

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
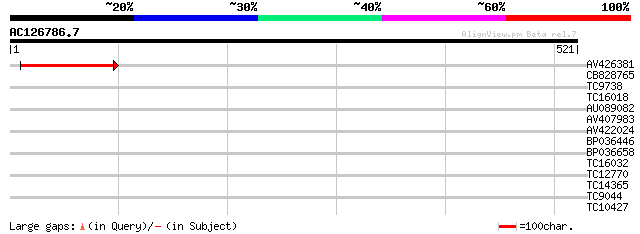
Query= AC126786.7 - phase: 0 /pseudo
(521 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
AV426381 88 3e-18
CB828765 39 0.002
TC9738 homologue to UP|Q9FVD4 (Q9FVD4) Ser/Thr specific protein ... 30 1.3
TC16018 UP|Q8MZB8 (Q8MZB8) AT15479p (CG31542-PA), partial (7%) 29 2.1
AU089082 29 2.1
AV407983 28 2.8
AV422024 28 2.8
BP036446 28 3.7
BP036658 28 3.7
TC16032 similar to BAC87886 (BAC87886) PGL-3, partial (4%) 28 4.8
TC12770 28 4.8
TC14365 similar to UP|Q9XFC1 (Q9XFC1) Stearoyl acyl carrier prot... 27 6.2
TC9044 similar to GB|AAR21619.1|38490051|AY457043 TATA-binding p... 27 8.1
TC10427 similar to UP|Q9AU06 (Q9AU06) NADPH-cytochrome P450 oxyd... 27 8.1
>AV426381
Length = 430
Score = 88.2 bits (217), Expect = 3e-18
Identities = 44/90 (48%), Positives = 62/90 (68%)
Frame = +2
Query: 11 RQLDVTPTWAVAAVCAIIVIISILLEKLIHKFASVFEERKKHALLEALEKIKAELMVLGF 70
++L+ TPTWAVA VC +++ ISI++E I E+R K AL EA+EKIK ELM++GF
Sbjct: 29 KKLEETPTWAVAVVCFVMLAISIIIEHGIEAIEKWLEKRHKKALHEAVEKIKGELMLMGF 208
Query: 71 ISLLLTFGQNYISKVCIPVKYSNTMLPCQP 100
IS LLT ++ IS +CI + ++T PC P
Sbjct: 209 ISFLLTVFKDPISNICISKQVASTWHPCHP 298
>CB828765
Length = 343
Score = 38.9 bits (89), Expect = 0.002
Identities = 15/27 (55%), Positives = 19/27 (69%)
Frame = +3
Query: 6 GGGDSRQLDVTPTWAVAAVCAIIVIIS 32
G S+ LD TPTWAVA VC + ++IS
Sbjct: 261 GSSSSKNLDQTPTWAVACVCTVFILIS 341
>TC9738 homologue to UP|Q9FVD4 (Q9FVD4) Ser/Thr specific protein
phosphatase 2A B regulatory subunit beta isoform,
partial (7%)
Length = 695
Score = 29.6 bits (65), Expect = 1.3
Identities = 16/38 (42%), Positives = 21/38 (55%)
Frame = +1
Query: 168 YIFSFSSWLSSM*YSVPLQ*HLVEQRLKVGKNGSKTIW 205
+ F FSS+ *YS PL HL+ + KV N S +W
Sbjct: 361 FFFQFSSF*FRP*YSFPLFLHLLLVKEKVQSNASYCVW 474
>TC16018 UP|Q8MZB8 (Q8MZB8) AT15479p (CG31542-PA), partial (7%)
Length = 425
Score = 28.9 bits (63), Expect = 2.1
Identities = 16/51 (31%), Positives = 24/51 (46%)
Frame = +1
Query: 93 NTMLPCQPLAERTADHPGEPALEPQGTEHEPTPNIPAGEGESKGEHHRRLL 143
N +P QP + + HP P +PQ TP +P+ S +H + LL
Sbjct: 31 NHGVPLQPRP*QLSRHPHPPPQQPQRRRRRTTPQLPS----STRQHRQPLL 171
>AU089082
Length = 300
Score = 28.9 bits (63), Expect = 2.1
Identities = 12/22 (54%), Positives = 13/22 (58%)
Frame = -1
Query: 149 FLGGGGGGPGCKPWTGYISYIF 170
FLGGGGGG G W G + F
Sbjct: 84 FLGGGGGGFGAGVWCGVVMLRF 19
>AV407983
Length = 417
Score = 28.5 bits (62), Expect = 2.8
Identities = 18/43 (41%), Positives = 19/43 (43%), Gaps = 6/43 (13%)
Frame = -1
Query: 129 AGEGESKGEHHR------RLLSYERRFLGGGGGGPGCKPWTGY 165
+GEG KG R R R LGGGGG G W GY
Sbjct: 261 SGEGRKKGWRRRGRV*RGRRSGNRRGVLGGGGGVLGRGVW*GY 133
>AV422024
Length = 460
Score = 28.5 bits (62), Expect = 2.8
Identities = 11/38 (28%), Positives = 21/38 (54%)
Frame = -3
Query: 91 YSNTMLPCQPLAERTADHPGEPALEPQGTEHEPTPNIP 128
++++ P PL + P EP ++ + +PTPN+P
Sbjct: 326 HNSSAFPPLPLCSASNLAPTEPRIDRTNSRSDPTPNLP 213
>BP036446
Length = 522
Score = 28.1 bits (61), Expect = 3.7
Identities = 17/50 (34%), Positives = 23/50 (46%), Gaps = 3/50 (6%)
Frame = +3
Query: 81 YISKVCIPVKYSNTMLPCQP--LAERTADHPGEP-ALEPQGTEHEPTPNI 127
Y SK C S++MLPC P + +A HP +L P P N+
Sbjct: 264 YCSKCCQTAPASSSMLPCTPPSAPDSSAPHPAPSCSLPPHTRARMPACNL 413
>BP036658
Length = 489
Score = 28.1 bits (61), Expect = 3.7
Identities = 10/18 (55%), Positives = 14/18 (77%)
Frame = -1
Query: 79 QNYISKVCIPVKYSNTML 96
QNY+ KVC+P+K S M+
Sbjct: 303 QNYMLKVCLPIKLSTNMV 250
>TC16032 similar to BAC87886 (BAC87886) PGL-3, partial (4%)
Length = 631
Score = 27.7 bits (60), Expect = 4.8
Identities = 20/55 (36%), Positives = 28/55 (50%), Gaps = 5/55 (9%)
Frame = +3
Query: 107 DHPGEPALEPQGTEHEPTPNIPAGE-----GESKGEHHRRLLSYERRFLGGGGGG 156
D E A+E + EHE T AGE ++K +H+ R Y+ + G GGGG
Sbjct: 72 DVEAENAVEVEYVEHEQTNPQVAGELNQPDVDAKDQHYPR-RGYQNQRGGRGGGG 233
>TC12770
Length = 588
Score = 27.7 bits (60), Expect = 4.8
Identities = 12/34 (35%), Positives = 18/34 (52%)
Frame = -3
Query: 104 RTADHPGEPALEPQGTEHEPTPNIPAGEGESKGE 137
R H EP+ + +G EH P +GE++GE
Sbjct: 580 RLQGHCHEPSPKTRGKEHATNMETPKQQGENRGE 479
>TC14365 similar to UP|Q9XFC1 (Q9XFC1) Stearoyl acyl carrier protein
desaturase Lldd3A20, partial (94%)
Length = 1539
Score = 27.3 bits (59), Expect = 6.2
Identities = 12/31 (38%), Positives = 17/31 (54%)
Frame = -3
Query: 2 GGGGGGGDSRQLDVTPTWAVAAVCAIIVIIS 32
GGGGG G W V VC+++V++S
Sbjct: 148 GGGGGLG---------WWGVVTVCSVVVVVS 83
>TC9044 similar to GB|AAR21619.1|38490051|AY457043 TATA-binding protein
associated factor 4 {Arabidopsis thaliana;} , partial
(5%)
Length = 580
Score = 26.9 bits (58), Expect = 8.1
Identities = 12/32 (37%), Positives = 15/32 (46%)
Frame = -3
Query: 96 LPCQPLAERTADHPGEPALEPQGTEHEPTPNI 127
LPC P A EP L P G+ P P++
Sbjct: 170 LPCFPAEPEAARKAVEPFLSPSGSSLVPLPDV 75
>TC10427 similar to UP|Q9AU06 (Q9AU06) NADPH-cytochrome P450 oxydoreductase
isoform 3, partial (17%)
Length = 565
Score = 26.9 bits (58), Expect = 8.1
Identities = 11/26 (42%), Positives = 14/26 (53%)
Frame = -3
Query: 131 EGESKGEHHRRLLSYERRFLGGGGGG 156
+G+ G+HH L E G GGGG
Sbjct: 287 DGDGGGKHHDELTILEEDGAGAGGGG 210
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.354 0.158 0.568
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 10,049,356
Number of Sequences: 28460
Number of extensions: 160476
Number of successful extensions: 3648
Number of sequences better than 10.0: 28
Number of HSP's better than 10.0 without gapping: 2072
Number of HSP's successfully gapped in prelim test: 60
Number of HSP's that attempted gapping in prelim test: 580
Number of HSP's gapped (non-prelim): 3007
length of query: 521
length of database: 4,897,600
effective HSP length: 94
effective length of query: 427
effective length of database: 2,222,360
effective search space: 948947720
effective search space used: 948947720
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 14 ( 7.1 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 37 (21.6 bits)
S2: 57 (26.6 bits)
Medicago: description of AC126786.7


