

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC126016.2 - phase: 0
(327 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
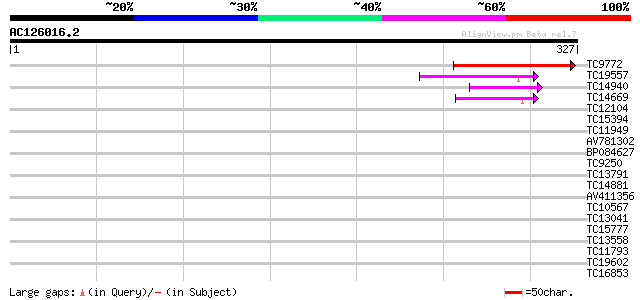
Score E
Sequences producing significant alignments: (bits) Value
TC9772 weakly similar to GB|AAG50239.1|12276054|AF304126 RING an... 143 3e-35
TC19557 similar to PIR|E86326|E86326 protein F18O14.3 [imported]... 45 2e-05
TC14940 42 2e-04
TC14669 similar to PIR|E86326|E86326 protein F18O14.3 [imported]... 40 5e-04
TC12104 37 0.005
TC15394 similar to UP|Q9FFQ8 (Q9FFQ8) Emb|CAB62360.1, partial (16%) 36 0.008
TC11949 36 0.010
AV781302 35 0.013
BP084627 35 0.017
TC9250 UP|Q9FLP6 (Q9FLP6) Ubiquitin-like protein SMT3-like (AT5g... 35 0.017
TC13791 weakly similar to UP|RO52_HUMAN (P19474) 52 kDa Ro prote... 35 0.022
TC14881 35 0.022
AV411356 34 0.038
TC10567 similar to UP|AAR99376 (AAR99376) Ring domain containing... 34 0.038
TC13041 similar to UP|Q9M8S4 (Q9M8S4) Zinc finger protein 1 (zfn... 33 0.050
TC15777 similar to UP|AAR99376 (AAR99376) Ring domain containing... 32 0.11
TC13558 similar to GB|AAD33770.1|4928919|AF138744 zinc finger pr... 32 0.15
TC11793 similar to UP|AAR99376 (AAR99376) Ring domain containing... 32 0.19
TC19602 similar to UP|Q9LG20 (Q9LG20) F14J16.17, partial (13%) 32 0.19
TC16853 similar to UP|Q93ZP3 (Q93ZP3) At1g32359/F27G20.10, parti... 32 0.19
>TC9772 weakly similar to GB|AAG50239.1|12276054|AF304126 RING and zinc
finger protein {Caenorhabditis elegans;} , partial (13%)
Length = 558
Score = 143 bits (361), Expect = 3e-35
Identities = 61/70 (87%), Positives = 66/70 (94%)
Frame = +3
Query: 257 EDDDEDALPFACFICRNPFVDPVSTKCKHYFCEHCALKHHAKNKKCFVCNQPTLGIFNVA 316
+++D+D+LPFACFICR PFVDPV TKCKHYFCEHCALKHHAKNKKCFVCNQPTLG FNVA
Sbjct: 3 DEEDDDSLPFACFICRQPFVDPVVTKCKHYFCEHCALKHHAKNKKCFVCNQPTLGTFNVA 182
Query: 317 HEIRKKMAED 326
EIRKKMAED
Sbjct: 183 QEIRKKMAED 212
>TC19557 similar to PIR|E86326|E86326 protein F18O14.3 [imported] -
Arabidopsis thaliana {Arabidopsis thaliana;}, partial
(24%)
Length = 457
Score = 45.1 bits (105), Expect = 2e-05
Identities = 25/72 (34%), Positives = 34/72 (46%), Gaps = 3/72 (4%)
Frame = +2
Query: 237 RKMRLATGEDAEEEGASLNDEDDDEDALPFACFICRNPFVDPVSTKCKHYFCEHCA---L 293
R+M GE ++ + DA F C IC DPV T C H FC C L
Sbjct: 218 RQMASGYGESTRAAPSTSCSGNSPNDAGDFECNICFELAQDPVITLCGHLFCWPCLYRWL 397
Query: 294 KHHAKNKKCFVC 305
HH+++++C VC
Sbjct: 398 HHHSQSQECPVC 433
>TC14940
Length = 781
Score = 41.6 bits (96), Expect = 2e-04
Identities = 16/42 (38%), Positives = 23/42 (54%)
Frame = +2
Query: 266 FACFICRNPFVDPVSTKCKHYFCEHCALKHHAKNKKCFVCNQ 307
F+C IC P V+ +ST+C H FC+ C + KC C +
Sbjct: 143 FSCPICMGPLVEEMSTRCGHIFCKACIKAAISAQNKCPTCRK 268
>TC14669 similar to PIR|E86326|E86326 protein F18O14.3 [imported] -
Arabidopsis thaliana {Arabidopsis thaliana;}, partial
(77%)
Length = 1120
Score = 40.0 bits (92), Expect = 5e-04
Identities = 19/51 (37%), Positives = 29/51 (56%), Gaps = 3/51 (5%)
Frame = +2
Query: 258 DDDEDALPFACFICRNPFVDPVSTKCKHYFCEHCALK---HHAKNKKCFVC 305
+++ DA F C IC + DP+ T C H FC C K H+++++C VC
Sbjct: 98 NNNSDAGNFECNICFDLAQDPIITLCGHLFCWPCLYKWLHFHSQSRECPVC 250
>TC12104
Length = 771
Score = 37.0 bits (84), Expect = 0.005
Identities = 13/38 (34%), Positives = 19/38 (49%)
Frame = +1
Query: 268 CFICRNPFVDPVSTKCKHYFCEHCALKHHAKNKKCFVC 305
C IC+ P+ CKH FCE C + + + C +C
Sbjct: 421 CAICQEKMHSPILLCCKHMFCEECVSEWFERERTCPLC 534
>TC15394 similar to UP|Q9FFQ8 (Q9FFQ8) Emb|CAB62360.1, partial (16%)
Length = 1093
Score = 36.2 bits (82), Expect = 0.008
Identities = 60/275 (21%), Positives = 113/275 (40%), Gaps = 19/275 (6%)
Frame = +2
Query: 23 KPVNRKNMRKRTIENEDNENDSNNEESLMHVQKKNTK------ADNKLFFSTGSSKSSA- 75
K +++N+ E++D+E+ S EE QKK +K + +K T S+K +
Sbjct: 29 KDESQENVAAPHKESDDDESKSEEEEDKSKAQKKTSKKIVKEGSGSKAGERTKSAKKTPV 208
Query: 76 -SAKPNEESEKPSFHFESSKEIQVQHDSKATATLETETDFSRDARAIRERALKQATESLK 134
S+K +++ K S + K+ +HDS + + +++ S+ + E+ K
Sbjct: 209 KSSKNVDKTPKKS----TPKQTAAEHDSASASLSKSKQPASKKRKTENEKP------DTK 358
Query: 135 GKSTSSEDV-KLYKGINNYTDHKAGFRREQTIASEKAGGSHGPLRASAHIRVSARFDYQP 193
GK++S + K K + ++ + + S +A + + + D+
Sbjct: 359 GKASSKKQTDKSSKALVKDQGKSKSGKKAKAVPSREA-------MHAVVVDILKEVDFNT 517
Query: 194 DICKDYKETGYCGYGDSCKFMHDRGDYKSGW-----QMEKEWDEAEKARKMRLATGEDAE 248
D +G MH + + K M E DE GE+A+
Sbjct: 518 ATLSDILRPLGTHFG--VDLMHRKAEVKDIITDVINNMSDEEDE-----------GEEAD 658
Query: 249 EEGASLNDE--DDDEDALPFACF-ICRN--PFVDP 278
+G + D+ DDD++ +A F +CRN P V P
Sbjct: 659 GDGDADKDDNGDDDDE*FSWAMFDLCRN*RPLVIP 763
Score = 35.4 bits (80), Expect = 0.013
Identities = 30/135 (22%), Positives = 57/135 (42%), Gaps = 7/135 (5%)
Frame = +2
Query: 2 EDSEAKSTENQQTEQVCSFFRKPV------NRKNMRKRTIENEDNENDSNNEESLMHVQK 55
+D E+KS E + + K + ++ R ++ + ++ N +++
Sbjct: 74 DDDESKSEEEEDKSKAQKKTSKKIVKEGSGSKAGERTKSAKKTPVKSSKNVDKTPKKSTP 253
Query: 56 KNTKADN-KLFFSTGSSKSSASAKPNEESEKPSFHFESSKEIQVQHDSKATATLETETDF 114
K T A++ S SK AS K E+EKP ++S + Q SKA + ++
Sbjct: 254 KQTAAEHDSASASLSKSKQPASKKRKTENEKPDTKGKASSKKQTDKSSKALVKDQGKSKS 433
Query: 115 SRDARAIRERALKQA 129
+ A+A+ R A
Sbjct: 434 GKKAKAVPSREAMHA 478
>TC11949
Length = 780
Score = 35.8 bits (81), Expect = 0.010
Identities = 18/55 (32%), Positives = 23/55 (41%), Gaps = 11/55 (20%)
Frame = +3
Query: 264 LPFACFICRNPFVDPVSTKCKHYFCEHCA-----------LKHHAKNKKCFVCNQ 307
+ C IC + DPVS C H FC CA LK +KC +C +
Sbjct: 117 IDLTCSICMDTVFDPVSLTCGHIFCYSCACSAASVTIVDGLKAAXPKEKCPLCRE 281
>AV781302
Length = 533
Score = 35.4 bits (80), Expect = 0.013
Identities = 13/22 (59%), Positives = 14/22 (63%)
Frame = +3
Query: 196 CKDYKETGYCGYGDSCKFMHDR 217
C Y TGYCGYG C+F H R
Sbjct: 345 CIYYLRTGYCGYGSRCRFNHPR 410
Score = 28.9 bits (63), Expect = 1.2
Identities = 18/53 (33%), Positives = 25/53 (46%)
Frame = +3
Query: 161 REQTIASEKAGGSHGPLRASAHIRVSARFDYQPDICKDYKETGYCGYGDSCKF 213
R I + + GG + P RA QP +C+ Y TG C +G SCK+
Sbjct: 414 RGAVIGAARTGGEY-PERAG-----------QP-VCQYYMRTGSCKFGSSCKY 533
>BP084627
Length = 327
Score = 35.0 bits (79), Expect = 0.017
Identities = 16/39 (41%), Positives = 23/39 (58%), Gaps = 1/39 (2%)
Frame = -2
Query: 268 CFICRNPFVDPVSTKCKHYFCEHCALK-HHAKNKKCFVC 305
C IC++ + V TKC H FC C LK ++++KC C
Sbjct: 326 CSICQDRTKEVVITKCYHLFCNTCMLKVTGSRHRKCPQC 210
>TC9250 UP|Q9FLP6 (Q9FLP6) Ubiquitin-like protein SMT3-like
(AT5g55160/MCO15_11) (Small ubiquitin-like modifier 2),
partial (22%)
Length = 1363
Score = 35.0 bits (79), Expect = 0.017
Identities = 22/87 (25%), Positives = 42/87 (47%), Gaps = 2/87 (2%)
Frame = +1
Query: 243 TGEDAEEEGASLNDEDDDEDAL-PF-ACFICRNPFVDPVSTKCKHYFCEHCALKHHAKNK 300
T ++ + G E +D++ PF AC +C +DP+S + H +C+ C L
Sbjct: 358 TYDEKRKLGYGTQKERLGKDSIKPFDACCLCLKSLIDPMSCQKGHLYCKECIL------- 516
Query: 301 KCFVCNQPTLGIFNVAHEIRKKMAEDK 327
+C + + + VAH ++K +D+
Sbjct: 517 QCLLSQKKDIQRRLVAHAAQQKQEKDE 597
Score = 34.3 bits (77), Expect = 0.029
Identities = 15/46 (32%), Positives = 24/46 (51%), Gaps = 4/46 (8%)
Frame = +1
Query: 266 FACFICRNPFVDPVS----TKCKHYFCEHCALKHHAKNKKCFVCNQ 307
+ C C+ + +S + C H FC+ CA + A +K C VCN+
Sbjct: 946 YICPSCKVTLTNTMSLVAISSCGHVFCKKCADRFMASDKVCLVCNK 1083
>TC13791 weakly similar to UP|RO52_HUMAN (P19474) 52 kDa Ro protein (Sjogren
syndrome type A antigen) (SS-A) (Ro(SS-A)) (52 kDa
ribonucleoprotein autoantigen Ro/SS-A), partial (5%)
Length = 545
Score = 34.7 bits (78), Expect = 0.022
Identities = 18/55 (32%), Positives = 23/55 (41%), Gaps = 11/55 (20%)
Frame = +3
Query: 264 LPFACFICRNPFVDPVSTKCKHYFCEHCA-----------LKHHAKNKKCFVCNQ 307
+ C IC + DPVS C H FC CA LK +KC +C +
Sbjct: 48 IDLTCPICLDTVFDPVSLTCGHIFCYMCACSAASVSIVDGLKSAVTKQKCPLCRE 212
>TC14881
Length = 609
Score = 34.7 bits (78), Expect = 0.022
Identities = 19/59 (32%), Positives = 26/59 (43%), Gaps = 1/59 (1%)
Frame = +3
Query: 251 GASLNDEDDDEDALPFACFICRNPFVDPVSTKCKHY-FCEHCALKHHAKNKKCFVCNQP 308
G+S + DD D C IC D C+H C CA + ++ KC +C QP
Sbjct: 51 GSSTAADFDDNDPQK-ECVICMTEPKDTAVLPCRHMCMCSDCAKELRLQSNKCPICRQP 224
>AV411356
Length = 358
Score = 33.9 bits (76), Expect = 0.038
Identities = 18/43 (41%), Positives = 22/43 (50%), Gaps = 3/43 (6%)
Frame = +2
Query: 266 FACFICRNPFVDPVSTKCKHYFCEHCA---LKHHAKNKKCFVC 305
F C IC + +PV T C H FC C L H+ K+C VC
Sbjct: 38 FDCNICLDLAREPVVTCCGHLFCWTCVYRWLHLHSDAKECPVC 166
>TC10567 similar to UP|AAR99376 (AAR99376) Ring domain containing protein,
partial (27%)
Length = 423
Score = 33.9 bits (76), Expect = 0.038
Identities = 15/42 (35%), Positives = 19/42 (44%)
Frame = +2
Query: 253 SLNDEDDDEDALPFACFICRNPFVDPVSTKCKHYFCEHCALK 294
++ D D F C IC + DPV T C H +C C K
Sbjct: 41 AIEDHSDRNGLSGFDCSICLDRVQDPVVTLCGHLYCWPCIYK 166
>TC13041 similar to UP|Q9M8S4 (Q9M8S4) Zinc finger protein 1 (zfn1), partial
(9%)
Length = 535
Score = 33.5 bits (75), Expect = 0.050
Identities = 11/22 (50%), Positives = 14/22 (63%)
Frame = +2
Query: 196 CKDYKETGYCGYGDSCKFMHDR 217
C Y TG+CG+G C+F H R
Sbjct: 458 CSYYLRTGFCGFGSRCRFNHPR 523
>TC15777 similar to UP|AAR99376 (AAR99376) Ring domain containing protein,
partial (34%)
Length = 566
Score = 32.3 bits (72), Expect = 0.11
Identities = 20/63 (31%), Positives = 28/63 (43%)
Frame = +1
Query: 232 EAEKARKMRLATGEDAEEEGASLNDEDDDEDALPFACFICRNPFVDPVSTKCKHYFCEHC 291
E + + +M G+DA E+ D + F C IC + DPV T C H +C C
Sbjct: 202 EDKSSLEMWKCGGDDAIEDS-------DKNGSGGFDCNICLDCVQDPVVTLCGHLYCWPC 360
Query: 292 ALK 294
K
Sbjct: 361 IYK 369
>TC13558 similar to GB|AAD33770.1|4928919|AF138744 zinc finger protein 2
{Arabidopsis thaliana;}, partial (8%)
Length = 553
Score = 32.0 bits (71), Expect = 0.15
Identities = 12/24 (50%), Positives = 16/24 (66%)
Frame = +1
Query: 192 QPDICKDYKETGYCGYGDSCKFMH 215
+PD C Y TG CGYG +C++ H
Sbjct: 481 EPD-CLYYLRTGMCGYGSNCRYNH 549
>TC11793 similar to UP|AAR99376 (AAR99376) Ring domain containing protein,
partial (23%)
Length = 527
Score = 31.6 bits (70), Expect = 0.19
Identities = 15/38 (39%), Positives = 18/38 (46%)
Frame = +3
Query: 257 EDDDEDALPFACFICRNPFVDPVSTKCKHYFCEHCALK 294
E D + F C IC + DPV T C H +C C K
Sbjct: 318 ESDRTSSGGFDCNICLDCVQDPVVTLCGHLYCWPCIYK 431
>TC19602 similar to UP|Q9LG20 (Q9LG20) F14J16.17, partial (13%)
Length = 548
Score = 31.6 bits (70), Expect = 0.19
Identities = 13/40 (32%), Positives = 20/40 (49%)
Frame = +2
Query: 268 CFICRNPFVDPVSTKCKHYFCEHCALKHHAKNKKCFVCNQ 307
C +C V+TK + C+ C L + K +KC C+Q
Sbjct: 218 CELCGTQQPKDVTTKYSTWSCKFCTLDNSVKMEKCSACDQ 337
>TC16853 similar to UP|Q93ZP3 (Q93ZP3) At1g32359/F27G20.10, partial (20%)
Length = 722
Score = 31.6 bits (70), Expect = 0.19
Identities = 10/26 (38%), Positives = 17/26 (64%)
Frame = +1
Query: 190 DYQPDICKDYKETGYCGYGDSCKFMH 215
+++ IC ++ TGYC +G+ C F H
Sbjct: 52 NWKTRICNKWEMTGYCPFGNKCHFAH 129
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.311 0.127 0.368
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 5,566,147
Number of Sequences: 28460
Number of extensions: 81131
Number of successful extensions: 525
Number of sequences better than 10.0: 81
Number of HSP's better than 10.0 without gapping: 476
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 510
length of query: 327
length of database: 4,897,600
effective HSP length: 91
effective length of query: 236
effective length of database: 2,307,740
effective search space: 544626640
effective search space used: 544626640
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (21.9 bits)
S2: 55 (25.8 bits)
Medicago: description of AC126016.2


