

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC126018.36 + phase: 0
(475 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
Score E
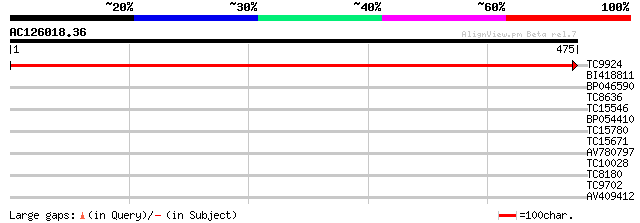
Sequences producing significant alignments: (bits) Value
TC9924 UP|RBL_LOTJA (Q9BBU1) Ribulose bisphosphate carboxylase l... 953 0.0
BI418811 29 1.5
BP046590 29 1.9
TC8636 28 3.3
TC15546 similar to GB|AAC98030.1|4056457|F5O8 ESTs gb|234051 and... 28 3.3
BP054410 28 4.3
TC15780 similar to UP|Q8VAD1 (Q8VAD1) Wsv494 (WSSV020), partial ... 28 4.3
TC15671 28 4.3
AV780797 27 7.3
TC10028 weakly similar to UP|Q9LXX7 (Q9LXX7) Cytochrome P450-lik... 27 9.5
TC8180 similar to UP|EXTN_TOBAC (P13983) Extensin precursor (Cel... 27 9.5
TC9702 27 9.5
AV409412 27 9.5
>TC9924 UP|RBL_LOTJA (Q9BBU1) Ribulose bisphosphate carboxylase large chain
precursor (RuBisCO large subunit) , complete
Length = 1592
Score = 953 bits (2464), Expect = 0.0
Identities = 459/475 (96%), Positives = 470/475 (98%)
Frame = +3
Query: 1 MSPQTETKATVGFKAGVKDYRLTYYTPDYETKDTDILAAFRVSPQPGVPAEEAGAAVAAE 60
MSPQTETKA+VGFKAGVKDY+LTYYTPDYETKDTDILAAFRV+PQPGVP EEAGAAVAAE
Sbjct: 165 MSPQTETKASVGFKAGVKDYKLTYYTPDYETKDTDILAAFRVTPQPGVPPEEAGAAVAAE 344
Query: 61 SSTGTWTTVWTDGLTSLDRYKGRCYHIEPVAGEESQFIAYVAYPLDLFEEGSVTNMFTSI 120
SSTGTWTTVWTDGLTSLDRYKGRCYHIEPV GEESQFIAYVAYPLDLFEEGSVTNMFTSI
Sbjct: 345 SSTGTWTTVWTDGLTSLDRYKGRCYHIEPVPGEESQFIAYVAYPLDLFEEGSVTNMFTSI 524
Query: 121 VGNVFGFKALRALRLEDLRIPVAYVKTFQGPPHGIQVERDKLNKYGRPLLGCTIKPKLGL 180
VGNVFGFKALRALRLEDLRIP AYVKTFQGPPHGIQVERDKLNKYGRPLLGCTIKPKLGL
Sbjct: 525 VGNVFGFKALRALRLEDLRIPNAYVKTFQGPPHGIQVERDKLNKYGRPLLGCTIKPKLGL 704
Query: 181 SAKNYGRAVYECLRGGLDFTKDDENVNSQPFMRWRDRFLFCAEAIYKAQAETGEIKGHYL 240
SAKNYGRAVYECLRGGLDFTKDDENVNSQPFMRWRDRFLFCAEA++KAQ ETGEIKGHYL
Sbjct: 705 SAKNYGRAVYECLRGGLDFTKDDENVNSQPFMRWRDRFLFCAEALFKAQEETGEIKGHYL 884
Query: 241 NATAGTCEDMMKRAVFARELGVPIVMHDYLTGGFTANTTLAHYCRDNGLLLHIHRAMHAV 300
NATAGTCE+MMKRAVFARELGVPIVMHDYLTGGFTANTTLAHYCRDNGLLLHIHRAMHAV
Sbjct: 885 NATAGTCEEMMKRAVFARELGVPIVMHDYLTGGFTANTTLAHYCRDNGLLLHIHRAMHAV 1064
Query: 301 IDRQKNHGMHFRVLAKALRMSGGDHIHAGTVVGKLEGERDITLGFVDLLRDDFVEKDRSR 360
IDRQKNHGMHFRVLAKALRMSGGDHIHAGTVVGKLEGER+ITLGFVDLLRDDFVEKDRSR
Sbjct: 1065IDRQKNHGMHFRVLAKALRMSGGDHIHAGTVVGKLEGEREITLGFVDLLRDDFVEKDRSR 1244
Query: 361 GIFFTQDWVSLPGVLPVASGGIHVWHMPALTEIFGDDSVLQFGGGTLGHPWGNAPGAVAN 420
GI+FTQDWVSLPGVLPVASGGIHVWHMPALTEIFGDDSVLQFGGGTLGHPWGNAPGAVAN
Sbjct: 1245GIYFTQDWVSLPGVLPVASGGIHVWHMPALTEIFGDDSVLQFGGGTLGHPWGNAPGAVAN 1424
Query: 421 RVALEACVQARNEGRDLAREGNEIIREATKWSPELAAACEVWKEIKFEFPAMDTI 475
RVALEACVQARNEGRDLAREGNEIIREA+KWSPELAAACE+WKEIKFEF AMDT+
Sbjct: 1425RVALEACVQARNEGRDLAREGNEIIREASKWSPELAAACEIWKEIKFEFQAMDTL 1589
>BI418811
Length = 462
Score = 29.3 bits (64), Expect = 1.5
Identities = 14/36 (38%), Positives = 23/36 (63%), Gaps = 1/36 (2%)
Frame = -3
Query: 280 LAHYCRDNGL-LLHIHRAMHAVIDRQKNHGMHFRVL 314
L H CR+ L LLH+HR + ++ R + H +H ++L
Sbjct: 412 LRHPCRNLLLFLLHLHRFIITLLRRHRGHLLHLKLL 305
>BP046590
Length = 498
Score = 28.9 bits (63), Expect = 1.9
Identities = 14/31 (45%), Positives = 17/31 (54%), Gaps = 4/31 (12%)
Frame = -3
Query: 184 NYGRAVYECLRGGLDF----TKDDENVNSQP 210
NYGR Y+C GG D+ E VN+QP
Sbjct: 376 NYGRTFYQCSYGGCDYFDWAEGPIEGVNNQP 284
>TC8636
Length = 563
Score = 28.1 bits (61), Expect = 3.3
Identities = 16/55 (29%), Positives = 27/55 (49%), Gaps = 2/55 (3%)
Frame = -3
Query: 280 LAHYCRDNGLLLHIHRAMHAVIDRQKNHGMHFRVLAKALRMSGG--DHIHAGTVV 332
+ +Y L+LH+ M + +RQ G V ++ LRMS +H+G V+
Sbjct: 489 ITYYT*HKSLILHLRIKMMILEERQTKFGTRIYVKSEKLRMSVSLIKKLHSGAVI 325
>TC15546 similar to GB|AAC98030.1|4056457|F5O8 ESTs gb|234051 and gb|F13722
come from this gene. {Arabidopsis thaliana;}, partial
(96%)
Length = 985
Score = 28.1 bits (61), Expect = 3.3
Identities = 19/77 (24%), Positives = 33/77 (42%), Gaps = 13/77 (16%)
Frame = +1
Query: 412 GNAPGAVANRVALEACV----------QARNEGRDLAREGNEIIREATK---WSPELAAA 458
G A GA ++ L C+ ARN+ DL +EG+ ++ K + + A
Sbjct: 202 GRADGAQPRQMRLAECLVGDETGMIIFTARNDQVDLMKEGSTVVMRNAKIDMYKGSMRLA 381
Query: 459 CEVWKEIKFEFPAMDTI 475
+ W ++ PA T+
Sbjct: 382 VDKWGRVEVAEPASFTV 432
>BP054410
Length = 572
Score = 27.7 bits (60), Expect = 4.3
Identities = 11/22 (50%), Positives = 14/22 (63%)
Frame = +3
Query: 103 YPLDLFEEGSVTNMFTSIVGNV 124
YPL L+ +G + N F S GNV
Sbjct: 165 YPLSLYMKGGILNRFLSSTGNV 230
>TC15780 similar to UP|Q8VAD1 (Q8VAD1) Wsv494 (WSSV020), partial (29%)
Length = 653
Score = 27.7 bits (60), Expect = 4.3
Identities = 8/23 (34%), Positives = 14/23 (60%)
Frame = -1
Query: 199 FTKDDENVNSQPFMRWRDRFLFC 221
+TK ++ QP + W D ++FC
Sbjct: 446 YTKGKHMISIQPLLNWSDSYIFC 378
>TC15671
Length = 546
Score = 27.7 bits (60), Expect = 4.3
Identities = 20/55 (36%), Positives = 25/55 (45%), Gaps = 3/55 (5%)
Frame = +2
Query: 403 GGGTLGHPWGNAPGAVANRVALEAC--VQARNEGRDLAREGNEII-REATKWSPE 454
GGG LG + A+ RVA E C ARN G + ++ E RE W E
Sbjct: 41 GGGVLGGEEADGGAAIGERVAEERCGGSPARNRGWKVQKD*EECSQREEH*WEKE 205
>AV780797
Length = 611
Score = 26.9 bits (58), Expect = 7.3
Identities = 11/21 (52%), Positives = 13/21 (61%)
Frame = -1
Query: 53 AGAAVAAESSTGTWTTVWTDG 73
AGAAV + T + T VW DG
Sbjct: 305 AGAAVTGDHHTNSGTAVWVDG 243
>TC10028 weakly similar to UP|Q9LXX7 (Q9LXX7) Cytochrome P450-like protein,
partial (16%)
Length = 874
Score = 26.6 bits (57), Expect = 9.5
Identities = 10/19 (52%), Positives = 14/19 (73%)
Frame = -2
Query: 136 EDLRIPVAYVKTFQGPPHG 154
E++ +PVA V+ F GP HG
Sbjct: 273 EEVGVPVAEVEEFAGPAHG 217
>TC8180 similar to UP|EXTN_TOBAC (P13983) Extensin precursor (Cell wall
hydroxyproline-rich glycoprotein), partial (3%)
Length = 575
Score = 26.6 bits (57), Expect = 9.5
Identities = 12/33 (36%), Positives = 19/33 (57%)
Frame = +2
Query: 433 EGRDLAREGNEIIREATKWSPELAAACEVWKEI 465
EG+ + E + +EA + PEL+AA E K +
Sbjct: 113 EGKKMEEEAKKAYQEALQLEPELSAAQEALKRL 211
>TC9702
Length = 927
Score = 26.6 bits (57), Expect = 9.5
Identities = 11/30 (36%), Positives = 19/30 (62%)
Frame = +3
Query: 444 IIREATKWSPELAAACEVWKEIKFEFPAMD 473
++R + K++ ++ CEV K I F FPA +
Sbjct: 504 LVRRSIKYARIVSELCEVLK*IGFRFPAFE 593
>AV409412
Length = 426
Score = 26.6 bits (57), Expect = 9.5
Identities = 17/47 (36%), Positives = 22/47 (46%)
Frame = -2
Query: 395 GDDSVLQFGGGTLGHPWGNAPGAVANRVALEACVQARNEGRDLAREG 441
GDD ++ GGG G G G V VA E + +GR+ EG
Sbjct: 404 GDD--VEGGGGGGGEEGGEEGGGVGGEVADEVDGEGDGDGREREGEG 270
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.321 0.138 0.427
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 8,040,381
Number of Sequences: 28460
Number of extensions: 105357
Number of successful extensions: 446
Number of sequences better than 10.0: 26
Number of HSP's better than 10.0 without gapping: 444
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 446
length of query: 475
length of database: 4,897,600
effective HSP length: 94
effective length of query: 381
effective length of database: 2,222,360
effective search space: 846719160
effective search space used: 846719160
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 57 (26.6 bits)
Medicago: description of AC126018.36


