

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC126784.6 - phase: 0
(135 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
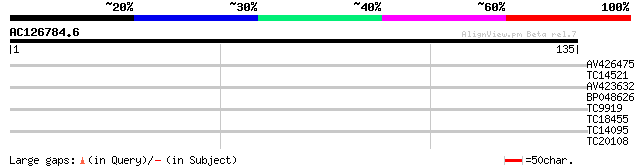
Score E
Sequences producing significant alignments: (bits) Value
AV426475 31 0.083
TC14521 similar to UP|O04894 (O04894) Transaldolase , partial (... 27 1.2
AV423632 27 1.2
BP048626 26 2.0
TC9919 25 4.6
TC18455 25 6.0
TC14095 homologue to UP|RS2_ARATH (P49688) 40S ribosomal protein... 24 7.8
TC20108 24 7.8
>AV426475
Length = 300
Score = 30.8 bits (68), Expect = 0.083
Identities = 11/30 (36%), Positives = 17/30 (56%)
Frame = +2
Query: 47 TGVVGEDGLPLQQSHFRGRLLEGTTLPLPH 76
TG +G GLP+ H ++ GT++ PH
Sbjct: 182 TGAIGPTGLPVDTRHVSFNIISGTSMSCPH 271
>TC14521 similar to UP|O04894 (O04894) Transaldolase , partial (86%)
Length = 1695
Score = 26.9 bits (58), Expect = 1.2
Identities = 12/33 (36%), Positives = 19/33 (57%)
Frame = -1
Query: 87 NSVESSNSWETSVTFNDITYWNHDSVPSNNDDF 119
+S SS+SWE + + T+W +PS +D F
Sbjct: 1260 DSTPSSSSWEPTKLQSTPTFWRALYIPSASDAF 1162
>AV423632
Length = 484
Score = 26.9 bits (58), Expect = 1.2
Identities = 13/41 (31%), Positives = 22/41 (52%)
Frame = +2
Query: 88 SVESSNSWETSVTFNDITYWNHDSVPSNNDDFSRAFHWIPV 128
S+ SS+S + ND+T H+++ +DD R W+ V
Sbjct: 245 SIPSSSSRDKFPKPNDLTSIAHNTLDRTDDDDGRGEQWVEV 367
>BP048626
Length = 357
Score = 26.2 bits (56), Expect = 2.0
Identities = 10/22 (45%), Positives = 14/22 (63%)
Frame = +3
Query: 25 LLPCCIKHDGPTEVSQYFKPKP 46
L+P C+KHD P++ K KP
Sbjct: 72 LVPRCLKHDHPSDTKTASKLKP 137
>TC9919
Length = 702
Score = 25.0 bits (53), Expect = 4.6
Identities = 9/24 (37%), Positives = 14/24 (57%)
Frame = +3
Query: 22 RVHLLPCCIKHDGPTEVSQYFKPK 45
RVH P C+K ++S+ KP+
Sbjct: 42 RVHQKPICVKRSATDQISRSSKPE 113
>TC18455
Length = 565
Score = 24.6 bits (52), Expect = 6.0
Identities = 13/34 (38%), Positives = 16/34 (46%)
Frame = -3
Query: 68 EGTTLPLPHNYSGFVLGKKNSVESSNSWETSVTF 101
E T L V+GKK S SW +S+TF
Sbjct: 284 ETATYELHRTEQNQVVGKKRPP*SMKSWSSSLTF 183
>TC14095 homologue to UP|RS2_ARATH (P49688) 40S ribosomal protein S2,
partial (88%)
Length = 1134
Score = 24.3 bits (51), Expect = 7.8
Identities = 23/72 (31%), Positives = 33/72 (44%), Gaps = 4/72 (5%)
Frame = -2
Query: 25 LLPCCIKHDGP-TEVSQYFKPKPTGVVGEDGLPLQQSHFRGRLLE---GTTLPLPHNYSG 80
L+P C+KHD P T Q ++ + GL S G LLE G T LP
Sbjct: 938 LVPRCLKHDHPMTPKLQANSSLHPFLLKDKGLCCWLSQKIGILLEWRFGETSLLPEFRCQ 759
Query: 81 FVLGKKNSVESS 92
+G + +++SS
Sbjct: 758 KTIGLQQTIKSS 723
>TC20108
Length = 368
Score = 24.3 bits (51), Expect = 7.8
Identities = 13/24 (54%), Positives = 16/24 (66%), Gaps = 1/24 (4%)
Frame = +1
Query: 112 VPSNNDDFSRAFHWIPV-AEAVSC 134
VPSN + S AFHW + AEA +C
Sbjct: 109 VPSN*EVSSTAFHWFLL*AEA*NC 180
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.316 0.136 0.429
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 2,830,724
Number of Sequences: 28460
Number of extensions: 40630
Number of successful extensions: 163
Number of sequences better than 10.0: 16
Number of HSP's better than 10.0 without gapping: 163
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 163
length of query: 135
length of database: 4,897,600
effective HSP length: 81
effective length of query: 54
effective length of database: 2,592,340
effective search space: 139986360
effective search space used: 139986360
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.5 bits)
S2: 50 (23.9 bits)
Medicago: description of AC126784.6


