

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC126018.1 + phase: 0 /pseudo
(596 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
Score E
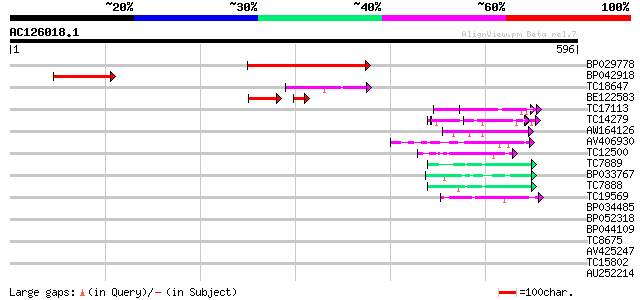
Sequences producing significant alignments: (bits) Value
BP029778 188 2e-48
BP042918 82 3e-16
TC18647 similar to UP|Q9ASR9 (Q9ASR9) At2g01320/F10A8.20, partia... 60 1e-09
BE122583 47 3e-07
TC17113 weakly similar to UP|Q9LFU8 (Q9LFU8) Proline-rich protei... 47 9e-06
TC14279 similar to UP|Q09085 (Q09085) Hydroxyproline-rich glycop... 46 2e-05
AW164126 43 2e-04
AV406930 43 2e-04
TC12500 similar to GB|AAH45354.1|28279536|BC045354 FNBP4 protein... 42 3e-04
TC7889 similar to UP|Q40336 (Q40336) Proline-rich cell wall prot... 42 3e-04
BP033767 41 5e-04
TC7888 similar to UP|Q40336 (Q40336) Proline-rich cell wall prot... 40 8e-04
TC19569 similar to UP|AAR11301 (AAR11301) Lectin-like receptor k... 40 8e-04
BP034485 40 0.001
BP052318 40 0.001
BP044109 40 0.001
TC8675 weakly similar to UP|Q9LFH0 (Q9LFH0) PDR9 ABC transporter... 40 0.001
AV425247 39 0.002
TC15802 similar to UP|Q9XIV1 (Q9XIV1) mRNA expressed in cucumber... 39 0.003
AU252214 39 0.003
>BP029778
Length = 438
Score = 188 bits (478), Expect = 2e-48
Identities = 91/129 (70%), Positives = 107/129 (82%)
Frame = -2
Query: 251 KKKGMVLPFEPHSITFDDIVYSVDMPVEMKEQGVREDRLVLLKGVSGAFRPGVLTALMGV 310
K KGM+LPF+ ++TF ++ Y VDMP E+++QG+ E +L LL VSG F PGVLTALMG
Sbjct: 398 KNKGMILPFQQFTVTFHNVNYYVDMPQEIRKQGIIETKLKLLSNVSGVFAPGVLTALMGS 219
Query: 311 SGAGKTTLMDVLAGRKTGGYIDGDIKVSGYPKKQETFARISGYCEHNDIHSPHVTVHESL 370
SGAGKTTLMDVLAGRKTGGYI+GDIK+SGYPK Q TFAR++GY E NDIHSP VTV ESL
Sbjct: 218 SGAGKTTLMDVLAGRKTGGYIEGDIKISGYPKVQHTFARVAGYVEQNDIHSPQVTVEESL 39
Query: 371 LYSAWLRLP 379
L+SA LRLP
Sbjct: 38 LFSALLRLP 12
>BP042918
Length = 412
Score = 81.6 bits (200), Expect = 3e-16
Identities = 33/65 (50%), Positives = 46/65 (70%)
Frame = -3
Query: 47 TIAKLPVYYKQRDLLFYPSWAYAIPSWILKIPVSLMEVSLWVFLTYYVIGFDPNVGRMFK 106
TI +LPV+YK RD LF+P+W Y +P+++L+IP+S+ E +WV +TYY GF P R FK
Sbjct: 410 TIQRLPVFYKHRDHLFHPAWTYTVPNFLLRIPISIFESLVWVAITYYTTGFAPEASRFFK 231
Query: 107 QFRCV 111
Q V
Sbjct: 230 QLLVV 216
>TC18647 similar to UP|Q9ASR9 (Q9ASR9) At2g01320/F10A8.20, partial (17%)
Length = 430
Score = 60.1 bits (144), Expect = 1e-09
Identities = 35/93 (37%), Positives = 54/93 (57%), Gaps = 3/93 (3%)
Frame = +1
Query: 291 LLKGVSGAFRPGVLTALMGVSGAGKTTLMDVLAGRKTGG---YIDGDIKVSGYPKKQETF 347
LLK VSG +PG L A+MG SG+GKTTL++VLAG+ ++ G ++ +G P + +
Sbjct: 106 LLKNVSGEAKPGRLLAIMGPSGSGKTTLLNVLAGQLAASPRLHLSGLLEFNGKPGSKNAY 285
Query: 348 ARISGYCEHNDIHSPHVTVHESLLYSAWLRLPS 380
Y D+ +TV E+L + L+LP+
Sbjct: 286 K--FAYVRQEDLFFSQLTVRETLSLATELQLPN 378
>BE122583
Length = 361
Score = 47.0 bits (110), Expect(2) = 3e-07
Identities = 19/34 (55%), Positives = 28/34 (81%)
Frame = +3
Query: 252 KKGMVLPFEPHSITFDDIVYSVDMPVEMKEQGVR 285
KKGMVLPF+P S+ F+++ Y ++MP EMK+QG +
Sbjct: 171 KKGMVLPFQPLSLAFENVNYYIEMPNEMKKQGFK 272
Score = 24.6 bits (52), Expect(2) = 3e-07
Identities = 11/17 (64%), Positives = 13/17 (75%)
Frame = +1
Query: 299 FRPGVLTALMGVSGAGK 315
F+ TAL+GVSGAGK
Sbjct: 310 FQASDFTALVGVSGAGK 360
>TC17113 weakly similar to UP|Q9LFU8 (Q9LFU8) Proline-rich protein, partial
(25%)
Length = 1077
Score = 47.0 bits (110), Expect = 9e-06
Identities = 39/113 (34%), Positives = 47/113 (41%), Gaps = 6/113 (5%)
Frame = +1
Query: 446 PPPFPAHLLHTSLSHYPSGFSPYFTLLDPFFFPSYPFTPSPLYSSFVFPPPFPMAVTRSP 505
PP P L P SP L DPF PS P PL+ + F PP P + +P
Sbjct: 463 PPLIPNPFQPPPLIPNPFQPSPPPLLPDPFQPPS----PPPLFPN-PFQPPSPPPLIPNP 627
Query: 506 LPSPAARPPTAI--FLPPITSSSPFPLFPHSHKLLPLF----PSPSPAVFFSP 552
P + PP F PP + P PLFP ++P P P P FF P
Sbjct: 628 FQPPPSPPPFIPNPFQPPPSKQPPSPLFPFPPIVIPGLTPPPPPPPPPPFFPP 786
Score = 44.7 bits (104), Expect = 4e-05
Identities = 34/91 (37%), Positives = 40/91 (43%), Gaps = 4/91 (4%)
Frame = +1
Query: 473 DPFFFPSYPFTPSPLYSSFVFPPPFPMAVTRSPL-PSPAARPPTAIFLPPITSSSPFPLF 531
D FFFP PF P PL + PP P PL P+P PP +P SP PL
Sbjct: 370 DFFFFPPIPFFPPPLIPNPFQPPLVPNPFQPPPLIPNPFQPPP---LIPNPFQPSPPPLL 540
Query: 532 PHSHK---LLPLFPSPSPAVFFSPSCAVPLV 559
P + PLFP+P F P PL+
Sbjct: 541 PDPFQPPSPPPLFPNP-----FQPPSPPPLI 618
Score = 31.2 bits (69), Expect = 0.50
Identities = 25/72 (34%), Positives = 28/72 (38%), Gaps = 1/72 (1%)
Frame = +1
Query: 444 PLPPPF-PAHLLHTSLSHYPSGFSPYFTLLDPFFFPSYPFTPSPLYSSFVFPPPFPMAVT 502
P PPPF P PS P+ ++ P P P P P F PPFP
Sbjct: 640 PSPPPFIPNPFQPPPSKQPPSPLFPFPPIVIPGLTPPPPPPPPP-----PFFPPFP---- 792
Query: 503 RSPLPSPAARPP 514
PL P RPP
Sbjct: 793 --PLFPPPGRPP 822
>TC14279 similar to UP|Q09085 (Q09085) Hydroxyproline-rich glycoprotein
(HRGP) (Fragment), partial (57%)
Length = 941
Score = 46.2 bits (108), Expect = 2e-05
Identities = 39/111 (35%), Positives = 45/111 (40%), Gaps = 4/111 (3%)
Frame = +1
Query: 440 PSFAPLPPPFPAHLLHTSLSHYPSGFSPYFTLLDPFFFPSYPFTPSPLYSSFVFPP---- 495
P +P PPP + SH P PY+ P P P P P Y PP
Sbjct: 100 PPPSPSPPPPYYYQSPPPPSHSPP--PPYYYKSPP---PPSPSPPPPYYYKSPPPPSPSP 264
Query: 496 PFPMAVTRSPLPSPAARPPTAIFLPPITSSSPFPLFPHSHKLLPLFPSPSP 546
P P P PSP PP PP +SSP P + H + P PSPSP
Sbjct: 265 PPPYYYKSPPPPSPIPHPPYYYKSPPPPTSSPPPPY---HYVSPPPPSPSP 408
Score = 43.9 bits (102), Expect = 7e-05
Identities = 38/121 (31%), Positives = 43/121 (35%), Gaps = 17/121 (14%)
Frame = +1
Query: 444 PLPPPFPAHLLHTSLSHYPSGFSPYFTLLDPFFFPSYPFTPSPLYSSFVFPP----PFPM 499
P P P P + + PS PY+ P P P P Y PP P P
Sbjct: 202 PSPSPPPPYYYKSPPPPSPSPPPPYYYKSPP---PPSPIPHPPYYYKSPPPPTSSPPPPY 372
Query: 500 AVTRSPLPSPAARPPTAIFLPPITSSSPFPLF-------------PHSHKLLPLFPSPSP 546
P PSP+ PP PP S SP P + P H P PSPSP
Sbjct: 373 HYVSPPPPSPSPPPPYHYASPPPPSPSPAPTYIYKSPPPPVKLPPPPYHYTSPPPPSPSP 552
Query: 547 A 547
A
Sbjct: 553 A 555
Score = 43.5 bits (101), Expect = 1e-04
Identities = 42/126 (33%), Positives = 54/126 (42%), Gaps = 10/126 (7%)
Frame = +1
Query: 443 APLPP---PFPAHLLHTSLSHYPSGFSPYFTLLDPFFFPSYPFTPSPLYSSFVFPP---P 496
+P PP P P + + PS PY+ P P P P P Y PP P
Sbjct: 142 SPPPPSHSPPPPYYYKSPPPPSPSPPPPYYYKSPP---PPSPSPPPPYYYKSPPPPSPIP 312
Query: 497 FPMAVTRSPLPSPAARPPTAIFL-PPITSSSPFPLFPHSHKLLPLFPSPSPA---VFFSP 552
P +SP P ++ PP ++ PP S SP P + H P PSPSPA ++ SP
Sbjct: 313 HPPYYYKSPPPPTSSPPPPYHYVSPPPPSPSPPPPY---HYASPPPPSPSPAPTYIYKSP 483
Query: 553 SCAVPL 558
V L
Sbjct: 484 PPPVKL 501
Score = 42.0 bits (97), Expect = 3e-04
Identities = 29/69 (42%), Positives = 33/69 (47%)
Frame = +1
Query: 478 PSYPFTPSPLYSSFVFPPPFPMAVTRSPLPSPAARPPTAIFLPPITSSSPFPLFPHSHKL 537
P P P P Y + PPP P P PSP+ PP PP S SP P P+ +K
Sbjct: 25 PPPPPVPKPYY--YKSPPPPPYYYKSPPPPSPSPPPPYYYQSPPPPSHSPPP--PYYYKS 192
Query: 538 LPLFPSPSP 546
P PSPSP
Sbjct: 193 PPP-PSPSP 216
Score = 39.7 bits (91), Expect = 0.001
Identities = 35/121 (28%), Positives = 46/121 (37%), Gaps = 12/121 (9%)
Frame = +1
Query: 444 PLPPPFPAHLLHTSLS----HYPSGFSPYFTLLDPFFFPSYP---FTPSPLYSSFVFPPP 496
P PPP P + S +Y S P + P+++ S P +P P Y PPP
Sbjct: 25 PPPPPVPKPYYYKSPPPPPYYYKSPPPPSPSPPPPYYYQSPPPPSHSPPPPYYYKSPPPP 204
Query: 497 FPMA-----VTRSPLPSPAARPPTAIFLPPITSSSPFPLFPHSHKLLPLFPSPSPAVFFS 551
P P PSP+ PP PP S P P + + P P P + S
Sbjct: 205 SPSPPPPYYYKSPPPPSPSPPPPYYYKSPPPPSPIPHPPYYYKSPPPPTSSPPPPYHYVS 384
Query: 552 P 552
P
Sbjct: 385 P 387
Score = 33.9 bits (76), Expect = 0.077
Identities = 35/120 (29%), Positives = 53/120 (44%), Gaps = 10/120 (8%)
Frame = +1
Query: 440 PSFAPLPPPFPAHLLHTSLSHYPSGFSPYFTLLDPFFFPSYPF-TPSPLYSSFVFPPPFP 498
P+ +P PPP+ HY S P + P+ + S P +PSP + PP P
Sbjct: 346 PTSSP-PPPY----------HYVSPPPPSPSPPPPYHYASPPPPSPSPAPTYIYKSPPPP 492
Query: 499 MAV-------TRSPLPSPAARPPTAIFL--PPITSSSPFPLFPHSHKLLPLFPSPSPAVF 549
+ + T P PSP+ PT I+ PP T S P P++ ++ SP P ++
Sbjct: 493 VKLPPPPYHYTSPPPPSPSP-APTYIYKSPPPPTKSPPPPVY--------IYASPPPPIY 645
>AW164126
Length = 387
Score = 42.7 bits (99), Expect = 2e-04
Identities = 32/107 (29%), Positives = 45/107 (41%), Gaps = 12/107 (11%)
Frame = +2
Query: 456 TSLSHYPSGF-----SPYFTLLDPFFFPSYPF---TPSPLYSSFVFPP----PFPMAVTR 503
+S+S PS +P FT+ P+++ S P +P P Y PP P P
Sbjct: 56 SSISSTPSTLLLQVPTPTFTISKPYYYKSPPPPSPSPKPYYYKSPPPPSPSPPPPYYYKS 235
Query: 504 SPLPSPAARPPTAIFLPPITSSSPFPLFPHSHKLLPLFPSPSPAVFF 550
P PSP PP PP S P P + +S P+ P P ++
Sbjct: 236 PPPPSPVPHPPYYYKSPPPPSPIPHPPYYYSSPPPPVKSPPPPTPYY 376
>AV406930
Length = 433
Score = 42.7 bits (99), Expect = 2e-04
Identities = 54/165 (32%), Positives = 72/165 (42%), Gaps = 14/165 (8%)
Frame = +1
Query: 401 LHLLTLLNEHDRGGGRFIDHAMLSAELTPLMTSLTVFHFPSFAPLPPPFPAHLLHTSLSH 460
LHL L + H H+ +SA +L HF AP PPP P +
Sbjct: 4 LHLPPLHSSHP-------PHSSISA------LALAPPHF--LAPPPPPQPPN-------- 114
Query: 461 YPSGFSPYFTLLDPFFFPSYPFTP--SPLYSSFVFPPPFPMAV--TRSPLPSPAARP--- 513
P S ++ ++P TP +PL SF P P P ++ + S LPSP+ P
Sbjct: 115 -PPSISQNLSI------STHPITPFHTPLSHSFNLPLPKPASIPLSFSALPSPSPSPSSS 273
Query: 514 PTAIFLPPIT-----SSSPF--PLFPHSHKLLPLFPSPSPAVFFS 551
PT I PP + S SP PLF S ++LPL S SP + S
Sbjct: 274 PTLIPPPPSSSRRRGSCSPMNSPLFVSSKRILPLL-STSPTLLSS 405
>TC12500 similar to GB|AAH45354.1|28279536|BC045354 FNBP4 protein {Danio
rerio;} , partial (5%)
Length = 678
Score = 42.0 bits (97), Expect = 3e-04
Identities = 45/115 (39%), Positives = 52/115 (45%), Gaps = 10/115 (8%)
Frame = +1
Query: 429 PLMTSLTVFHFPSFAPLPPPFPAHLLHTSLSHYPSGFSPYFTLLD-PFFFPSYPFTPSPL 487
PL+ S T APLPPP HL TSL+ PS S T+ P P P +P P
Sbjct: 370 PLLMSAT-----DRAPLPPP-STHL--TSLTT-PSPPSSRTTITQTPLPSPPQPPSPPPK 522
Query: 488 YSSFVF-PPPFPM-AVTRSPLP-------SPAARPPTAIFLPPITSSSPFPLFPH 533
Y PPP PM ++ R+PLP S A PPT PP P P PH
Sbjct: 523 YGVATIPPPPLPMSSIDRTPLPLLSTPLTSLDAPPPTPTLSPP---PPPPPTPPH 678
Score = 35.0 bits (79), Expect = 0.035
Identities = 35/110 (31%), Positives = 45/110 (40%), Gaps = 3/110 (2%)
Frame = +1
Query: 440 PSFAPLPPPFPAHLLHTSLSHYPSGFSPYFTLLDPFFFPSYPFTPSPLYSSFVFPPPFPM 499
PS P PP PA ++ S SG + + L P P P P Y PP P+
Sbjct: 214 PSLPPTTPP-PASVIAGSA--IMSGEASFPCSLPS---PPQPPPPPPKYGVTTIPPSPPL 375
Query: 500 ---AVTRSPLPSPAARPPTAIFLPPITSSSPFPLFPHSHKLLPLFPSPSP 546
A R+PLP P+ L +T+ SP P S + P PSP
Sbjct: 376 LMSATDRAPLPPPSTH------LTSLTTPSP----PSSRTTITQTPLPSP 495
Score = 29.3 bits (64), Expect = 1.9
Identities = 21/65 (32%), Positives = 26/65 (39%), Gaps = 12/65 (18%)
Frame = +1
Query: 494 PPPFPMA-VTRSPLPSPAARPPTAIFLPPITSSSPFPLFPHSHKL-----------LPLF 541
PP PM+ R+PLP P+ P + LPP P S + LP
Sbjct: 4 PPSTPMSSADRAPLPPPSTPPKSLKALPPPPPPPSAPFIAGSAIISKVSKSVGDVSLPCS 183
Query: 542 PSPSP 546
PSP P
Sbjct: 184 PSPPP 198
Score = 27.3 bits (59), Expect = 7.2
Identities = 16/52 (30%), Positives = 22/52 (41%), Gaps = 1/52 (1%)
Frame = +1
Query: 494 PPPFPMAVTRSPLPSPAARP-PTAIFLPPITSSSPFPLFPHSHKLLPLFPSP 544
PP +T++PLPSP P P + P P+ LPL +P
Sbjct: 448 PPSSRTTITQTPLPSPPQPPSPPPKYGVATIPPPPLPMSSIDRTPLPLLSTP 603
>TC7889 similar to UP|Q40336 (Q40336) Proline-rich cell wall protein,
partial (66%)
Length = 1158
Score = 42.0 bits (97), Expect = 3e-04
Identities = 31/116 (26%), Positives = 45/116 (38%), Gaps = 2/116 (1%)
Frame = +1
Query: 440 PSFAPLPPPFPAHLLHTSLSHYPSGFSPYFTLLDPFFFPSYPFTPSP--LYSSFVFPPPF 497
P + P PP P +P + P +P P+ P P + V+PPP+
Sbjct: 169 PPYVPKPPVVPV--------------TPPYVPKPPIVYPP-PYVPKPPIVKPPIVYPPPY 303
Query: 498 PMAVTRSPLPSPAARPPTAIFLPPITSSSPFPLFPHSHKLLPLFPSPSPAVFFSPS 553
T + P PP + LPP+ S P P + P+P+P V PS
Sbjct: 304 --VPTPPIVKPPIVYPPPYVPLPPVVPSPPIVTPPTPTPPIVTPPTPTPPVVTPPS 465
Score = 32.7 bits (73), Expect = 0.17
Identities = 32/121 (26%), Positives = 42/121 (34%), Gaps = 1/121 (0%)
Frame = +1
Query: 440 PSFAPLPPPFPAHLLHTSLSHYPSGFSPYFTLLDPFFFPSYPFTPSP-LYSSFVFPPPFP 498
P + P PP P P P + P P P+ P P + S P P
Sbjct: 10 PPYVPKPPHVPK----------PPIVKPPYVPKPPVVKPP-PYVPKPPVVSPPYVPKPPV 156
Query: 499 MAVTRSPLPSPAARPPTAIFLPPITSSSPFPLFPHSHKLLPLFPSPSPAVFFSPSCAVPL 558
+ VT +P P P T ++P P P P + P P P V P P+
Sbjct: 157 VPVTPPYVPKPPVVPVTPPYVPKPPIVYPPPYVPKPPIVKPPIVYPPPYVPTPPIVKPPI 336
Query: 559 V 559
V
Sbjct: 337 V 339
Score = 30.4 bits (67), Expect = 0.85
Identities = 28/100 (28%), Positives = 37/100 (37%)
Frame = +1
Query: 428 TPLMTSLTVFHFPSFAPLPPPFPAHLLHTSLSHYPSGFSPYFTLLDPFFFPSYPFTPSPL 487
TP + + + P + PLPP P+ + T P+ P T P TP+P
Sbjct: 310 TPPIVKPPIVYPPPYVPLPPVVPSPPIVTP----PTPTPPIVT----------PPTPTPP 447
Query: 488 YSSFVFPPPFPMAVTRSPLPSPAARPPTAIFLPPITSSSP 527
V PP P T P P P P PP + P
Sbjct: 448 ----VVTPPSPPTETPCPPPPPVVPSP-----PPAQPTCP 540
Score = 28.9 bits (63), Expect = 2.5
Identities = 31/129 (24%), Positives = 45/129 (34%), Gaps = 9/129 (6%)
Frame = +1
Query: 440 PSFAPLPPPFPAHLLHTSLSHYPSGFSPYFTLLDPFFFPSYPFTPSPLYSSFVFPPPFPM 499
P + P PP P +P + P + P+ P P V+PPP+
Sbjct: 130 PPYVPKPPVVPV--------------TPPYVPKPPVVPVTPPYVPKP---PIVYPPPY-- 252
Query: 500 AVTRSPLPSPAARPPTAIFLPPITSSSPF--------PLFPHSHKLLPLFP-SPSPAVFF 550
+PP I PPI P+ P + +PL P PSP +
Sbjct: 253 ----------VPKPP--IVKPPIVYPPPYVPTPPIVKPPIVYPPPYVPLPPVVPSPPIVT 396
Query: 551 SPSCAVPLV 559
P+ P+V
Sbjct: 397 PPTPTPPIV 423
>BP033767
Length = 541
Score = 41.2 bits (95), Expect = 5e-04
Identities = 38/121 (31%), Positives = 48/121 (39%), Gaps = 5/121 (4%)
Frame = +1
Query: 438 HFPSFAPLPPPFPAHLLH-----TSLSHYPSGFSPYFTLLDPFFFPSYPFTPSPLYSSFV 492
H F PL P H H T +H+P P + P S P T L+ +
Sbjct: 160 HKAPFHPLNPQTHHHHHHLPHLQTQNTHFPPPPQPPPPKMPPHL--SSPHTH--LHHHLL 327
Query: 493 FPPPFPMAVTRSPLPSPAARPPTAIFLPPITSSSPFPLFPHSHKLLPLFPSPSPAVFFSP 552
PPP P PSP PP++ PP +P P S LP P+PSP+ P
Sbjct: 328 LPPPSP--------PSPPTSPPSSSLKPP----NPNPPTSSSASPLPPSPAPSPSSLSPP 471
Query: 553 S 553
S
Sbjct: 472 S 474
>TC7888 similar to UP|Q40336 (Q40336) Proline-rich cell wall protein,
partial (60%)
Length = 913
Score = 40.4 bits (93), Expect = 8e-04
Identities = 32/120 (26%), Positives = 46/120 (37%), Gaps = 6/120 (5%)
Frame = +3
Query: 440 PSFAPLPPPFPAHLLHTSLSHYPSGFSPYFT----LLDPFFFPSYPFTPSP--LYSSFVF 493
P + P PP P + PY + P +P P+ P P + V+
Sbjct: 561 PPYVPKPPVVPVTPPYVPKPPVVPVTPPYVPKPPIVKPPIVYPP-PYVPKPPIVKPPIVY 737
Query: 494 PPPFPMAVTRSPLPSPAARPPTAIFLPPITSSSPFPLFPHSHKLLPLFPSPSPAVFFSPS 553
PPP+ T + P PP + LPP+ S P P + P+P+P V PS
Sbjct: 738 PPPY--VPTPPIVKPPIVYPPPYVPLPPVVPSPPIVTPPTPTPPIVTPPTPTPPVVTPPS 911
Score = 32.7 bits (73), Expect = 0.17
Identities = 24/91 (26%), Positives = 33/91 (35%)
Frame = +3
Query: 467 PYFTLLDPFFFPSYPFTPSPLYSSFVFPPPFPMAVTRSPLPSPAARPPTAIFLPPITSSS 526
PY P P +P P++ PP P + + P+ P PP I PP+
Sbjct: 126 PYCPYPAPSKPPKHPIVKPPVHK----PPQHP-PIVKPPVHKPPQHPP--IVKPPVHKPP 284
Query: 527 PFPLFPHSHKLLPLFPSPSPAVFFSPSCAVP 557
+P P PSP P + P P
Sbjct: 285 KYPPSHGPEPCPPPKPSPKPPHYPKPPVVKP 377
Score = 30.8 bits (68), Expect = 0.65
Identities = 32/122 (26%), Positives = 46/122 (37%), Gaps = 9/122 (7%)
Frame = +3
Query: 440 PSFAPLPPPF-PAHLLHTSLSHYPSGFSPYFTLLDPFFFPSYPFTPSPLYSSFVFPP--P 496
P P PP P H+ + P P ++ P + P P P P V PP P
Sbjct: 342 PPHYPKPPVVKPPHVPKPPVVKPPHVPKP--PIVKPPYVPKPPHVPKP---PIVKPPYVP 506
Query: 497 FPMAVTRSP-LPSPAARPPTAIFLPPITSSSP-----FPLFPHSHKLLPLFPSPSPAVFF 550
P V P +P P P + PP+ +P P+ P + +P P P + +
Sbjct: 507 KPPVVKPPPYVPKPPVVSPPYVPKPPVVPVTPPYVPKPPVVPVTPPYVPKPPIVKPPIVY 686
Query: 551 SP 552
P
Sbjct: 687 PP 692
Score = 30.0 bits (66), Expect = 1.1
Identities = 34/126 (26%), Positives = 44/126 (33%), Gaps = 6/126 (4%)
Frame = +3
Query: 440 PSFAPLPPPFPAHLLHTSLSHYPSGFSPYFTLLDPFFFPSYPFTPSP-LYSSFVFPPPFP 498
P + P PP P P P + P P P+ P P + S P P
Sbjct: 441 PPYVPKPPHVPK----------PPIVKPPYVPKPPVVKPP-PYVPKPPVVSPPYVPKPPV 587
Query: 499 MAVTRSPLPSPAARPPTAIFL--PPITSSS---PFPLFPHSHKLLPLFPSPSPAVFFSPS 553
+ VT +P P P T ++ PPI P P P + P P P V P
Sbjct: 588 VPVTPPYVPKPPVVPVTPPYVPKPPIVKPPIVYPPPYVPKPPIVKPPIVYPPPYVPTPPI 767
Query: 554 CAVPLV 559
P+V
Sbjct: 768 VKPPIV 785
>TC19569 similar to UP|AAR11301 (AAR11301) Lectin-like receptor kinase 1;1,
partial (16%)
Length = 486
Score = 40.4 bits (93), Expect = 8e-04
Identities = 40/115 (34%), Positives = 50/115 (42%), Gaps = 6/115 (5%)
Frame = +1
Query: 453 LLHTSLSHYPSGFSPYFTLLDPFFFPSYPFTPSPLYSSFVFPPPFPMAVTRSP--LPSPA 510
LLHTS S S Y P P++ + SP +S PPP P VT +P PS +
Sbjct: 34 LLHTS----SSSSSSYSPPPPPSQQPTHSPSTSP--TSTTTPPPSPTKVTENPPTAPSTS 195
Query: 511 ARPPTAIF----LPPITSSSPFPLFPHSHKLLPLFPSPSPAVFFSPSCAVPLVLA 561
+ PT+ PP S+S HS P PSPSP PS P +A
Sbjct: 196 TKSPTSSASEEPSPPNPSTSTTQQPKHSPTSPPASPSPSP-----PSTKPPTAMA 345
Score = 36.6 bits (83), Expect = 0.012
Identities = 38/130 (29%), Positives = 49/130 (37%), Gaps = 10/130 (7%)
Frame = +1
Query: 438 HFPSFAPL----PPPFPAHLLHTSLSHYPSGFSPYFTLLDPFFFPSYPFTPSPLYSSFVF 493
H PS +P PPP P T ++ P T P P+P S+
Sbjct: 103 HSPSTSPTSTTTPPPSP-----TKVTENPPTAPSTSTKSPTSSASEEPSPPNPSTSTTQQ 267
Query: 494 PPPFPMAVTRSPLPSP-AARPPTAIFLP-----PITSSSPFPLFPHSHKLLPLFPSPSPA 547
P P + SP PSP + +PPTA+ P T S P P S P + S
Sbjct: 268 PKHSPTSPPASPSPSPPSTKPPTAMASPSTSPLSATKSHPTPPAAPSASSTPPPTTTSLK 447
Query: 548 VFFSPSCAVP 557
SPS + P
Sbjct: 448 TMSSPSSSTP 477
>BP034485
Length = 485
Score = 40.0 bits (92), Expect = 0.001
Identities = 43/117 (36%), Positives = 47/117 (39%), Gaps = 5/117 (4%)
Frame = -2
Query: 442 FAPLPPPFPAHL--LHTSLSHYPSGFSPYFTLLDPFFFPSYPFT-PSPLYSSFVFPPPFP 498
F P PPP P L L S S F P FFPS+P P PL F+F F
Sbjct: 349 FPPPPPPLPPLLP*LGMSKSWKSKSILSNFPPPPPPFFPSFPLLFPPPL---FLFLFLFL 179
Query: 499 MAVTRSPLPSPAARPPTAIFL--PPITSSSPFPLFPHSHKLLPLFPSPSPAVFFSPS 553
AV SP P P PP +F P + P P L L SP P F PS
Sbjct: 178 FAV--SPPPPP---PPFLLFFPRPKNLTRKPPPFLSLVFFFLSLLLSPPPPPFCFPS 23
>BP052318
Length = 460
Score = 39.7 bits (91), Expect = 0.001
Identities = 30/85 (35%), Positives = 40/85 (46%), Gaps = 2/85 (2%)
Frame = +2
Query: 479 SYPFTPSPL--YSSFVFPPPFPMAVTRSPLPSPAARPPTAIFLPPITSSSPFPLFPHSHK 536
S+ TP+P ++ PPP P A T P P P+ RPP + PP P PL H +
Sbjct: 131 SWTVTPTPPRHITATSSPPPPPSAPTSPPSPYPSLRPPIS---PP-----PSPLPRHLRR 286
Query: 537 LLPLFPSPSPAVFFSPSCAVPLVLA 561
P PS + SP + PL L+
Sbjct: 287 WTPWPSPPSLPSWPSPFLSCPLALS 361
>BP044109
Length = 497
Score = 39.7 bits (91), Expect = 0.001
Identities = 32/106 (30%), Positives = 42/106 (39%), Gaps = 11/106 (10%)
Frame = +2
Query: 450 PAHLLHTSLSHYPSGFSPYFTLLDPFFFPSYPFTPSPLYSSFVFPPPFPMAVTRS----- 504
P H TSLSH S +L FFPS P P P + P A ++S
Sbjct: 35 PPHTATTSLSHSSST-----SLSSSHFFPSPPPPPPPPNKNKTTPSHASNATSKSTPRTQ 199
Query: 505 ------PLPSPAARPPTAIFLPPITSSSPFPLFPHSHKLLPLFPSP 544
P P+ + +PP + F ++S P SH P PSP
Sbjct: 200 TPTTPPPSPTSSPKPPPSTFATKPSTSPPETPSSSSHGPAPTLPSP 337
>TC8675 weakly similar to UP|Q9LFH0 (Q9LFH0) PDR9 ABC transporter, partial
(13%)
Length = 727
Score = 39.7 bits (91), Expect = 0.001
Identities = 17/56 (30%), Positives = 33/56 (58%)
Frame = +3
Query: 53 VYYKQRDLLFYPSWAYAIPSWILKIPVSLMEVSLWVFLTYYVIGFDPNVGRMFKQF 108
V Y++R Y WAY+ ++++P S ++ +L+V +TY +IG + ++F F
Sbjct: 18 VLYRERFAGMYSPWAYSFAQVLIEVPYSFIQAALYVIITYPMIGHYWSAYKIFWSF 185
>AV425247
Length = 417
Score = 39.3 bits (90), Expect = 0.002
Identities = 30/90 (33%), Positives = 40/90 (44%), Gaps = 7/90 (7%)
Frame = +3
Query: 470 TLLDPFFFPSY-PFTPSPLYSSFVFPPP-FPMAVTRSPLPSPAARPPTAIFLPPITSSSP 527
TL+DP ++ P +P+ ++ PPP ++ + SP PSP PPT P P
Sbjct: 126 TLIDPPVHHAHSPPSPTTYSHPYLCPPPQSTLSPSPSPSPSPLLLPPTLPLPSPFAHLLP 305
Query: 528 FPLFPHSHKLLPL-----FPSPSPAVFFSP 552
SH LPL F SP FSP
Sbjct: 306 LL*HSFSHPPLPLLTPTSFAQSSPRPIFSP 395
Score = 33.1 bits (74), Expect = 0.13
Identities = 26/80 (32%), Positives = 31/80 (38%), Gaps = 12/80 (15%)
Frame = +2
Query: 472 LDPFFFPSYPFTPSPLYSSFVFPPPFPMAVTRSPLPS--------PAARPPTAIFLPPIT 523
L PF P P S PFP +T SP P+ P++ P A+ PP
Sbjct: 158 LSPFSHHLLPPLSLPPSSIHTISFPFPFPITPSPSPNTPPPLTLCPSSPSPLALIFPPTA 337
Query: 524 SSSP----FPLFPHSHKLLP 539
S S P P SH L P
Sbjct: 338 SPSHPHLICPKQPTSHLLSP 397
Score = 29.3 bits (64), Expect = 1.9
Identities = 23/76 (30%), Positives = 31/76 (40%)
Frame = +1
Query: 427 LTPLMTSLTVFHFPSFAPLPPPFPAHLLHTSLSHYPSGFSPYFTLLDPFFFPSYPFTPSP 486
L PL F FP +P P P P L +S +H+P+ P+ P P PS
Sbjct: 223 LLPLPLPHHPFSFPQHSPSPHPLPIFSLSSS-THFPTHRFPF---SPPPHLPKAAHVPSS 390
Query: 487 LYSSFVFPPPFPMAVT 502
L P P+ +T
Sbjct: 391 L--------PTPLEIT 414
>TC15802 similar to UP|Q9XIV1 (Q9XIV1) mRNA expressed in cucumber hypocotyls
(Arabinogalactan protein), partial (26%)
Length = 795
Score = 38.5 bits (88), Expect = 0.003
Identities = 36/131 (27%), Positives = 48/131 (36%), Gaps = 1/131 (0%)
Frame = +2
Query: 428 TPLMTSLTVFHFPSFAPLPPPFPAHLLHTSLSHYPSGFSPYFTLLDPFFFPSYPFT-PSP 486
+P S T AP+ P PA + + +P T P P T P P
Sbjct: 134 SPATPSATTPVSTPVAPVASPKPAATTPAASPSTATAPAPATTT------PVTPVTSPPP 295
Query: 487 LYSSFVFPPPFPMAVTRSPLPSPAARPPTAIFLPPITSSSPFPLFPHSHKLLPLFPSPSP 546
PPP + V+ P SP A PTA+ + ++ P S P+PSP
Sbjct: 296 AAVPVASPPPAAVPVSSPPAKSPPAPAPTAVPVAAPVTTPEVPAPAPSKTKKDAAPAPSP 475
Query: 547 AVFFSPSCAVP 557
V SP P
Sbjct: 476 VVPDSPPAGAP 508
>AU252214
Length = 350
Score = 38.5 bits (88), Expect = 0.003
Identities = 25/62 (40%), Positives = 31/62 (49%)
Frame = -2
Query: 483 TPSPLYSSFVFPPPFPMAVTRSPLPSPAARPPTAIFLPPITSSSPFPLFPHSHKLLPLFP 542
+P P SS PP F VT + P PP+ L + SSSP PL P+ H L P +
Sbjct: 262 SPLPPPSSRPLPPSF*RRVTLNLPPLSRTDPPS---L*SLCSSSPIPLMPYEHGLPPPWS 92
Query: 543 SP 544
SP
Sbjct: 91 SP 86
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.331 0.145 0.488
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 12,848,248
Number of Sequences: 28460
Number of extensions: 279015
Number of successful extensions: 4514
Number of sequences better than 10.0: 440
Number of HSP's better than 10.0 without gapping: 3603
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 4133
length of query: 596
length of database: 4,897,600
effective HSP length: 95
effective length of query: 501
effective length of database: 2,193,900
effective search space: 1099143900
effective search space used: 1099143900
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.9 bits)
S2: 58 (26.9 bits)
Medicago: description of AC126018.1


