

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC126782.2 - phase: 0 /pseudo
(912 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
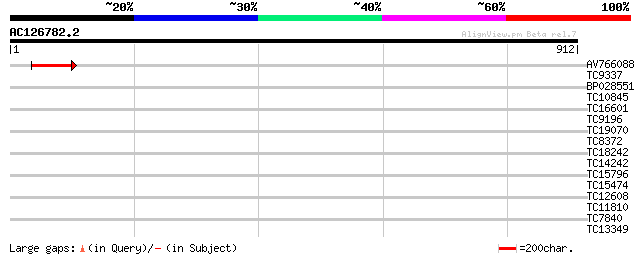
AV766088 61 9e-10
TC9337 similar to UP|PME_PRUPE (Q43062) Pectinesterase PPE8B pre... 34 0.12
BP028551 31 1.0
TC10845 weakly similar to UP|WRK6_ARATH (Q9C519) WRKY transcript... 30 1.3
TC16601 similar to PIR|S42728|S42728 phosphodiesterase (clone p4... 30 2.2
TC9196 weakly similar to UP|Q9LUC0 (Q9LUC0) Gb|AAD10662.1, parti... 28 5.0
TC19070 weakly similar to UP|Q9LRP6 (Q9LRP6) DNA-binding protein... 28 6.5
TC8372 similar to UP|O04334 (O04334) At2g30610 protein, partial ... 28 6.5
TC18242 weakly similar to UP|O70495 (O70495) Plenty-of-prolines-... 28 6.5
TC14242 homologue to UP|PSAF_SPIOL (P12355) Photosystem I reacti... 28 6.5
TC15796 similar to UP|Q948T1 (Q948T1) Two-pore calcium channel, ... 28 6.5
TC15474 UP|Q7Y1B9 (Q7Y1B9) Ammonium transporter, complete 28 8.5
TC12608 28 8.5
TC11810 28 8.5
TC7840 similar to UP|AAQ72788 (AAQ72788) Hypersensitive-induced ... 28 8.5
TC13349 28 8.5
>AV766088
Length = 501
Score = 60.8 bits (146), Expect = 9e-10
Identities = 35/73 (47%), Positives = 46/73 (62%)
Frame = -1
Query: 35 ILIFVIHVLLVNMLNCLS*ILIIEVCCHLIFYIVMYGPLLF*VQLVTNIMFFSWIIIQIF 94
I+ FV V L NML C +LI+ + C L FY V+YG LLF LV + MF+SW+I +I
Sbjct: 219 IVRFVSLVCLANMLGCHLFLLIMLL*CLLTFYTVIYGLLLFYRPLVIDFMFYSWMISRIS 40
Query: 95 CGLFQCHINPKFI 107
CG F IN +F+
Sbjct: 39 CGRFL*VINLRFL 1
>TC9337 similar to UP|PME_PRUPE (Q43062) Pectinesterase PPE8B precursor
(Pectin methylesterase) (PE) , partial (54%)
Length = 1145
Score = 33.9 bits (76), Expect = 0.12
Identities = 20/42 (47%), Positives = 26/42 (61%)
Frame = +1
Query: 35 ILIFVIHVLLVNMLNCLS*ILIIEVCCHLIFYIVMYGPLLF* 76
+++FV V NM CL L + + HLIF IV+YG LLF*
Sbjct: 1018 LILFVTLVF*ANMFGCLLVHLKLLL*DHLIFCIVIYGRLLF* 1143
>BP028551
Length = 235
Score = 30.8 bits (68), Expect = 1.0
Identities = 17/46 (36%), Positives = 24/46 (51%)
Frame = -1
Query: 565 DLPTTCHLLVFLKANVITLFSCTRKTLIWHIFYSRLMILFSPRPQM 610
D P T +L F + NVI+ F C H+ YS ++ L RPQ+
Sbjct: 151 DYPFTLNLNFFHEKNVISFFIC-------HLLYSYIVSLLKIRPQL 35
>TC10845 weakly similar to UP|WRK6_ARATH (Q9C519) WRKY transcription factor
6 (WRKY DNA-binding protein 6) (AtWRKY6), partial (17%)
Length = 635
Score = 30.4 bits (67), Expect = 1.3
Identities = 28/95 (29%), Positives = 43/95 (44%), Gaps = 14/95 (14%)
Frame = +1
Query: 303 TLS*TMSCPLILYNI*WTK---PKLAHLILNWLISQTKSLLLVQTHQISPHLHNL----- 354
TLS + L++Y I WTK L HL LN +S +LL + + SP +L
Sbjct: 28 TLSLSSCTYLLVYLILWTKDGVSLLIHLPLNPSLSFHPNLLHLSSTPCSPFTDSLSILPA 207
Query: 355 ------TPLVPSNKIFLPYFNQLYKLTQNLSLGVN 383
TPL+ + K+ + L + T L L ++
Sbjct: 208 TPTMNNTPLMTAGKLSAKSISSLIETTHRLLLAIS 312
>TC16601 similar to PIR|S42728|S42728 phosphodiesterase (clone p4-6) - mouse
{Mus musculus;}, partial (23%)
Length = 716
Score = 29.6 bits (65), Expect = 2.2
Identities = 25/80 (31%), Positives = 35/80 (43%)
Frame = +1
Query: 335 QTKSLLLVQTHQISPHLHNLTPLVPSNKIFLPYFNQLYKLTQNLSLGVNMGFLNQTKNIM 394
+T S LL +HQ+ HLH+L NQL++ T L F + K
Sbjct: 298 KTFS*LLQPSHQLQVHLHHL--------------NQLHQSTTRL-------FCSLAKWGK 414
Query: 395 VCTPMLQNPLYLGTLCLHLK 414
C+P + +G LCLH K
Sbjct: 415 TCSPWI-----IGILCLHSK 459
>TC9196 weakly similar to UP|Q9LUC0 (Q9LUC0) Gb|AAD10662.1, partial (25%)
Length = 799
Score = 28.5 bits (62), Expect = 5.0
Identities = 16/36 (44%), Positives = 20/36 (55%)
Frame = -1
Query: 351 LHNLTPLVPSNKIFLPYFNQLYKLTQNLSLGVNMGF 386
LH+LT + K FL FN Q LS G+N+GF
Sbjct: 175 LHHLTMFIVFRKSFLLIFNGSSCTLQLLSHGLNIGF 68
>TC19070 weakly similar to UP|Q9LRP6 (Q9LRP6) DNA-binding protein
(AT3g15590/MQD17_5), partial (20%)
Length = 744
Score = 28.1 bits (61), Expect = 6.5
Identities = 18/55 (32%), Positives = 29/55 (52%), Gaps = 6/55 (10%)
Frame = +1
Query: 210 LILSHLLK-----CCITKILLTITYECLVVYVFLSFL-HQKYTNFNLVPQNVCFL 258
L L HL K C I ++ + + ++C + F+S + H K+ F L + VCFL
Sbjct: 544 LKLMHLRKLQFPICLIEEVSMFLLFKCNYLSQFISNIEHSKFVCFVLYSKFVCFL 708
>TC8372 similar to UP|O04334 (O04334) At2g30610 protein, partial (98%)
Length = 1366
Score = 28.1 bits (61), Expect = 6.5
Identities = 15/48 (31%), Positives = 22/48 (45%)
Frame = +3
Query: 556 NKLLELGVSDLPTTCHLLVFLKANVITLFSCTRKTLIWHIFYSRLMIL 603
N L G S T CH+ F N+ L SC TL +S ++++
Sbjct: 420 NSFLAGGFS*FMTCCHMCDFHYCNIPCLISCRIATLSATFLFSEVLLM 563
>TC18242 weakly similar to UP|O70495 (O70495) Plenty-of-prolines-101,
partial (5%)
Length = 490
Score = 28.1 bits (61), Expect = 6.5
Identities = 14/46 (30%), Positives = 25/46 (53%)
Frame = +2
Query: 144 CVKLMELFFVCLVLTHLLKMVNLNEKYGPLTILFELY*FTRHYHLH 189
C KL F+C+ L HL+ + NE ++ +E++ ++ Y LH
Sbjct: 311 CWKLQ*AIFLCMSLNHLVMYIYTNELSNFISQSYEVH-ISQSYRLH 445
>TC14242 homologue to UP|PSAF_SPIOL (P12355) Photosystem I reaction centre
subunit III, chloroplast precursor (Light-harvesting
complex I 17 kDa protein) (PSI-F), partial (28%)
Length = 558
Score = 28.1 bits (61), Expect = 6.5
Identities = 19/60 (31%), Positives = 27/60 (44%), Gaps = 9/60 (15%)
Frame = -2
Query: 324 LAHLILNWLISQTKSLLLVQT--HQISPHLH-------NLTPLVPSNKIFLPYFNQLYKL 374
+ H L W + + LL+ HQ SPH+H LT + N+ Y NQL K+
Sbjct: 431 IQHQQLPWFYKKAEHPLLLSLFHHQFSPHIHLPRV**SPLTTINKPNQTKPNYLNQLKKI 252
>TC15796 similar to UP|Q948T1 (Q948T1) Two-pore calcium channel, partial
(42%)
Length = 1524
Score = 28.1 bits (61), Expect = 6.5
Identities = 18/58 (31%), Positives = 30/58 (51%)
Frame = +2
Query: 726 WKTT*MLFDVSCATFKAQANMVCIFIIHPLLP*FHIQMLIEVGAPTPDGLLLDTVSFL 783
W+ +*+L+ V C T + + C+FI H + +++E P LLL T+ FL
Sbjct: 491 WQLS*LLYQV*CHT*GSYF-VCCVFIAHLVYRFLGESLMLETLIWKPQTLLLMTICFL 661
>TC15474 UP|Q7Y1B9 (Q7Y1B9) Ammonium transporter, complete
Length = 1950
Score = 27.7 bits (60), Expect = 8.5
Identities = 18/61 (29%), Positives = 32/61 (51%), Gaps = 2/61 (3%)
Frame = -2
Query: 406 LGTLCLHLKTQIGKWSWMMNIML*LKIGR--GT*YLDHLMLMLFGPCGFLDIKKNLTVLL 463
LGT C+H++ +IG++ + GR G ++L+ PCG L I++N + +
Sbjct: 806 LGTHCVHVRLRIGRFG---------RTGRF*GLR*TRPIVLLCHRPCGLLRIQRNQSRRI 654
Query: 464 R 464
R
Sbjct: 653 R 651
>TC12608
Length = 735
Score = 27.7 bits (60), Expect = 8.5
Identities = 15/42 (35%), Positives = 28/42 (65%)
Frame = +3
Query: 545 YAFFANHSMGLNKLLELGVSDLPTTCHLLVFLKANVITLFSC 586
YA+ +HSM L +++ + V +P+ +LL+FL N+I ++ C
Sbjct: 465 YAY--HHSMQLWRIIIIRV--IPSR*YLLIFLNFNIIYIYIC 578
>TC11810
Length = 640
Score = 27.7 bits (60), Expect = 8.5
Identities = 15/36 (41%), Positives = 24/36 (66%)
Frame = +3
Query: 2 HSFGMIVWVIREKMF*ILLGKIS*LNVITLENYILI 37
HSF M+V +I ++MF*+L ++ *L ++ E Y I
Sbjct: 465 HSFHMLVDIILDRMF*VLP*ELL*LLILAHEYYCRI 572
>TC7840 similar to UP|AAQ72788 (AAQ72788) Hypersensitive-induced response
protein, complete
Length = 1585
Score = 27.7 bits (60), Expect = 8.5
Identities = 26/116 (22%), Positives = 50/116 (42%), Gaps = 5/116 (4%)
Frame = -2
Query: 272 YCQIKLLFVVMSYLMKILSHMQSCTFLNLIHTLS*TMSCPLILYNI*WTKPKLA-----H 326
+C + L ++ + + + SH QSC+ ++LIH T LI +NI H
Sbjct: 813 FCPLNLQYL-LCFSFLLCSHPQSCSSIDLIHGFLDT----LIWFNIHNKSLNYLIAIS*H 649
Query: 327 LILNWLISQTKSLLLVQTHQISPHLHNLTPLVPSNKIFLPYFNQLYKLTQNLSLGV 382
+ + ++ + +L+ +I HL N P + + ++ NL LG+
Sbjct: 648 SLFKFFLNCLCNFILLLKSRIQLHLRNTCP------------DNIKNISLNLCLGI 517
>TC13349
Length = 512
Score = 27.7 bits (60), Expect = 8.5
Identities = 18/70 (25%), Positives = 38/70 (53%), Gaps = 8/70 (11%)
Frame = +1
Query: 8 VWVIREKMF*ILLGKIS*LNVITLENYILIFVIH--VLLVNML------NCLS*ILIIEV 59
V V+ K+ +++ ++S + V+ + ++I I V+L+N L +L++E
Sbjct: 100 VAVVIMKIMVMIMARVSMMMVVVMRTLVMIIRIWR*VVLLNQL*EHHA*TYDMMVLVVEF 279
Query: 60 CCHLIFYIVM 69
C HL+ ++VM
Sbjct: 280 CHHLLCFVVM 309
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.346 0.153 0.515
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 19,229,949
Number of Sequences: 28460
Number of extensions: 325762
Number of successful extensions: 3748
Number of sequences better than 10.0: 33
Number of HSP's better than 10.0 without gapping: 3667
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 3746
length of query: 912
length of database: 4,897,600
effective HSP length: 99
effective length of query: 813
effective length of database: 2,080,060
effective search space: 1691088780
effective search space used: 1691088780
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.5 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 38 (21.7 bits)
S2: 59 (27.3 bits)
Medicago: description of AC126782.2


