

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
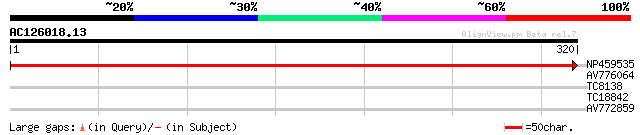
Query= AC126018.13 - phase: 0
(320 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
NP459535 cytochrome f [Lotus japonicus] 607 e-175
AV776064 29 1.2
TC8138 weakly similar to UP|Q84630 (Q84630) A316R protein, parti... 27 4.6
TC18842 similar to UP|Q8RVQ6 (Q8RVQ6) SEC6, partial (31%) 27 6.0
AV772859 27 6.0
>NP459535 cytochrome f [Lotus japonicus]
Length = 963
Score = 607 bits (1566), Expect = e-175
Identities = 304/320 (95%), Positives = 314/320 (98%)
Frame = +1
Query: 1 MQTRNAFSWIKEEITRSISVLLMIYIITRAPISNAYPIFAQQGYENPREATGRIVCANCH 60
MQTRNAFS+IKEEITRSISVLL+IYII RAPISNAYPIFAQQGYENPREATGRIVCANCH
Sbjct: 1 MQTRNAFSYIKEEITRSISVLLVIYIIIRAPISNAYPIFAQQGYENPREATGRIVCANCH 180
Query: 61 LANKPVDIEVPQAVLPDTVFEAVVRIPYDMQVKQVLANGKKGALNVGAVLILPEGFELAP 120
LANKPVDIEVPQ VLPDTVFEAVVRIPYDMQVKQVLANGK+GALNVGAVLILPEGFELAP
Sbjct: 181 LANKPVDIEVPQTVLPDTVFEAVVRIPYDMQVKQVLANGKRGALNVGAVLILPEGFELAP 360
Query: 121 PDRISPEIKEKIGNLSFQSYRPTKKNILVVGPVPGKKYSEITFPILSPDPATKRDVHFLK 180
DRISPEIKEK+GNLSFQSYRPTKKNILVVGPVPG+KYSEITFPILSPDPAT RDV+FLK
Sbjct: 361 TDRISPEIKEKMGNLSFQSYRPTKKNILVVGPVPGQKYSEITFPILSPDPATNRDVNFLK 540
Query: 181 YPIYVGGNRGRGQIYPDGSKSNNNVYNATATGIVNKIIRKEKGGYEITIVDASDGREVID 240
YPIYVGGNRGRGQIYPDGSKSNNNVYNAT +GI+NKIIRK+KGGYEITIVDASDGREVID
Sbjct: 541 YPIYVGGNRGRGQIYPDGSKSNNNVYNATTSGIINKIIRKDKGGYEITIVDASDGREVID 720
Query: 241 IIPPGPELLVSEGESMKLDQPLTSNPNVGGFGQGDAEIVLQDPLRVQGLLFFLASIILAQ 300
IIPPGPELLVSEGES+KLDQPLTSNPNVGGFGQGDAEIVLQDPLRVQGLLFFLASII AQ
Sbjct: 721 IIPPGPELLVSEGESIKLDQPLTSNPNVGGFGQGDAEIVLQDPLRVQGLLFFLASIIFAQ 900
Query: 301 IFLVLKKKQFEKVQLSEMNF 320
IFLVLKKKQFEKVQLSEMNF
Sbjct: 901 IFLVLKKKQFEKVQLSEMNF 960
>AV776064
Length = 418
Score = 28.9 bits (63), Expect = 1.2
Identities = 21/71 (29%), Positives = 36/71 (50%), Gaps = 5/71 (7%)
Frame = -2
Query: 93 KQVLANGKKGALNVGAVLILPEGFELAPPDR-ISPEIKEKIGNLSFQSYRPTKK----NI 147
+++L G+KG +G +L +PE P +R E++ K+ L ++ RP K+ +
Sbjct: 366 RRLLRGGRKG*SRIGNLLPVPE-----PGNRHTHNELENKVSRLEEENERPRKRKELEQM 202
Query: 148 LVVGPVPGKKY 158
L P P KY
Sbjct: 201 LPCTPPPEPKY 169
>TC8138 weakly similar to UP|Q84630 (Q84630) A316R protein, partial (36%)
Length = 928
Score = 26.9 bits (58), Expect = 4.6
Identities = 15/44 (34%), Positives = 19/44 (43%)
Frame = +3
Query: 141 RPTKKNILVVGPVPGKKYSEITFPILSPDPATKRDVHFLKYPIY 184
+P K I V P+P K P+L P P K + PIY
Sbjct: 375 KPIPKPIFVKKPIPFYKPVPKVIPVLKPIPIYKPVPKVIPVPIY 506
>TC18842 similar to UP|Q8RVQ6 (Q8RVQ6) SEC6, partial (31%)
Length = 711
Score = 26.6 bits (57), Expect = 6.0
Identities = 12/29 (41%), Positives = 20/29 (68%), Gaps = 1/29 (3%)
Frame = -3
Query: 158 YSEITFPI-LSPDPATKRDVHFLKYPIYV 185
Y+ T+P LSP+P+ ++FLK P+Y+
Sbjct: 226 YAHSTYPCSLSPEPSIFAVLNFLKRPLYL 140
>AV772859
Length = 329
Score = 26.6 bits (57), Expect = 6.0
Identities = 11/22 (50%), Positives = 15/22 (68%)
Frame = +2
Query: 187 GNRGRGQIYPDGSKSNNNVYNA 208
G+R G+IYP SKS + YN+
Sbjct: 38 GSRATGKIYPHKSKSQSG*YNS 103
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.318 0.140 0.401
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 4,727,036
Number of Sequences: 28460
Number of extensions: 58743
Number of successful extensions: 252
Number of sequences better than 10.0: 10
Number of HSP's better than 10.0 without gapping: 250
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 252
length of query: 320
length of database: 4,897,600
effective HSP length: 90
effective length of query: 230
effective length of database: 2,336,200
effective search space: 537326000
effective search space used: 537326000
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 55 (25.8 bits)
Medicago: description of AC126018.13


