

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC126781.3 + phase: 0
(77 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
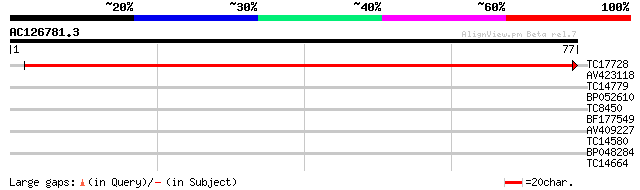
TC17728 similar to UP|Q9M011 (Q9M011) Light-inducible protein AT... 96 2e-21
AV423118 26 1.6
TC14779 similar to UP|O65833 (O65833) TCTR2 protein, partial (6%) 25 2.0
BP052610 25 2.6
TC8450 similar to UP|PDA1_VIGUN (O04865) Phospholipase D alpha 1... 24 4.5
BF177549 24 4.5
AV409227 24 4.5
TC14580 weakly similar to UP|RK21_SPIOL (P24613) 50S ribosomal p... 24 5.9
BP048284 23 7.7
TC14664 weakly similar to AAR24650 (AAR24650) At5g63160, partial... 23 7.7
>TC17728 similar to UP|Q9M011 (Q9M011) Light-inducible protein ATLS1,
complete
Length = 640
Score = 95.5 bits (236), Expect = 2e-21
Identities = 44/75 (58%), Positives = 59/75 (78%)
Frame = +2
Query: 3 ILVNGGVAIAFGGTEEPAAYGELISIGGLDPTVNAKLSSTIAQIIQTNLHIHSSRFYIKF 62
I++ G V ++FGGTE+PAAYGEL+SIGGL+P VN KLS+ IA I++T L + SRF++KF
Sbjct: 176 IVLKGSVPVSFGGTEQPAAYGELVSIGGLNPDVNKKLSAAIASILETKLSVPKSRFFLKF 355
Query: 63 SDVQPSFVGFNGSTF 77
D + S G+NGSTF
Sbjct: 356 YDTKGSNFGWNGSTF 400
>AV423118
Length = 472
Score = 25.8 bits (55), Expect = 1.6
Identities = 20/74 (27%), Positives = 32/74 (43%), Gaps = 13/74 (17%)
Frame = +2
Query: 2 CILVNGGVAIAF--GGTEEPAAYG-----------ELISIGGLDPTVNAKLSSTIAQIIQ 48
C++ G AIA G E AYG +L + G + +S I+ ++
Sbjct: 143 CVIATGYDAIALKSGWDEYGIAYGRPTENVHIRRVDLQASSGSALAFGSDMSGGISNVLV 322
Query: 49 TNLHIHSSRFYIKF 62
N H+H+S I+F
Sbjct: 323 ENAHLHNSNGGIEF 364
>TC14779 similar to UP|O65833 (O65833) TCTR2 protein, partial (6%)
Length = 572
Score = 25.4 bits (54), Expect = 2.0
Identities = 11/19 (57%), Positives = 14/19 (72%)
Frame = +3
Query: 9 VAIAFGGTEEPAAYGELIS 27
V A+GG+E PAA+G IS
Sbjct: 72 VCTAYGGSEAPAAFGYPIS 128
>BP052610
Length = 404
Score = 25.0 bits (53), Expect = 2.6
Identities = 12/29 (41%), Positives = 17/29 (58%)
Frame = +2
Query: 26 ISIGGLDPTVNAKLSSTIAQIIQTNLHIH 54
I IGG DPT + L+ + +Q+ Q H H
Sbjct: 164 IKIGG-DPTFSFVLAFSFSQVHQAGFHFH 247
>TC8450 similar to UP|PDA1_VIGUN (O04865) Phospholipase D alpha 1 (PLD
alpha 1) (Choline phosphatase 1)
(Phosphatidylcholine-hydrolyzing phospholipase D 1) ,
partial (73%)
Length = 2062
Score = 24.3 bits (51), Expect = 4.5
Identities = 19/75 (25%), Positives = 30/75 (39%), Gaps = 8/75 (10%)
Frame = +2
Query: 5 VNGGVAIAFGGTEEPAAYGELISIGGLDPTVNAKLSSTIAQIIQ--------TNLHIHSS 56
++GG A F + E AA LIS G D ++ + I+ N + S
Sbjct: 761 IDGGAAFGFPESPEDAARAGLIS--GKDNIIDRSIQDAYIHAIRRAKNFIYIENQYFLGS 934
Query: 57 RFYIKFSDVQPSFVG 71
F D++P +G
Sbjct: 935 SFAWSAEDIKPEDIG 979
>BF177549
Length = 440
Score = 24.3 bits (51), Expect = 4.5
Identities = 9/17 (52%), Positives = 13/17 (75%)
Frame = +2
Query: 49 TNLHIHSSRFYIKFSDV 65
+++HIHS FYIK + V
Sbjct: 296 SSIHIHSPLFYIKMTTV 346
>AV409227
Length = 427
Score = 24.3 bits (51), Expect = 4.5
Identities = 11/28 (39%), Positives = 16/28 (56%)
Frame = +1
Query: 14 GGTEEPAAYGELISIGGLDPTVNAKLSS 41
GG++ P YG + S G P+V + SS
Sbjct: 184 GGSDPPPGYGNVGSFKGASPSVPSWSSS 267
>TC14580 weakly similar to UP|RK21_SPIOL (P24613) 50S ribosomal protein L21,
chloroplast precursor (CL21) (CS-L7), partial (42%)
Length = 938
Score = 23.9 bits (50), Expect = 5.9
Identities = 10/19 (52%), Positives = 13/19 (67%)
Frame = +2
Query: 36 NAKLSSTIAQIIQTNLHIH 54
N STIA+++ TN HIH
Sbjct: 548 NINERSTIAELMGTNSHIH 604
>BP048284
Length = 168
Score = 23.5 bits (49), Expect = 7.7
Identities = 13/48 (27%), Positives = 23/48 (47%)
Frame = -2
Query: 15 GTEEPAAYGELISIGGLDPTVNAKLSSTIAQIIQTNLHIHSSRFYIKF 62
G ++ + L S+ GLD +++ +LH HS+ Y+KF
Sbjct: 146 GFDQVGLFIFLFSLDGLD------------KVVYMSLHDHSTYLYLKF 39
>TC14664 weakly similar to AAR24650 (AAR24650) At5g63160, partial (24%)
Length = 589
Score = 23.5 bits (49), Expect = 7.7
Identities = 11/18 (61%), Positives = 12/18 (66%)
Frame = -3
Query: 24 ELISIGGLDPTVNAKLSS 41
ELIS GG DPT +L S
Sbjct: 56 ELISAGGCDPTRKVQLLS 3
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.321 0.139 0.402
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 1,125,996
Number of Sequences: 28460
Number of extensions: 10499
Number of successful extensions: 61
Number of sequences better than 10.0: 20
Number of HSP's better than 10.0 without gapping: 61
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 61
length of query: 77
length of database: 4,897,600
effective HSP length: 53
effective length of query: 24
effective length of database: 3,389,220
effective search space: 81341280
effective search space used: 81341280
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 48 (23.1 bits)
Medicago: description of AC126781.3


