

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
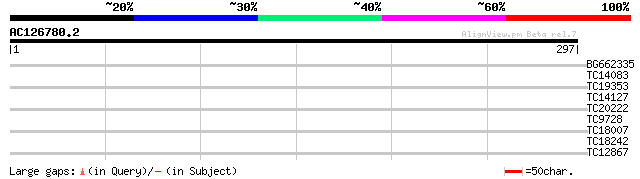
Query= AC126780.2 - phase: 0 /pseudo
(297 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
BG662335 37 0.003
TC14083 similar to UP|CRTC_RICCO (P93508) Calreticulin precursor... 29 0.84
TC19353 homologue to UP|AX2D_PHAAU (O24542) Auxin-induced protei... 27 4.1
TC14127 similar to UP|Q84UB0 (Q84UB0) Transcription factor Myb1,... 27 4.1
TC20222 similar to UP|Q9LFU8 (Q9LFU8) Proline-rich protein, part... 26 7.1
TC9728 similar to UP|Q94F45 (Q94F45) AT4g18020/T6K21_200, partia... 26 7.1
TC18007 similar to PIR|T01245|T01245 N-acetyltransferase homolog... 26 9.2
TC18242 weakly similar to UP|O70495 (O70495) Plenty-of-prolines-... 26 9.2
TC12867 similar to GB|AAL91627.1|19715609|AY075613 AT3g59630/T16... 26 9.2
>BG662335
Length = 336
Score = 37.4 bits (85), Expect = 0.003
Identities = 22/59 (37%), Positives = 31/59 (52%), Gaps = 1/59 (1%)
Frame = +1
Query: 100 GCILGLRVSPIHRRKGVGLKLVTSIEEWM-LTNGADYAFLATEKNNNASKNLFTNKCNY 157
G I L V HR+ G+ KL+T+ + M GA+Y L K+N A+ NL+T Y
Sbjct: 22 GHITSLAVLRTHRKLGLATKLMTAAQNAMEQVFGAEYVSLHVRKSNRAAFNLYTETLGY 198
>TC14083 similar to UP|CRTC_RICCO (P93508) Calreticulin precursor, partial
(92%)
Length = 1693
Score = 29.3 bits (64), Expect = 0.84
Identities = 18/47 (38%), Positives = 25/47 (52%)
Frame = +1
Query: 175 TNHISKKDVKIDKISIDQAISFYTRILKTKELYPLDMDIILKEKLSL 221
TNH+ KKDV + DQ YT IL+ Y + +D + K+ SL
Sbjct: 574 TNHLIKKDVPCE---TDQLTHVYTFILRPDATYSILIDNVEKQTGSL 705
>TC19353 homologue to UP|AX2D_PHAAU (O24542) Auxin-induced protein 22D
(Indole-3-acetic acid induced protein ARG13), partial
(41%)
Length = 560
Score = 26.9 bits (58), Expect = 4.1
Identities = 16/56 (28%), Positives = 23/56 (40%)
Frame = +1
Query: 200 ILKTKELYPLDMDIILKEKLSLGTWVSYYKDEGFKLNIEDIITHKSTTSSSWIIFS 255
ILK K L+M ++K K+ G W+ + L K W+IFS
Sbjct: 76 ILKEKATMDLNMSQLMKIKMVTGCWLEMFHGTCLCLPARG*EL*KDQKQRGWVIFS 243
>TC14127 similar to UP|Q84UB0 (Q84UB0) Transcription factor Myb1, partial
(21%)
Length = 556
Score = 26.9 bits (58), Expect = 4.1
Identities = 11/24 (45%), Positives = 15/24 (61%)
Frame = -3
Query: 143 NNNASKNLFTNKCNYFNFTSLIIF 166
NNN N F N+ +YF+F L I+
Sbjct: 473 NNNXPSNQFKNQQSYFHFNRLRIY 402
>TC20222 similar to UP|Q9LFU8 (Q9LFU8) Proline-rich protein, partial (5%)
Length = 521
Score = 26.2 bits (56), Expect = 7.1
Identities = 21/48 (43%), Positives = 28/48 (57%), Gaps = 1/48 (2%)
Frame = +3
Query: 23 KVVGKLERNC-TEINGTTKKGFSIFTNMMSNGDPLSRIRFYPLHVMLV 69
KV L R+ T IN TTK+G SIF+ + D L I + PL V+L+
Sbjct: 231 KVESLLARDMETSINNTTKEGRSIFSTV----DTL--IHYTPLPVVLL 356
>TC9728 similar to UP|Q94F45 (Q94F45) AT4g18020/T6K21_200, partial (5%)
Length = 636
Score = 26.2 bits (56), Expect = 7.1
Identities = 16/54 (29%), Positives = 24/54 (43%), Gaps = 12/54 (22%)
Frame = +2
Query: 156 NYFNFTSLIIFLHP------------PTSFPTNHISKKDVKIDKISIDQAISFY 197
++FN S +IF H P F T+HI + KI K+S+D +
Sbjct: 212 SFFNTLSTLIFQHSFTQSNSCVPTVRPI*FSTSHIFYDE*KIVKVSVDSICQLF 373
>TC18007 similar to PIR|T01245|T01245 N-acetyltransferase homolog F16M14.6 -
Arabidopsis thaliana {Arabidopsis thaliana;}, partial
(66%)
Length = 456
Score = 25.8 bits (55), Expect = 9.2
Identities = 16/52 (30%), Positives = 25/52 (47%)
Frame = +2
Query: 100 GCILGLRVSPIHRRKGVGLKLVTSIEEWMLTNGADYAFLATEKNNNASKNLF 151
G I L V +R KG+ +LVT + M+ +G + L E N + L+
Sbjct: 299 GYIAMLVVIKPYRGKGIATELVTRSIQVMMESGCEEVTLEAEVTNKGALALY 454
>TC18242 weakly similar to UP|O70495 (O70495) Plenty-of-prolines-101,
partial (5%)
Length = 490
Score = 25.8 bits (55), Expect = 9.2
Identities = 11/28 (39%), Positives = 14/28 (49%)
Frame = -2
Query: 140 TEKNNNASKNLFTNKCNYFNFTSLIIFL 167
T N N+F N C Y +F + IFL
Sbjct: 129 TGNQNEKKNNVFWNSCFYLDFYYVCIFL 46
>TC12867 similar to GB|AAL91627.1|19715609|AY075613 AT3g59630/T16L24_180
{Arabidopsis thaliana;}, partial (16%)
Length = 788
Score = 25.8 bits (55), Expect = 9.2
Identities = 20/64 (31%), Positives = 31/64 (48%), Gaps = 10/64 (15%)
Frame = +3
Query: 145 NASKNLFTNK-----CNYFNFTSLIIFLHPPTSFPTNHISKKDVKID-----KISIDQAI 194
+A KN T K C YF+F +LI PPT+ H S V++ + ++D
Sbjct: 66 SACKNKITVKSVIVMCKYFSFLTLISPNPPPTTASLLHPSS*GVQLRGSGNLREAVDFLC 245
Query: 195 SFYT 198
+FY+
Sbjct: 246 NFYS 257
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.321 0.137 0.405
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 5,477,097
Number of Sequences: 28460
Number of extensions: 79125
Number of successful extensions: 410
Number of sequences better than 10.0: 18
Number of HSP's better than 10.0 without gapping: 410
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 410
length of query: 297
length of database: 4,897,600
effective HSP length: 90
effective length of query: 207
effective length of database: 2,336,200
effective search space: 483593400
effective search space used: 483593400
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 55 (25.8 bits)
Medicago: description of AC126780.2


