

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
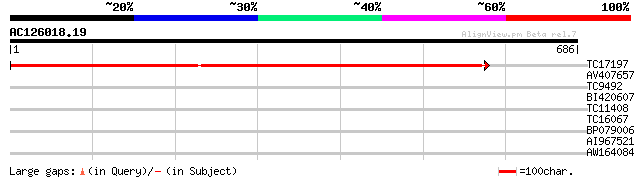
Query= AC126018.19 - phase: 0
(686 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
TC17197 UP|RPOC_LOTJA (Q9BBS8) DNA-directed RNA polymerase beta'... 1106 0.0
AV407657 31 0.76
TC9492 homologue to UP|Q9MBA2 (Q9MBA2) MinD (Septum site-determi... 28 3.8
BI420607 28 4.9
TC11408 28 4.9
TC16067 UP|Q9ZRY4 (Q9ZRY4) NDX1 homeobox protein, complete 28 6.4
BP079006 28 6.4
AI967521 27 8.4
AW164084 27 8.4
>TC17197 UP|RPOC_LOTJA (Q9BBS8) DNA-directed RNA polymerase beta' chain
(PEP) (Plastid-encoded RNA polymerase beta' subunit)
(RNA polymerase beta' subunit) , partial (84%)
Length = 1728
Score = 1106 bits (2861), Expect = 0.0
Identities = 546/580 (94%), Positives = 559/580 (96%)
Frame = +1
Query: 1 MIDQYKHQHLRIGSVSPEQISAWAKKILPNGEIVGEVTKPYTLHYKTNKPEKDGLFCERI 60
MIDQYKHQ LRIGSVSP+QISAWA KILPNGEIVGEVTKPYT HYKTNKPEKDGLFCERI
Sbjct: 1 MIDQYKHQQLRIGSVSPQQISAWATKILPNGEIVGEVTKPYTFHYKTNKPEKDGLFCERI 180
Query: 61 FGPIKSGICACGNYRVIGDKKDQPKFCEQCGVEFVDSRVRRYQMGYIKLACPVTHVWYLK 120
FGPIKSGIC CGNYRVIGDKKD+PKFCEQCGVEFVDSR+RRYQMGYIKLACPVTHVWYLK
Sbjct: 181 FGPIKSGICTCGNYRVIGDKKDEPKFCEQCGVEFVDSRIRRYQMGYIKLACPVTHVWYLK 360
Query: 121 RLPSYIASLLDKPLKELEGLVYCDFSFARPVVKKPTFLRLRGSFEYEIQSWKYSIPLFFA 180
RLPSYIA+LLDKPLKELEGLVYCDFSFARPVVKKPTFLRLRG FEYEIQSWKYSIPLFFA
Sbjct: 361 RLPSYIANLLDKPLKELEGLVYCDFSFARPVVKKPTFLRLRGLFEYEIQSWKYSIPLFFA 540
Query: 181 TQGFDTFRNREISSGAGAIREQLVDLDLRIIMDSSLVEWKELGEEGSADNENENEWEDRK 240
T GFDTFRNREISSGAGAIREQLVDLDLRI++DSSLVEWKELGEE S NENEWEDRK
Sbjct: 541 THGFDTFRNREISSGAGAIREQLVDLDLRIVIDSSLVEWKELGEERST--HNENEWEDRK 714
Query: 241 VGRRKNFLVRRMELVKHFIRTNIEPEWMVLSLLPVLPPELRPIIQIDGGKLMSSDINELY 300
VGRRKNFLVRRMEL KHFIRTNIEPEWMVL LLPVLPPELRPIIQIDGGKLMSSDINELY
Sbjct: 715 VGRRKNFLVRRMELAKHFIRTNIEPEWMVLCLLPVLPPELRPIIQIDGGKLMSSDINELY 894
Query: 301 RRVIYRNNTLIDLLTTSRSTPGELVMCQEKLVQEAVDTLLDNGIRGQPMRDGHNKVYKSF 360
RRVIYRNNTLIDLLTTSRSTPGELVMCQEKLVQEAVDTLLDNGIRGQPM+DGHNKVYKSF
Sbjct: 895 RRVIYRNNTLIDLLTTSRSTPGELVMCQEKLVQEAVDTLLDNGIRGQPMKDGHNKVYKSF 1074
Query: 361 SDIIEGKEGRFRETLLGKRVDYSGRSVIVVGPSLSLHRCGLPREIAIELFQTFLIRGLIR 420
SDIIEGKEGRFRETLLGKRVDYSGRSVIVVGPSLSLHRCGLPREIAIELFQTFLIR LIR
Sbjct: 1075SDIIEGKEGRFRETLLGKRVDYSGRSVIVVGPSLSLHRCGLPREIAIELFQTFLIRSLIR 1254
Query: 421 KHFASNIGVAKSKIREKEPIVWEILQEVMRGHPVLLNRAPTLHRLGIQAFQPILVEGRAI 480
KHFASNIGVAKSKIREKEPIVWEILQEVMRGHP+LLNRAPTLHRLGIQAFQPILVEGRAI
Sbjct: 1255KHFASNIGVAKSKIREKEPIVWEILQEVMRGHPILLNRAPTLHRLGIQAFQPILVEGRAI 1434
Query: 481 CLHPLVCKGFNADFDGDQMAVHVPLSLEAQAEARLLMFSHTNLLSPAIGDPISVPTQDML 540
CLHPLVCKGFNADFDGDQMA+HVPLSLEAQAEARLLMFSHTNLLSPAIGDPI+VPTQDML
Sbjct: 1435CLHPLVCKGFNADFDGDQMAIHVPLSLEAQAEARLLMFSHTNLLSPAIGDPIAVPTQDML 1614
Query: 541 IGLYVLTSGNRRGICANRYNPFNCRNSKNEKISNNNSKYM 580
IGLY+LTSGNRRGICANRYN N +NSKNEKI + KYM
Sbjct: 1615IGLYILTSGNRRGICANRYNRSNWKNSKNEKI--RDKKYM 1728
>AV407657
Length = 426
Score = 30.8 bits (68), Expect = 0.76
Identities = 14/31 (45%), Positives = 21/31 (67%)
Frame = +2
Query: 372 RETLLGKRVDYSGRSVIVVGPSLSLHRCGLP 402
++ +LGKR D S R+V+V P L+L G+P
Sbjct: 329 KDVVLGKRNDCSFRTVVVGDPDLALSEIGVP 421
>TC9492 homologue to UP|Q9MBA2 (Q9MBA2) MinD (Septum site-determining MinD)
(At5g24020), partial (33%)
Length = 633
Score = 28.5 bits (62), Expect = 3.8
Identities = 18/57 (31%), Positives = 30/57 (52%), Gaps = 3/57 (5%)
Frame = +3
Query: 443 EILQEVMRGHPVLLNRAPTLHRLGIQAFQPILVEG---RAICLHPLVCKGFNADFDG 496
E+++ RG P++LN+ PTL L + LVE +A+ + +GF + F G
Sbjct: 159 EVIRSTNRGFPLVLNKPPTLAGLAFEQAAWRLVEQDTMQAVMVEEEPKRGFFSFFGG 329
>BI420607
Length = 408
Score = 28.1 bits (61), Expect = 4.9
Identities = 17/58 (29%), Positives = 34/58 (58%), Gaps = 1/58 (1%)
Frame = -1
Query: 220 KELGEEGSAD-NENENEWEDRKVGRRKNFLVRRMELVKHFIRTNIEPEWMVLSLLPVL 276
++ G+EG + ++ E EW +++ +R+ R + FI+ N+ E++VLS+L L
Sbjct: 213 QDTGKEGRREWSKIEEEW*EKQERKRERERERECCVFFLFIQLNMRLEFLVLSILRFL 40
>TC11408
Length = 580
Score = 28.1 bits (61), Expect = 4.9
Identities = 12/30 (40%), Positives = 17/30 (56%)
Frame = -1
Query: 560 NPFNCRNSKNEKISNNNSKYMKKKEPFFCN 589
+PF CRNS + KI N+S + + CN
Sbjct: 157 HPFTCRNSSSNKIHVNSSSRVIRCRLLHCN 68
>TC16067 UP|Q9ZRY4 (Q9ZRY4) NDX1 homeobox protein, complete
Length = 3271
Score = 27.7 bits (60), Expect = 6.4
Identities = 10/33 (30%), Positives = 18/33 (54%)
Frame = -1
Query: 552 RGICANRYNPFNCRNSKNEKISNNNSKYMKKKE 584
R I N +PF CR + N K++ + +++E
Sbjct: 100 RWILLNNLHPFKCRRASNSKLNRRRRRRERERE 2
>BP079006
Length = 388
Score = 27.7 bits (60), Expect = 6.4
Identities = 9/22 (40%), Positives = 14/22 (62%)
Frame = -3
Query: 482 LHPLVCKGFNADFDGDQMAVHV 503
+HP+ C+GF D G+Q H+
Sbjct: 221 VHPVACRGFQVDQSGNQCTRHM 156
>AI967521
Length = 436
Score = 27.3 bits (59), Expect = 8.4
Identities = 21/65 (32%), Positives = 29/65 (44%)
Frame = +1
Query: 167 EIQSWKYSIPLFFATQGFDTFRNREISSGAGAIREQLVDLDLRIIMDSSLVEWKELGEEG 226
E+QSW YS P + F + R SG +R++ +D D I + V GE G
Sbjct: 253 EVQSWPYSFP---DSDDFSKWDERGNVSGRLLVRDRYIDDDY-ISAKGAYVGLAPPGEVG 420
Query: 227 SADNE 231
S E
Sbjct: 421 SWQEE 435
>AW164084
Length = 394
Score = 27.3 bits (59), Expect = 8.4
Identities = 16/51 (31%), Positives = 21/51 (40%), Gaps = 12/51 (23%)
Frame = +3
Query: 151 VVKKPTFLRLRGSFEYEIQSW------------KYSIPLFFATQGFDTFRN 189
V+KKP L + G F+Y I W K P+F+ TF N
Sbjct: 153 VLKKPLCLDVEGPFKYGIYQWALPVISSNSLAVKILYPIFWGLMTLSTFGN 305
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.323 0.141 0.428
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 12,409,354
Number of Sequences: 28460
Number of extensions: 179492
Number of successful extensions: 756
Number of sequences better than 10.0: 18
Number of HSP's better than 10.0 without gapping: 751
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 755
length of query: 686
length of database: 4,897,600
effective HSP length: 96
effective length of query: 590
effective length of database: 2,165,440
effective search space: 1277609600
effective search space used: 1277609600
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.9 bits)
S2: 58 (26.9 bits)
Medicago: description of AC126018.19


