

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
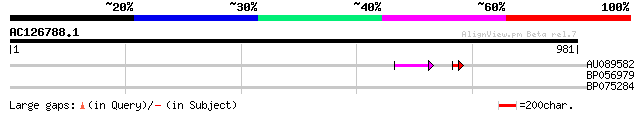
Query= AC126788.1 - phase: 0 /pseudo
(981 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
AU089582 38 3e-05
BP056979 28 5.4
BP075284 28 5.4
>AU089582
Length = 383
Score = 38.1 bits (87), Expect(2) = 3e-05
Identities = 28/68 (41%), Positives = 38/68 (55%)
Frame = +3
Query: 666 LSWRRFIFGTL*SCMVFRRVLYQIEIRGSLLDSGRVSKMLWARN*G*VRLIILRRMVSRR 725
L+ RF + L CMV+ + +QIE S G + K+LW +* *V L IL+ MVS R
Sbjct: 15 LNMLRFTWMRLFPCMVYLCL*FQIEELNSHHIFGGLFKLLWELD*K*VPLFILKLMVSPR 194
Query: 726 ERYNP*RI 733
+ *RI
Sbjct: 195 GLFRS*RI 218
Score = 27.3 bits (59), Expect(2) = 3e-05
Identities = 12/18 (66%), Positives = 12/18 (66%)
Frame = +1
Query: 767 WHRLKPCMVEGAELLCVG 784
WH LKPCMV A L VG
Sbjct: 319 WHHLKPCMVGDAGHL*VG 372
>BP056979
Length = 508
Score = 28.5 bits (62), Expect = 5.4
Identities = 17/37 (45%), Positives = 24/37 (63%)
Frame = -3
Query: 701 VSKMLWARN*G*VRLIILRRMVSRRERYNP*RIY*GC 737
VSK R *V ++IL+RM+S RE ++P + *GC
Sbjct: 110 VSKQCLLRGPR*VVIVILQRMISNRE-HDPQELI*GC 3
>BP075284
Length = 358
Score = 28.5 bits (62), Expect = 5.4
Identities = 13/23 (56%), Positives = 15/23 (64%)
Frame = +2
Query: 604 NIRSLQGC*HHWMYRSGNGIVFL 626
NIRS + C HHW RSG +V L
Sbjct: 131 NIRSTEACKHHW--RSGGMVVLL 193
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.373 0.166 0.653
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 18,802,031
Number of Sequences: 28460
Number of extensions: 286237
Number of successful extensions: 3961
Number of sequences better than 10.0: 6
Number of HSP's better than 10.0 without gapping: 2117
Number of HSP's successfully gapped in prelim test: 142
Number of HSP's that attempted gapping in prelim test: 1757
Number of HSP's gapped (non-prelim): 2421
length of query: 981
length of database: 4,897,600
effective HSP length: 99
effective length of query: 882
effective length of database: 2,080,060
effective search space: 1834612920
effective search space used: 1834612920
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 13 ( 7.0 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 36 (22.0 bits)
S2: 60 (27.7 bits)
Medicago: description of AC126788.1


