

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
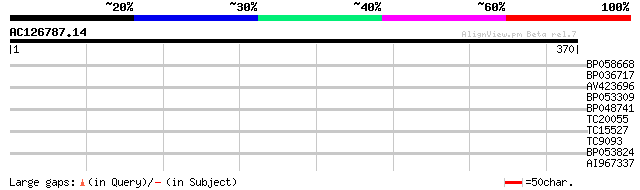
Query= AC126787.14 - phase: 0
(370 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
BP058668 29 1.4
BP036717 28 1.9
AV423696 28 1.9
BP053309 27 4.2
BP048741 27 5.4
TC20055 27 5.4
TC15527 similar to UP|AAR87166 (AAR87166) Expressed protein (Wit... 27 7.1
TC9093 UP|Q9SE27 (Q9SE27) Vacuolar H+-ATP synthase 16kDa proteol... 27 7.1
BP053824 27 7.1
AI967337 26 9.3
>BP058668
Length = 444
Score = 28.9 bits (63), Expect = 1.4
Identities = 13/24 (54%), Positives = 17/24 (70%)
Frame = +3
Query: 67 NDDVVFVEDRGLCVSLLHEIAWEV 90
+D+VV LCV+LLHEI+W V
Sbjct: 102 SDNVVATIRSLLCVNLLHEISWNV 173
>BP036717
Length = 477
Score = 28.5 bits (62), Expect = 1.9
Identities = 28/72 (38%), Positives = 32/72 (43%), Gaps = 10/72 (13%)
Frame = -1
Query: 22 MAGWHLSRNK--VLFFSGA--------LFITLAVGVHLTPYFPSVSDFMSSVKSSNDDVV 71
+ G HLS N VL FSG L +T V L P V D V + +D V
Sbjct: 327 LRGIHLS*NDTVVLQFSGGGCVLRRERLTVTAPWSVKLHHDKPVVLDDSGEVPLNQNDDV 148
Query: 72 FVEDRGLCVSLL 83
FV RGL V LL
Sbjct: 147 FVIHRGLLVRLL 112
>AV423696
Length = 427
Score = 28.5 bits (62), Expect = 1.9
Identities = 16/51 (31%), Positives = 23/51 (44%)
Frame = +1
Query: 110 FWSWSRLNSVSECEFQRLKRKDVMDLLNGSWVMIVGDSQARIFTLSLLSLV 160
+W W +EC R + + L+ V VGDS AR SLL ++
Sbjct: 70 YWRWKP----NECNIPRFEPHTFLQLIRNKHVAFVGDSLARNQLESLLCML 210
>BP053309
Length = 484
Score = 27.3 bits (59), Expect = 4.2
Identities = 16/49 (32%), Positives = 28/49 (56%), Gaps = 1/49 (2%)
Frame = -2
Query: 243 LLRNSVT-SMLPVSPKFGNDEPAMGSVSSVRSPHLFWLGMPTLINSMLN 290
L+R S + S++P+S + ND+P +G +S W L+N++LN
Sbjct: 423 LIRKSDSPSVIPISAQCFNDKPIVGIISPESGTTFVW-----LLNTILN 292
>BP048741
Length = 558
Score = 26.9 bits (58), Expect = 5.4
Identities = 12/28 (42%), Positives = 17/28 (59%), Gaps = 1/28 (3%)
Frame = +2
Query: 333 LNCGIKCTDDGMHYDGAVY-EAELHIMF 359
L+CG+ CT +H GA + E E I+F
Sbjct: 419 LHCGVFCTPQWLHRRGAAWSEQEFRILF 502
>TC20055
Length = 556
Score = 26.9 bits (58), Expect = 5.4
Identities = 14/43 (32%), Positives = 27/43 (62%), Gaps = 2/43 (4%)
Frame = +2
Query: 179 DYHID--IDEMGLKLDFIWAPYPNNLTDLVMRFKKKHLYPDVL 219
D+H+ ++ G +L ++A Y +L D V + +KHL+P++L
Sbjct: 296 DFHVAKFVEFKGQELG-LFAIYDGHLGDTVPAYLQKHLFPNIL 421
>TC15527 similar to UP|AAR87166 (AAR87166) Expressed protein (With
alternative splicing), partial (8%)
Length = 562
Score = 26.6 bits (57), Expect = 7.1
Identities = 13/45 (28%), Positives = 23/45 (50%)
Frame = -1
Query: 169 VRSFLFKRHSDYHIDIDEMGLKLDFIWAPYPNNLTDLVMRFKKKH 213
V +FK+H DY+I D F + Y + L + + +K++H
Sbjct: 490 VMEIIFKKHIDYNIARDHY-----FCFHHYTDELEVITLSYKQRH 371
>TC9093 UP|Q9SE27 (Q9SE27) Vacuolar H+-ATP synthase 16kDa proteolipid
subunit (Vacuolar ATP synthase proteolipid subunit) ,
complete
Length = 1062
Score = 26.6 bits (57), Expect = 7.1
Identities = 10/22 (45%), Positives = 14/22 (63%)
Frame = +1
Query: 206 VMRFKKKHLYPDVLVMGSGLWH 227
V+R ++ P L+ GSGLWH
Sbjct: 106 VLRLSRRRRCPCFLLYGSGLWH 171
>BP053824
Length = 395
Score = 26.6 bits (57), Expect = 7.1
Identities = 10/22 (45%), Positives = 13/22 (58%)
Frame = +3
Query: 221 MGSGLWHMLHITNASDYGGLLG 242
+G G +H +H T GGLLG
Sbjct: 93 LGVGWYHFVHFTRGHTLGGLLG 158
>AI967337
Length = 461
Score = 26.2 bits (56), Expect = 9.3
Identities = 17/66 (25%), Positives = 32/66 (47%)
Frame = +3
Query: 71 VFVEDRGLCVSLLHEIAWEVMPSKVFDPLLRNNSLNYDKFWSWSRLNSVSECEFQRLKRK 130
+F L V+ + +++ V+ + F + R LNY KFW+ + N+ + +L +
Sbjct: 84 MFPPPSSLVVTAMSVVSFVVLANSGFSEI-RGKHLNYSKFWNANSGNAAAAQRHIKLSGR 260
Query: 131 DVMDLL 136
M LL
Sbjct: 261 AGMLLL 278
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.324 0.139 0.427
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 6,830,686
Number of Sequences: 28460
Number of extensions: 99410
Number of successful extensions: 448
Number of sequences better than 10.0: 20
Number of HSP's better than 10.0 without gapping: 445
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 448
length of query: 370
length of database: 4,897,600
effective HSP length: 92
effective length of query: 278
effective length of database: 2,279,280
effective search space: 633639840
effective search space used: 633639840
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.0 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (22.0 bits)
S2: 56 (26.2 bits)
Medicago: description of AC126787.14


