

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
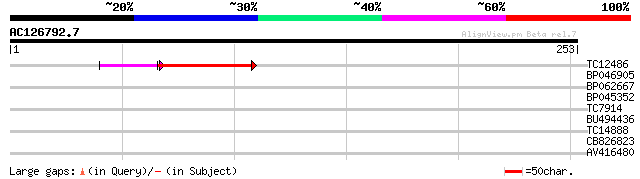
Query= AC126792.7 - phase: 0
(253 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
TC12486 47 2e-08
BP046905 28 1.2
BP062667 27 2.6
BP045352 27 3.4
TC7914 similar to UP|Q8VYW9 (Q8VYW9) Aminotransferase 1, complete 27 4.4
BU494436 26 7.5
TC14888 25 9.8
CB826823 25 9.8
AV416480 25 9.8
>TC12486
Length = 742
Score = 47.4 bits (111), Expect(2) = 2e-08
Identities = 22/44 (50%), Positives = 30/44 (68%)
Frame = +2
Query: 67 FADDVVLVGDSREEVNGRLETWRQALEAYGFCLSRSKTEYMECN 110
F + V V + RE++N RL RQ L+ YGF L++SK EYMEC+
Sbjct: 116 FCELYVEVRELREDINARLAICRQTLQTYGF*LNKSKAEYMECS 247
Score = 26.9 bits (58), Expect(2) = 2e-08
Identities = 14/29 (48%), Positives = 17/29 (58%)
Frame = +3
Query: 41 LSPYIFTLVLDVLTEHIQELAPRCMLFAD 69
LS Y F +VLD+L E I CMLF +
Sbjct: 39 LSHYHFIVVLDMLIEQI*ATMQLCMLFVN 125
>BP046905
Length = 538
Score = 28.5 bits (62), Expect = 1.2
Identities = 19/66 (28%), Positives = 32/66 (47%), Gaps = 5/66 (7%)
Frame = -3
Query: 82 NGRLETWRQALEAYGFC-----LSRSKTEYMECNFSGRRSKSTLEVKVGDHIIPQVTRFK 136
NG ET + E Y F + + EYM SG+ K +L+ I +++R+
Sbjct: 506 NGVAETLQIKEERYDFSEFLPEMIEERPEYMSRQLSGKEWKPSLDTIKEKKIKKKLSRWF 327
Query: 137 YLESFV 142
+L+SF+
Sbjct: 326 FLKSFI 309
>BP062667
Length = 528
Score = 27.3 bits (59), Expect = 2.6
Identities = 11/29 (37%), Positives = 17/29 (57%)
Frame = -2
Query: 2 IAYIRAIKDMYEGASTSVRTQDGTTEDFP 30
+ ++ I D +GAS++V T TT FP
Sbjct: 407 LGFVETIADALDGASSAVLTSASTTGSFP 321
>BP045352
Length = 434
Score = 26.9 bits (58), Expect = 3.4
Identities = 8/23 (34%), Positives = 16/23 (68%)
Frame = -1
Query: 199 WAVKSQHENQVSVAEMRMLRWMS 221
W++KSQ ++++ +LRW+S
Sbjct: 407 WSIKSQRSKHMTISLNELLRWLS 339
>TC7914 similar to UP|Q8VYW9 (Q8VYW9) Aminotransferase 1, complete
Length = 1532
Score = 26.6 bits (57), Expect = 4.4
Identities = 14/31 (45%), Positives = 18/31 (57%)
Frame = -1
Query: 223 KTRHDRIRNDIIRERERESGGSTYSRKVGRK 253
KT H+R NDI ++ E GS + KVG K
Sbjct: 1526 KTTHNRCSNDIYQDNE----GSNNASKVGNK 1446
>BU494436
Length = 449
Score = 25.8 bits (55), Expect = 7.5
Identities = 12/46 (26%), Positives = 20/46 (43%)
Frame = +3
Query: 154 HRIQVGWLKWRRASCVLCDKKVPFKLKGKFYRTAVRSALLYGTECW 199
H+ + GW W+R C + ++L G +LY +CW
Sbjct: 120 HQCRSGWEGWQRIVCTCHWQAGCYQL*GDDAGYCFLDRVLYQDDCW 257
>TC14888
Length = 573
Score = 25.4 bits (54), Expect = 9.8
Identities = 10/18 (55%), Positives = 12/18 (66%)
Frame = -1
Query: 97 FCLSRSKTEYMECNFSGR 114
FCLS T+ C+FSGR
Sbjct: 237 FCLSHLGTDSYSCHFSGR 184
>CB826823
Length = 429
Score = 25.4 bits (54), Expect = 9.8
Identities = 12/19 (63%), Positives = 16/19 (84%)
Frame = +1
Query: 235 RERERESGGSTYSRKVGRK 253
RERERE GG+T R++GR+
Sbjct: 316 RERERERGGTT--RELGRR 366
>AV416480
Length = 410
Score = 25.4 bits (54), Expect = 9.8
Identities = 14/41 (34%), Positives = 20/41 (48%)
Frame = +1
Query: 89 RQALEAYGFCLSRSKTEYMECNFSGRRSKSTLEVKVGDHII 129
R L AYG + S +EY++ N S S +V DH +
Sbjct: 208 RGGLGAYGGKILGSSSEYVQSNISRYFSDPQYYFQVNDHYV 330
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.321 0.135 0.405
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 4,286,699
Number of Sequences: 28460
Number of extensions: 58991
Number of successful extensions: 260
Number of sequences better than 10.0: 18
Number of HSP's better than 10.0 without gapping: 258
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 259
length of query: 253
length of database: 4,897,600
effective HSP length: 88
effective length of query: 165
effective length of database: 2,393,120
effective search space: 394864800
effective search space used: 394864800
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.9 bits)
S2: 54 (25.4 bits)
Medicago: description of AC126792.7


