEA (P35100) ATP-dependent Clp prot... 152 2e-37
TC8407 homologue to UP|CLPA_PEA (P35100) ATP-dependent Clp prote... 151 4e-37
TC8408 homologue to UP|CLPA_PEA (P35100) ATP-dependent Clp prote... 147 7e-36
TC10932 homologue to UP|Q39889 (Q39889) Heat shock protein , par... 119 3e-27
AV766174 118 4e-27
TC15018 weakly similar to UP|ERD1_ARATH (P42762) ERD1 protein, c... 103 2e-22
TC8927 similar to UP|Q9LLI0 (Q9LLI0) ClpB, partial (12%) 74 8e-14
TC15017 69 4e-12
TC16833 similar to UP|Q9LF37 (Q9LF37) ClpB heat shock protein-li... 59 5e-09
TC11285 55 4e-08
TC14421 similar to UP|Q39889 (Q39889) Heat shock protein , parti... 52 3e-07
TC15374 similar to UP|Q9LF37 (Q9LF37) ClpB heat shock protein-li... 50 1e-06
>TC17658 homologue to UP|CLPA_PEA (P35100) ATP-dependent Clp protease
ATP-binding subunit clpA homolog, chloroplast precursor
, partial (33%)
Length = 908
Score = 547 bits (1410), Expect = e-156
Identities = 277/302 (91%), Positives = 294/302 (96%)
Frame = +1
Query: 460 LRYTDEALVAAAELSHQYISDRFLPDKAIDLIDEAGSRVRLQHAQLPEEARGLEKEVRQI 519
LRYTDE+LVAAA+LS+QYISDRFLPDKAIDLIDEAGSRVRLQHAQLPEEAR L+KEVR I
Sbjct: 1 LRYTDESLVAAAQLSYQYISDRFLPDKAIDLIDEAGSRVRLQHAQLPEEARELDKEVRLI 180
Query: 520 VKEKDEAVRNQEFEKAGELRDKEMDLKTQISALIEKNKEMNKAESEAGDVGALVTEVDIQ 579
+KEK+E+VRNQ+FEKAGELRDKEMDLK QISAL+EK KEM+KAE+EAGD G +VTEVDIQ
Sbjct: 181 IKEKEESVRNQDFEKAGELRDKEMDLKAQISALVEKGKEMSKAETEAGDAGPVVTEVDIQ 360
Query: 580 HIVASWTGIPVDKVSVDESDRLLKMEDTLHKRIIGQHEAVEAISRAIRRARVGLKNPNRP 639
HIV++WTGIPV+KVS DESDRLLKMEDTLHKRIIGQ EAV+AISRAIRRARVGLKNPNRP
Sbjct: 361 HIVSAWTGIPVEKVSTDESDRLLKMEDTLHKRIIGQDEAVKAISRAIRRARVGLKNPNRP 540
Query: 640 IASFIFSGPTGVGKSELAKALASYYFGSEEAMIRLDMSEFMERHTVSKLIGSPPGYVGYT 699
IASFIFSGPTGVGKSELAKALA+YYFGSEEAMIRLDMSEFMERHTVSKLIGSPPGYVGYT
Sbjct: 541 IASFIFSGPTGVGKSELAKALAAYYFGSEEAMIRLDMSEFMERHTVSKLIGSPPGYVGYT 720
Query: 700 EGGQLTEAVRRRPYTVVLFDEIEKAHPDVFNMMLQILEDGRLTDSKGRTVDFKNTLLIMT 759
EGGQLTEAVRRRPYTVVLFDEIEKAHPDVFNMMLQILEDGRLTDSKGRTVDFKNTLLIMT
Sbjct: 721 EGGQLTEAVRRRPYTVVLFDEIEKAHPDVFNMMLQILEDGRLTDSKGRTVDFKNTLLIMT 900
Query: 760 SN 761
SN
Sbjct: 901 SN 906
>TC16764 homologue to UP|CLPA_PEA (P35100) ATP-dependent Clp protease
ATP-binding subunit clpA homolog, chloroplast precursor
, partial (24%)
Length = 653
Score = 385 bits (990), Expect = e-107
Identities = 197/217 (90%), Positives = 208/217 (95%)
Frame = +3
Query: 415 EYRKHIEKDPALERRFQPVKVPEPTVPETIQILKGLRERYEIHHKLRYTDEALVAAAELS 474
EYRKHIEKDPALERRFQPVKVPEPTV ETIQILKGLRERYEIHHKLRYTD+ALVAAA LS
Sbjct: 3 EYRKHIEKDPALERRFQPVKVPEPTVDETIQILKGLRERYEIHHKLRYTDDALVAAAHLS 182
Query: 475 HQYISDRFLPDKAIDLIDEAGSRVRLQHAQLPEEARGLEKEVRQIVKEKDEAVRNQEFEK 534
+QYISDRFLPDKAIDLIDEAGSRVRL+HAQLPEEAR L+KEVRQIVKEKDEAVRNQ+FEK
Sbjct: 183 YQYISDRFLPDKAIDLIDEAGSRVRLRHAQLPEEARELDKEVRQIVKEKDEAVRNQDFEK 362
Query: 535 AGELRDKEMDLKTQISALIEKNKEMNKAESEAGDVGALVTEVDIQHIVASWTGIPVDKVS 594
AGELRD+EMDLKTQISALIEK KEM+KAESEA D G +VTEVDIQHIVASWTGIPVDKVS
Sbjct: 363 AGELRDREMDLKTQISALIEKGKEMSKAESEAADGGPVVTEVDIQHIVASWTGIPVDKVS 542
Query: 595 VDESDRLLKMEDTLHKRIIGQHEAVEAISRAIRRARV 631
DESDRLLKME+TLHKR+IGQ EAV+AISRAIRRARV
Sbjct: 543 TDESDRLLKMEETLHKRVIGQDEAVKAISRAIRRARV 653
>TC15538 similar to UP|CLPA_PEA (P35100) ATP-dependent Clp protease
ATP-binding subunit clpA homolog, chloroplast precursor
, partial (21%)
Length = 658
Score = 286 bits (733), Expect = 8e-78
Identities = 154/196 (78%), Positives = 167/196 (84%)
Frame = +1
Query: 1 MSRALAQSINVPGLVAGRRHVNNNKGAARSRRSVRMMFTTRTASPRLSSYSGLRTLNSLD 60
M+R LAQSINVP LVA +RH N+KG+ +S+RS +MM +T+ RL+ +SGLRT N LD
Sbjct: 82 MARVLAQSINVPDLVARQRH-GNHKGSGKSKRSAKMMCALQTSGFRLAGFSGLRTYNPLD 258
Query: 61 SMLRPGQDFHSKVLTQIGTNRAKGGRGSRCVTKAMFERFTEKAIKVIMLAQEEARRLGHN 120
+MLRPG DFHSKV + R K RG V KAMFERFTEKAIKVIMLAQEEARRLGHN
Sbjct: 259 TMLRPGLDFHSKVSIATSSRRGKATRG---VPKAMFERFTEKAIKVIMLAQEEARRLGHN 429
Query: 121 FVGTEQILLGLIGEGTGIAAKVLKSMGINLKDARVEVEKIIGRGSGFVAVEIPFTPRAKR 180
FVGTEQILLGLIGEGTGIAAKVLKSMGINLKDARVEVEKIIGRGSGFVAVEIPFTPRAKR
Sbjct: 430 FVGTEQILLGLIGEGTGIAAKVLKSMGINLKDARVEVEKIIGRGSGFVAVEIPFTPRAKR 609
Query: 181 VLELSLEEARQLGHNY 196
VLELSLEEARQLGHNY
Sbjct: 610 VLELSLEEARQLGHNY 657
Score = 72.8 bits (177), Expect = 2e-13
Identities = 32/74 (43%), Positives = 55/74 (74%)
Frame = +1
Query: 174 FTPRAKRVLELSLEEARQLGHNYIGSEHLLLGLLREGEGVAARVLENLGADPTNIRTQVI 233
FT +A +V+ L+ EEAR+LGHN++G+E +LLGL+ EG G+AA+VL+++G + + R +V
Sbjct: 364 FTEKAIKVIMLAQEEARRLGHNFVGTEQILLGLIGEGTGIAAKVLKSMGINLKDARVEVE 543
Query: 234 RMVGEGADSVGATV 247
+++G G+ V +
Sbjct: 544 KIIGRGSGFVAVEI 585
>AV421116
Length = 421
Score = 271 bits (694), Expect = 3e-73
Identities = 138/140 (98%), Positives = 140/140 (99%)
Frame = +1
Query: 703 QLTEAVRRRPYTVVLFDEIEKAHPDVFNMMLQILEDGRLTDSKGRTVDFKNTLLIMTSNV 762
+LTEAVRRRPYTVVLFDEIEKAHPDVFNMMLQILEDGRLTDSKGRTVDFKNTLLIMTSNV
Sbjct: 1 RLTEAVRRRPYTVVLFDEIEKAHPDVFNMMLQILEDGRLTDSKGRTVDFKNTLLIMTSNV 180
Query: 763 GSSVIEKGGRRIGFDLDYDEKDSSYNRIKSLVTEELKQYFRPEFLNRLDEMIVFRQLTKL 822
GSSVIEKGGR+IGFDLDYDEKDSSYNRIKSLVTEELKQYFRPEFLNRLDEMIVFRQLTKL
Sbjct: 181 GSSVIEKGGRKIGFDLDYDEKDSSYNRIKSLVTEELKQYFRPEFLNRLDEMIVFRQLTKL 360
Query: 823 EVKEIADIMLKEVFERLKTK 842
EVKEIADIMLKEVFERLKTK
Sbjct: 361 EVKEIADIMLKEVFERLKTK 420
>AV406949
Length = 443
Score = 247 bits (631), Expect = 5e-66
Identities = 127/147 (86%), Positives = 133/147 (90%), Gaps = 1/147 (0%)
Frame = +1
Query: 176 PRAKRVLELSLEEARQLGHNYIGSEHLLLGLLREGEGVAARVLENLGADPTNIRTQVIRM 235
PRAKRV ELSLEEARQLGHNYIGSEHLLLGLLREGEGVAARVL++LGADP NIR QVIRM
Sbjct: 1 PRAKRVPELSLEEARQLGHNYIGSEHLLLGLLREGEGVAARVLQDLGADPNNIRAQVIRM 180
Query: 236 VGEGA-DSVGATVGSGSSNNKMPTLEEYGTNLTKLAEEGKLDPVVGRQPQIERVTQILGR 294
VGE ++ G VG G S+NKMPTLEEYGTNLTKLA EGKLDPVVGRQ QIERV QILGR
Sbjct: 181 VGESNNETAGVAVGRGGSSNKMPTLEEYGTNLTKLANEGKLDPVVGRQQQIERVIQILGR 360
Query: 295 RTKNNPCLIGEPGVGKTAIAEGLAQRI 321
RTKNNPCL+GEPGVGKTAIAEGLAQRI
Sbjct: 361 RTKNNPCLVGEPGVGKTAIAEGLAQRI 441
Score = 63.9 bits (154), Expect = 1e-10
Identities = 29/68 (42%), Positives = 49/68 (71%)
Frame = +1
Query: 102 KAIKVIMLAQEEARRLGHNFVGTEQILLGLIGEGTGIAAKVLKSMGINLKDARVEVEKII 161
+A +V L+ EEAR+LGHN++G+E +LLGL+ EG G+AA+VL+ +G + + R +V +++
Sbjct: 4 RAKRVPELSLEEARQLGHNYIGSEHLLLGLLREGEGVAARVLQDLGADPNNIRAQVIRMV 183
Query: 162 GRGSGFVA 169
G + A
Sbjct: 184 GESNNETA 207
>TC17248 similar to UP|CLPA_PEA (P35100) ATP-dependent Clp protease
ATP-binding subunit clpA homolog, chloroplast precursor
, partial (11%)
Length = 515
Score = 187 bits (476), Expect = 5e-48
Identities = 100/141 (70%), Positives = 115/141 (80%), Gaps = 1/141 (0%)
Frame = +3
Query: 1 MSRALAQSINVPGLVAGRRHVNNNKGAARSRRSVRMMFTTRTASP-RLSSYSGLRTLNSL 59
MSR LAQSIN+PGLVAGR H NN+ + +S+R V+MM+TT A RLS +SGLR+ N+L
Sbjct: 102 MSRVLAQSINIPGLVAGRTHGRNNR-STKSKRPVKMMYTTLQAPALRLSGFSGLRSFNNL 278
Query: 60 DSMLRPGQDFHSKVLTQIGTNRAKGGRGSRCVTKAMFERFTEKAIKVIMLAQEEARRLGH 119
D+MLRPG DFHSKV T ++ R SRCV KAMFERFTEKAIKVIMLAQEEARR+GH
Sbjct: 279 DTMLRPGLDFHSKVFAT--TTMSRKARASRCVPKAMFERFTEKAIKVIMLAQEEARRMGH 452
Query: 120 NFVGTEQILLGLIGEGTGIAA 140
NFVGTEQ+LLGLIGEGTGIAA
Sbjct: 453 NFVGTEQVLLGLIGEGTGIAA 515
Score = 53.9 bits (128), Expect = 1e-07
Identities = 23/42 (54%), Positives = 35/42 (82%)
Frame = +3
Query: 174 FTPRAKRVLELSLEEARQLGHNYIGSEHLLLGLLREGEGVAA 215
FT +A +V+ L+ EEAR++GHN++G+E +LLGL+ EG G+AA
Sbjct: 390 FTEKAIKVIMLAQEEARRMGHNFVGTEQVLLGLIGEGTGIAA 515
>CB829689
Length = 558
Score = 187 bits (474), Expect = 9e-48
Identities = 88/142 (61%), Positives = 115/142 (80%), Gaps = 1/142 (0%)
Frame = +1
Query: 255 KMPTLEEYGTNLTKLAEEGKLDPVVGRQPQIERVTQILGRRTKNNPCLIGEPGVGKTAIA 314
K LE+YG +LT +A+ GKLDPV+GR +I R QIL RRTKNNP LIGEPGVGKTAI+
Sbjct: 133 KYEALEKYGKDLTAMAKAGKLDPVIGRDDEIRRCIQILSRRTKNNPVLIGEPGVGKTAIS 312
Query: 315 EGLAQRIANGDVPETIEGKKVITLDMGLLVAGTKYRGEFEERLKKLMEEIKQSD-EIILF 373
EGLAQRI GDVP+ + +++I+LDMG L+AG KYRGEFE+RLK +++E+ +SD + +LF
Sbjct: 313 EGLAQRIVQGDVPQALMNRRLISLDMGALIAGAKYRGEFEDRLKAVLKEVTESDGQTVLF 492
Query: 374 IDEVHTLIGAGAAEGAIDAANI 395
IDE+HT++GAGA GA+DA N+
Sbjct: 493 IDEIHTVVGAGATNGAMDAGNL 558
>TC19529 similar to UP|CLPA_PEA (P35100) ATP-dependent Clp protease
ATP-binding subunit clpA homolog, chloroplast precursor
, partial (8%)
Length = 560
Score = 155 bits (391), Expect = 4e-38
Identities = 81/125 (64%), Positives = 97/125 (76%)
Frame = -1
Query: 1 MSRALAQSINVPGLVAGRRHVNNNKGAARSRRSVRMMFTTRTASPRLSSYSGLRTLNSLD 60
MSR LAQSI +PGLV GR H +NN+ + SRRS++MM T + + R+S +SGLRT N+LD
Sbjct: 365 MSRVLAQSITIPGLVCGRSHGHNNR-STMSRRSLKMMSTLQAPALRMSGFSGLRTYNNLD 189
Query: 61 SMLRPGQDFHSKVLTQIGTNRAKGGRGSRCVTKAMFERFTEKAIKVIMLAQEEARRLGHN 120
+MLRPG DF SKV G ++ R SRC+ KAMFERFTEKAIKVIML+QEEARRLGHN
Sbjct: 188 TMLRPGLDFRSKVF---GVTTSRKARASRCIPKAMFERFTEKAIKVIMLSQEEARRLGHN 18
Query: 121 FVGTE 125
FVGTE
Sbjct: 17 FVGTE 3
Score = 38.1 bits (87), Expect = 0.006
Identities = 16/27 (59%), Positives = 23/27 (84%)
Frame = -1
Query: 174 FTPRAKRVLELSLEEARQLGHNYIGSE 200
FT +A +V+ LS EEAR+LGHN++G+E
Sbjct: 83 FTEKAIKVIMLSQEEARRLGHNFVGTE 3
>TC14762 homologue to UP|CLPA_PEA (P35100) ATP-dependent Clp protease
ATP-binding subunit clpA homolog, chloroplast precursor
, partial (9%)
Length = 588
Score = 152 bits (385), Expect = 2e-37
Identities = 76/85 (89%), Positives = 82/85 (96%)
Frame = +1
Query: 842 KEIELSVTERFRERVVDEGYNPSYGARPLRRAIMRLLEDSMAEKMLAREIKEGDSVIVDA 901
+EIEL VTERFR+RVVDEGY+PSYGARPLRRAIMRLLEDSMAEKMLAREIKEGDSVIVD
Sbjct: 1 QEIELPVTERFRDRVVDEGYDPSYGARPLRRAIMRLLEDSMAEKMLAREIKEGDSVIVDV 180
Query: 902 DSDGNVIVLNGSTGAPDSLPDALPV 926
DSDGNVIVLNGS+G P+SLP+ALPV
Sbjct: 181 DSDGNVIVLNGSSGTPESLPEALPV 255
>TC8407 homologue to UP|CLPA_PEA (P35100) ATP-dependent Clp protease
ATP-binding subunit clpA homolog, chloroplast precursor
, partial (8%)
Length = 603
Score = 151 bits (382), Expect = 4e-37
Identities = 78/86 (90%), Positives = 83/86 (95%), Gaps = 2/86 (2%)
Frame = +2
Query: 843 EIELSVTERFRERVVDEGYNPSYGARPLRRAIMRLLEDSMAEKMLAREIKEGDSVIVDAD 902
EI+LSVTERF+ERVVDEGYNPSYGARPLRRAIMRLLEDSMAEKMLAREIKEGDSVIVDAD
Sbjct: 2 EIDLSVTERFKERVVDEGYNPSYGARPLRRAIMRLLEDSMAEKMLAREIKEGDSVIVDAD 181
Query: 903 SDGNVIVLNGS--TGAPDSLPDALPV 926
SDGNVIVLNGS +GAP+SLP+ LPV
Sbjct: 182 SDGNVIVLNGSSDSGAPESLPEVLPV 259
>TC8408 homologue to UP|CLPA_PEA (P35100) ATP-dependent Clp protease
ATP-binding subunit clpA homolog, chloroplast precursor
, partial (9%)
Length = 498
Score = 147 bits (371), Expect = 7e-36
Identities = 73/83 (87%), Positives = 81/83 (96%)
Frame = +3
Query: 844 IELSVTERFRERVVDEGYNPSYGARPLRRAIMRLLEDSMAEKMLAREIKEGDSVIVDADS 903
I+LSVTERFR+RVV+EGY+PSYGARPLRRAIMRLLEDSMAEKMLA EIKEGDSVI+DADS
Sbjct: 3 IDLSVTERFRDRVVEEGYDPSYGARPLRRAIMRLLEDSMAEKMLAGEIKEGDSVIIDADS 182
Query: 904 DGNVIVLNGSTGAPDSLPDALPV 926
DG VIVLNGS+GAP+SLP+ALPV
Sbjct: 183 DGKVIVLNGSSGAPESLPEALPV 251
>TC10932 homologue to UP|Q39889 (Q39889) Heat shock protein , partial (28%)
Length = 1001
Score = 119 bits (297), Expect = 3e-27
Identities = 84/253 (33%), Positives = 131/253 (51%), Gaps = 6/253 (2%)
Frame = +3
Query: 97 ERFTEKAIKVIMLAQEEARRLGHNFVGTEQILLGLIGEGTGIAAKVLKSMGINLKDARVE 156
++FT K + + A E A GH + + LI + GI + + + + AR
Sbjct: 210 DKFTHKTNEALAGAHELAMSSGHAQMTPLHLASTLISDPNGIFFQAISNSS-GEESARA- 383
Query: 157 VEKIIGRGSGFVAV------EIPFTPRAKRVLELSLEEARQLGHNYIGSEHLLLGLLREG 210
VE+++ + + EIP + + + + + G ++ + L+LG+L +
Sbjct: 384 VERVLNQALKKLPSQSPPPDEIPASTTLIKAIRRAQAAQKSRGDTHLAVDQLILGILEDS 563
Query: 211 EGVAARVLENLGADPTNIRTQVIRMVGEGADSVGATVGSGSSNNKMPTLEEYGTNLTKLA 270
+ +L+ G +++++ ++ G+ VG V S S + L+ YG +L + A
Sbjct: 564 Q--IGDLLKEAGVAAAKVKSELDKLRGK----VGKKVESASGDTTFQALKTYGRDLVEQA 725
Query: 271 EEGKLDPVVGRQPQIERVTQILGRRTKNNPCLIGEPGVGKTAIAEGLAQRIANGDVPETI 330
GKLDPV+GR +I RV +IL RRTKNNP LIGEPGVGKTA+ EGLAQRI GDVP
Sbjct: 726 --GKLDPVIGRDEEIRRVVRILSRRTKNNPVLIGEPGVGKTAVVEGLAQRIVRGDVPS-- 893
Query: 331 EGKKVITLDMGLL 343
I L +GLL
Sbjct: 894 -----IFLMLGLL 917
Score = 40.8 bits (94), Expect = 0.001
Identities = 18/23 (78%), Positives = 20/23 (86%)
Frame = +2
Query: 334 KVITLDMGLLVAGTKYRGEFEER 356
++I LDMG LVAG KYRGEFEER
Sbjct: 908 RLIALDMGALVAGAKYRGEFEER 976
>AV766174
Length = 416
Score = 118 bits (296), Expect = 4e-27
Identities = 54/84 (64%), Positives = 73/84 (86%), Gaps = 1/84 (1%)
Frame = -2
Query: 302 LIGEPGVGKTAIAEGLAQRIANGDVPETIEGKKVITLDMGLLVAGTKYRGEFEERLKKLM 361
LIGEPGVGKTAI+EGLAQRI GDVP+ + +++I+LDMG L+AG KYRGEFE+RLK ++
Sbjct: 412 LIGEPGVGKTAISEGLAQRIVQGDVPQALMNRRLISLDMGALIAGAKYRGEFEDRLKAVL 233
Query: 362 EEIKQSD-EIILFIDEVHTLIGAG 384
+E+ +SD + +LFIDE+HT++GAG
Sbjct: 232 KEVTESDGQTVLFIDEIHTVVGAG 161
>TC15018 weakly similar to UP|ERD1_ARATH (P42762) ERD1 protein, chloroplast
precursor, partial (10%)
Length = 598
Score = 103 bits (256), Expect = 2e-22
Identities = 47/102 (46%), Positives = 75/102 (73%)
Frame = +2
Query: 810 LDEMIVFRQLTKLEVKEIADIMLKEVFERLKTKEIELSVTERFRERVVDEGYNPSYGARP 869
+DE++VFR L K ++ EI D++L++V +R+ + I+L V+E + V EGYNP+YGARP
Sbjct: 2 IDEIVVFRSLEKSQLLEILDVLLQDVKKRIMSLGIDLEVSESVKNLVCKEGYNPTYGARP 181
Query: 870 LRRAIMRLLEDSMAEKMLAREIKEGDSVIVDADSDGNVIVLN 911
L+RAI L+ED ++E L + K+GD+V++D D +GN++V N
Sbjct: 182 LKRAITSLIEDPLSEAFLCGKCKQGDTVLIDLDVNGNLLVTN 307
>TC8927 similar to UP|Q9LLI0 (Q9LLI0) ClpB, partial (12%)
Length = 619
Score = 74.3 bits (181), Expect = 8e-14
Identities = 37/91 (40%), Positives = 60/91 (65%)
Frame = +2
Query: 812 EMIVFRQLTKLEVKEIADIMLKEVFERLKTKEIELSVTERFRERVVDEGYNPSYGARPLR 871
E IVF+ L E+ +I ++ ++ V RLK K+I+L T+ E + G++P++GARP++
Sbjct: 2 EYIVFQPLDSKEISKIVELQMERVKNRLKQKKIDLHYTQEALELLSVLGFDPNFGARPVK 181
Query: 872 RAIMRLLEDSMAEKMLAREIKEGDSVIVDAD 902
R I L+E+ +A +L + KE DS+IVDAD
Sbjct: 182 RVIQPLVENEIAMGVLRGDFKEDDSIIVDAD 274
>TC15017
Length = 1155
Score = 68.6 bits (166), Expect = 4e-12
Identities = 41/128 (32%), Positives = 71/128 (55%), Gaps = 3/128 (2%)
Frame = +2
Query: 98 RFTEKAIKVIMLAQEEARRLGHNFVGTEQILLGLIGEGTGIAAKVLKSMGINLKDARVEV 157
+++ ++IK + + EAR+L + GTE +L+G++ EGT AAK L++ GI L R E
Sbjct: 263 KWSARSIKSFAMGELEARKLKYPTTGTEALLMGILVEGTSKAAKFLRANGITLFKVRDET 442
Query: 158 EKIIGRGS--GFVAVEIPFTPRAKRVLELSLEE-ARQLGHNYIGSEHLLLGLLREGEGVA 214
+++G+ F P T A++ L+ +++E + G I HLLLG+ + E
Sbjct: 443 VELLGKSDMYFFSPEHPPLTEPARKALDWAIDEKLKSGGEGEINVAHLLLGIWSQKESAG 622
Query: 215 ARVLENLG 222
++L LG
Sbjct: 623 QQILAGLG 646
>TC16833 similar to UP|Q9LF37 (Q9LF37) ClpB heat shock protein-like, partial
(10%)
Length = 510
Score = 58.5 bits (140), Expect = 5e-09
Identities = 25/80 (31%), Positives = 54/80 (67%)
Frame = +3
Query: 823 EVKEIADIMLKEVFERLKTKEIELSVTERFRERVVDEGYNPSYGARPLRRAIMRLLEDSM 882
++ I + L+ V +R+ +++++ VTE + + + GY+P+YGARP++R I + +E+ +
Sbjct: 9 QISSIVRLQLERVQKRIADRKMKIQVTEAAIKLLGNLGYDPNYGARPVKRVIQQNVENEL 188
Query: 883 AEKMLAREIKEGDSVIVDAD 902
A+ +L E K+ D+++VD +
Sbjct: 189 AKGILRGEFKDEDTILVDTE 248
>TC11285
Length = 456
Score = 55.5 bits (132), Expect = 4e-08
Identities = 27/70 (38%), Positives = 45/70 (63%)
Frame = +1
Query: 94 AMFERFTEKAIKVIMLAQEEARRLGHNFVGTEQILLGLIGEGTGIAAKVLKSMGINLKDA 153
A +++ KAIK + + EAR+L + GTE +L+G++ EGT +AAK L++ G+ L
Sbjct: 238 AEIPKWSSKAIKSFAMGELEARKLKYPTTGTEALLMGVLIEGTNVAAKFLRANGVTLFKV 417
Query: 154 RVEVEKIIGR 163
R E K++G+
Sbjct: 418 RDETVKLLGK 447
Score = 39.7 bits (91), Expect = 0.002
Identities = 23/69 (33%), Positives = 41/69 (59%), Gaps = 1/69 (1%)
Frame = +1
Query: 171 EIP-FTPRAKRVLELSLEEARQLGHNYIGSEHLLLGLLREGEGVAARVLENLGADPTNIR 229
EIP ++ +A + + EAR+L + G+E LL+G+L EG VAA+ L G +R
Sbjct: 241 EIPKWSSKAIKSFAMGELEARKLKYPTTGTEALLMGVLIEGTNVAAKFLRANGVTLFKVR 420
Query: 230 TQVIRMVGE 238
+ ++++G+
Sbjct: 421 DETVKLLGK 447
>TC14421 similar to UP|Q39889 (Q39889) Heat shock protein , partial (14%)
Length = 731
Score = 52.4 bits (124), Expect = 3e-07
Identities = 29/100 (29%), Positives = 53/100 (53%), Gaps = 9/100 (9%)
Frame = +1
Query: 835 VFERLKTKEIELSVTERFRERVVDEGYNPSYGARPLRRAIMRLLEDSMAEKMLAREIKEG 894
V RL + I ++VT+ + ++ E Y+P YGARP+RR + R + ++ ++ EI E
Sbjct: 1 VASRLAERGIAMAVTDAALDYILAESYDPVYGARPIRRWLERKVVTELSRMLIRDEIDEN 180
Query: 895 DSVIVDADSDGNVIV---------LNGSTGAPDSLPDALP 925
+V +DA + G+ +V +N +TG + +P
Sbjct: 181 STVYIDAGTKGSELVYRVEKNGGLVNAATGQKSDILIQIP 300
>TC15374 similar to UP|Q9LF37 (Q9LF37) ClpB heat shock protein-like, partial
(9%)
Length = 574
Score = 50.4 bits (119), Expect = 1e-06
Identities = 20/66 (30%), Positives = 46/66 (69%)
Frame = +2
Query: 837 ERLKTKEIELSVTERFRERVVDEGYNPSYGARPLRRAIMRLLEDSMAEKMLAREIKEGDS 896
+R+ +++++ VT+ + + GY+P+YGARP++R I + +E+ +A+ +L E K+ D+
Sbjct: 5 KRIADRKMKIHVTDAAVQVLGSLGYDPNYGARPVKRVIQQNVENELAKGILRGEFKDEDT 184
Query: 897 VIVDAD 902
+++D +
Sbjct: 185 ILIDTE 202
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.316 0.136 0.370
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 10,643,435
Number of Sequences: 28460
Number of extensions: 113094
Number of successful extensions: 612
Number of sequences better than 10.0: 68
Number of HSP's better than 10.0 without gapping: 588
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 601
length of query: 926
length of database: 4,897,600
effective HSP length: 99
effective length of query: 827
effective length of database: 2,080,060
effective search space: 1720209620
effective search space used: 1720209620
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.6 bits)
S2: 60 (27.7 bits)
Medicago: description of AC126794.6


