

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC126786.5 - phase: 0
(487 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
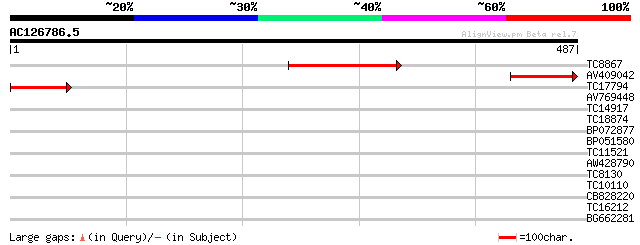
Score E
Sequences producing significant alignments: (bits) Value
TC8867 weakly similar to GB|AAM98094.1|22655006|AY139776 At1g764... 129 1e-30
AV409042 112 1e-25
TC17794 weakly similar to PIR|T04482|T04482 ribophorin I homolog... 51 4e-07
AV769448 36 0.012
TC14917 similar to GB|AAP37839.1|30725634|BT008480 At5g64750 {Ar... 31 0.52
TC18874 similar to UP|Q9FJJ0 (Q9FJJ0) Genomic DNA, chromosome 5,... 29 2.0
BP072877 28 2.6
BP051580 28 3.4
TC11521 similar to UP|O23144 (O23144) Proton pump interactor (AT... 28 4.4
AW428790 27 5.7
TC8130 similar to UP|Q9XFM6 (Q9XFM6) Progesterone-binding protei... 27 5.7
TC10110 weakly similar to UP|Q43854 (Q43854) Peroxidase precurso... 27 7.5
CB828220 27 9.8
TC16212 27 9.8
BG662281 27 9.8
>TC8867 weakly similar to GB|AAM98094.1|22655006|AY139776
At1g76400/F15M4_10 {Arabidopsis thaliana;}, partial
(14%)
Length = 649
Score = 129 bits (324), Expect = 1e-30
Identities = 60/97 (61%), Positives = 76/97 (77%)
Frame = +3
Query: 240 LVVKVVLPEGSKDISPSVPFPVKERHETKFSHLDIAGRPVVVLEKNNAVPEHNEHFQVYY 299
L+VKVVLPEGSKD + +PF V++ ETK+S+LD+ GR VV LEK N VPEHN FQVYY
Sbjct: 6 LIVKVVLPEGSKDPAAVIPFEVEQHLETKYSYLDVVGRTVVALEKRNVVPEHNTPFQVYY 185
Query: 300 KFNRLSMLREPLMLISGFFFLFLASIVYMHADLSISK 336
FN + ML EPLML+S FFFLF++++ Y+H D+SI K
Sbjct: 186 NFNPIFMLAEPLMLVSVFFFLFMSAVAYLHMDISIRK 296
>AV409042
Length = 337
Score = 112 bits (281), Expect = 1e-25
Identities = 55/57 (96%), Positives = 57/57 (99%)
Frame = -1
Query: 431 KERDLQEKLMAKHSTVVDCYEKKLGGREIENRIASHQQKITALKQEIDDLMDLIDEI 487
KERDLQEKLMAKHSTVVDCYEKKLGGREIENRIASHQQKITAL+QEIDDLMD+IDEI
Sbjct: 334 KERDLQEKLMAKHSTVVDCYEKKLGGREIENRIASHQQKITALRQEIDDLMDVIDEI 164
>TC17794 weakly similar to PIR|T04482|T04482 ribophorin I homolog - barley
(fragment) {Hordeum vulgare;}, partial (7%)
Length = 573
Score = 51.2 bits (121), Expect = 4e-07
Identities = 24/53 (45%), Positives = 36/53 (67%)
Frame = +1
Query: 1 MDILTVFTHILQPFPEKITQADIQLLLFQESAQYLSPYPVKVQSLNVKLPEAR 53
+++L TH L+PFP +I+Q++ QL F++SA LSPY VK Q+ +K P R
Sbjct: 415 LEVLYRLTHSLEPFPVEISQSESQLAYFRDSAILLSPYHVKQQTTFIKTPSTR 573
>AV769448
Length = 447
Score = 36.2 bits (82), Expect = 0.012
Identities = 14/19 (73%), Positives = 17/19 (88%)
Frame = +3
Query: 1 MDILTVFTHILQPFPEKIT 19
+D+L VFTH L+PFPEKIT
Sbjct: 390 LDVLAVFTHTLEPFPEKIT 446
>TC14917 similar to GB|AAP37839.1|30725634|BT008480 At5g64750 {Arabidopsis
thaliana;}, partial (17%)
Length = 1051
Score = 30.8 bits (68), Expect = 0.52
Identities = 17/40 (42%), Positives = 22/40 (54%), Gaps = 1/40 (2%)
Frame = +1
Query: 48 KLPEARIESYTKLENTKLQGSELKY-GPYENLPPFSYLPI 86
K PE I+ + LE T+ + LK G Y NLP + LPI
Sbjct: 313 KQPEPMIKLHCDLEATRQNSTSLKM*GSYNNLPSINQLPI 432
>TC18874 similar to UP|Q9FJJ0 (Q9FJJ0) Genomic DNA, chromosome 5, TAC
clone:K19B1, partial (33%)
Length = 543
Score = 28.9 bits (63), Expect = 2.0
Identities = 22/84 (26%), Positives = 40/84 (47%), Gaps = 9/84 (10%)
Frame = -1
Query: 333 SISKTSASYLAKIQWEEVQATIHQVHSIVS--RCLTTHDKLEAS-------LRDLSRTGD 383
++ +TS++ LA+ + +++++ S S R L T K ++ L D R+
Sbjct: 486 TLHRTSSTRLARFPFATIESSVGTTPSFTSLTRLLCTRVKWYSASAESCLVLSDSDRSAC 307
Query: 384 IQGCKATRKSVDSSLKELTKELKS 407
I GC A R + L + +LKS
Sbjct: 306 ISGCNAPRTPIAILLSSSSAKLKS 235
>BP072877
Length = 433
Score = 28.5 bits (62), Expect = 2.6
Identities = 14/36 (38%), Positives = 21/36 (57%)
Frame = -1
Query: 387 CKATRKSVDSSLKELTKELKSPVTFLQSSPQAAQIL 422
C +RKS D+S+ + K ++SP L SSP + L
Sbjct: 424 CVESRKSEDNSIHAVGKRVRSPRPMLFSSPSCSSEL 317
>BP051580
Length = 569
Score = 28.1 bits (61), Expect = 3.4
Identities = 11/39 (28%), Positives = 24/39 (61%)
Frame = +3
Query: 3 ILTVFTHILQPFPEKITQADIQLLLFQESAQYLSPYPVK 41
I++ F H+ + F + +I++ L+ + ++SPYPV+
Sbjct: 447 IVSWFEHLCEYFSMSMRAPNIEIHLYMLNIVFISPYPVR 563
>TC11521 similar to UP|O23144 (O23144) Proton pump interactor
(AT4g27500/F27G19_100), partial (21%)
Length = 1005
Score = 27.7 bits (60), Expect = 4.4
Identities = 10/26 (38%), Positives = 19/26 (72%)
Frame = +2
Query: 382 GDIQGCKATRKSVDSSLKELTKELKS 407
GD+ G K R+++ S +K++ +ELK+
Sbjct: 917 GDLDGVKKERQAIRSKIKQIDEELKA 994
>AW428790
Length = 327
Score = 27.3 bits (59), Expect = 5.7
Identities = 16/55 (29%), Positives = 27/55 (49%)
Frame = +1
Query: 283 EKNNAVPEHNEHFQVYYKFNRLSMLREPLMLISGFFFLFLASIVYMHADLSISKT 337
E+N P+ + + F R SML +L++ F+F I Y+ + +SKT
Sbjct: 19 EENIVSPDRFSYIWAFSYFKRFSMLVFCQILLTQFYF*LCDIISYLWYPIVLSKT 183
>TC8130 similar to UP|Q9XFM6 (Q9XFM6) Progesterone-binding protein-like,
partial (70%)
Length = 977
Score = 27.3 bits (59), Expect = 5.7
Identities = 15/46 (32%), Positives = 24/46 (51%)
Frame = -2
Query: 327 YMHADLSISKTSASYLAKIQWEEVQATIHQVHSIVSRCLTTHDKLE 372
Y DLS+ KTS L ++++ +H+VH + C TH + E
Sbjct: 484 YPQ*DLSLQKTSLLKLC*HPCQQIKHKVHRVHK--TSCSETHHRSE 353
>TC10110 weakly similar to UP|Q43854 (Q43854) Peroxidase precursor ,
partial (34%)
Length = 497
Score = 26.9 bits (58), Expect = 7.5
Identities = 14/38 (36%), Positives = 20/38 (51%)
Frame = -2
Query: 365 LTTHDKLEASLRDLSRTGDIQGCKATRKSVDSSLKELT 402
+T H +LEASL + RT GC +K + S + T
Sbjct: 151 MTVHSQLEASLLQVPRTQRRHGCIGKQKVMKKSRAKST 38
>CB828220
Length = 533
Score = 26.6 bits (57), Expect = 9.8
Identities = 14/31 (45%), Positives = 22/31 (70%)
Frame = +2
Query: 394 VDSSLKELTKELKSPVTFLQSSPQAAQILPK 424
++ S+ ++ KELKSP T + SS QA+ +L K
Sbjct: 35 INQSIIQVLKELKSPETLMASS-QASLLLQK 124
>TC16212
Length = 549
Score = 26.6 bits (57), Expect = 9.8
Identities = 10/23 (43%), Positives = 15/23 (64%)
Frame = +2
Query: 315 SGFFFLFLASIVYMHADLSISKT 337
SGF FL +VY H+ +S+ K+
Sbjct: 329 SGFVFLLCIEMVYFHSSISLCKS 397
>BG662281
Length = 399
Score = 26.6 bits (57), Expect = 9.8
Identities = 10/28 (35%), Positives = 15/28 (52%)
Frame = -2
Query: 173 ISTSSLWGDSKKTELEIEPRYPLFGGWK 200
+S LWGDS + +E P + G W+
Sbjct: 161 VSVRFLWGDSMRVAVECGPEFFASGMWR 78
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.318 0.136 0.390
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 7,547,443
Number of Sequences: 28460
Number of extensions: 99527
Number of successful extensions: 520
Number of sequences better than 10.0: 30
Number of HSP's better than 10.0 without gapping: 520
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 520
length of query: 487
length of database: 4,897,600
effective HSP length: 94
effective length of query: 393
effective length of database: 2,222,360
effective search space: 873387480
effective search space used: 873387480
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 57 (26.6 bits)
Medicago: description of AC126786.5


