

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
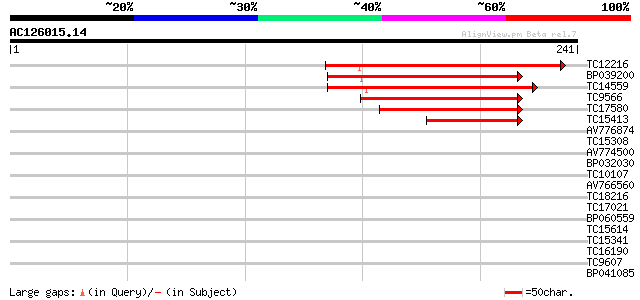
Query= AC126015.14 + phase: 0 /pseudo
(241 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
TC12216 similar to GB|AAP13420.1|30023774|BT006312 At3g45600 {Ar... 103 2e-23
BP039200 102 5e-23
TC14559 weakly similar to UP|Q9SUD4 (Q9SUD4) Senescence-associat... 100 2e-22
TC9566 weakly similar to UP|Q9FIQ5 (Q9FIQ5) Senescence-associate... 73 4e-14
TC17580 weakly similar to UP|Q9SUD4 (Q9SUD4) Senescence-associat... 64 2e-11
TC15413 similar to UP|Q9SUD4 (Q9SUD4) Senescence-associated prot... 54 3e-08
AV776874 28 1.4
TC15308 similar to UP|Q41725 (Q41725) TED3, partial (16%) 28 1.4
AV774500 27 2.4
BP032030 27 2.4
TC10107 similar to UP|Q9MB24 (Q9MB24) S1-3 protein (Fragment), p... 27 3.1
AV766560 27 3.1
TC18216 similar to GB|CAA28991.1|34041|HSKER65A keratin type II ... 27 3.1
TC17021 homologue to UP|Q9SSP4 (Q9SSP4) F3N23.2 protein (At1g728... 27 4.1
BP060559 27 4.1
TC15614 similar to PIR|T00553|T00553 disease resistance response... 27 4.1
TC15341 similar to UP|Q9M0H8 (Q9M0H8) Predicted proline-rich pro... 26 5.4
TC16190 weakly similar to UP|Q9LQ75 (Q9LQ75) T1N6.22 protein, pa... 26 5.4
TC9607 similar to UP|Q8S9J2 (Q8S9J2) At2g33840/T1B8.14, partial ... 26 5.4
BP041085 26 5.4
>TC12216 similar to GB|AAP13420.1|30023774|BT006312 At3g45600 {Arabidopsis
thaliana;}, partial (30%)
Length = 570
Score = 103 bits (258), Expect = 2e-23
Identities = 46/110 (41%), Positives = 69/110 (61%), Gaps = 8/110 (7%)
Frame = +2
Query: 135 NTGCCKPPTACNY--------NMEAVMMTQDSDCYKWSNEPTLLCYECDSCKAGVLEDIR 186
N+ + T C Y N+ + +M+ + DC KWSN+ LCY CDSCKAGVL ++
Sbjct: 2 NSARARAATDCGYIYQNETVWNLGSGLMSANPDCSKWSNDQGFLCYRCDSCKAGVLASLK 181
Query: 187 RNWHKLSVLTVTMLILLIGIYSIGCCAFRNARRAETDYPYGENRMTKVRP 236
++W K+SV+ + ++I+L+ +Y I A+RN +R + D PYGE RMTK P
Sbjct: 182 KSWRKVSVINIVVMIILVIVYIIAYAAYRNNKRMDNDEPYGEARMTKSHP 331
>BP039200
Length = 553
Score = 102 bits (255), Expect = 5e-23
Identities = 42/90 (46%), Positives = 59/90 (64%), Gaps = 7/90 (7%)
Frame = -1
Query: 136 TGCCKPPTACNYN-------MEAVMMTQDSDCYKWSNEPTLLCYECDSCKAGVLEDIRRN 188
+GCCKPPT C Y + + D DC +WSN+ T LCY CDSCKAG+L ++R+
Sbjct: 445 SGCCKPPTQCAYTFVNPTYWISPINTAADMDCLQWSNDQTQLCYGCDSCKAGLLANLRKE 266
Query: 189 WHKLSVLTVTMLILLIGIYSIGCCAFRNAR 218
W + V+ + L++LI +Y +GCCAFRNA+
Sbjct: 265 WKRAXVVLIITLVVLIVVYMVGCCAFRNAK 176
>TC14559 weakly similar to UP|Q9SUD4 (Q9SUD4) Senescence-associated
protein-like, partial (92%)
Length = 1125
Score = 100 bits (249), Expect = 2e-22
Identities = 42/97 (43%), Positives = 65/97 (66%), Gaps = 8/97 (8%)
Frame = +2
Query: 136 TGCCKPPTACNYNME--------AVMMTQDSDCYKWSNEPTLLCYECDSCKAGVLEDIRR 187
+GCCKP CN+ E A ++DC W+N+ T+LC+ C SCKAG+L++I+
Sbjct: 626 SGCCKPSDDCNFTYENPTTWTKPANATYTNTDCDAWNNDQTILCFNCQSCKAGLLQNIKT 805
Query: 188 NWHKLSVLTVTMLILLIGIYSIGCCAFRNARRAETDY 224
+W K++V+ + LI LI +YS+GCCAFRN+RR +++
Sbjct: 806 DWKKVAVVNIIFLIFLIIVYSVGCCAFRNSRRDNSNW 916
>TC9566 weakly similar to UP|Q9FIQ5 (Q9FIQ5) Senescence-associated protein
5-like protein, partial (28%)
Length = 675
Score = 73.2 bits (178), Expect = 4e-14
Identities = 28/69 (40%), Positives = 46/69 (66%)
Frame = +1
Query: 150 EAVMMTQDSDCYKWSNEPTLLCYECDSCKAGVLEDIRRNWHKLSVLTVTMLILLIGIYSI 209
E V D DC W+N+ LCY C++CKAG+L ++R+ W K +++ + +++LI +Y I
Sbjct: 25 EPVNPMADPDCNLWNNDQNQLCYNCNACKAGLLGNLRKEWRKANIILIVAVVVLICVYII 204
Query: 210 GCCAFRNAR 218
C AF+NA+
Sbjct: 205 ACSAFKNAQ 231
>TC17580 weakly similar to UP|Q9SUD4 (Q9SUD4) Senescence-associated
protein-like, partial (12%)
Length = 526
Score = 63.9 bits (154), Expect = 2e-11
Identities = 26/61 (42%), Positives = 37/61 (60%)
Frame = +3
Query: 158 SDCYKWSNEPTLLCYECDSCKAGVLEDIRRNWHKLSVLTVTMLILLIGIYSIGCCAFRNA 217
SDC W N+ LCY+C++CK G L +IR W L++ +L L+ IY +GC A +N
Sbjct: 27 SDCKTWDNKQDKLCYDCNACKGGFLPNIRNQWRHLTIFNGLVLALVTTIYVLGCYAIKNN 206
Query: 218 R 218
R
Sbjct: 207 R 209
>TC15413 similar to UP|Q9SUD4 (Q9SUD4) Senescence-associated protein-like,
partial (15%)
Length = 564
Score = 53.5 bits (127), Expect = 3e-08
Identities = 22/41 (53%), Positives = 33/41 (79%)
Frame = +3
Query: 178 KAGVLEDIRRNWHKLSVLTVTMLILLIGIYSIGCCAFRNAR 218
+AG+L++I+ +W K++V+ V L+ LI +YSIGCCAFRN R
Sbjct: 48 QAGLLQNIKTDWKKVAVVNVIFLVFLIIVYSIGCCAFRNNR 170
Score = 27.7 bits (60), Expect = 1.8
Identities = 11/30 (36%), Positives = 15/30 (49%), Gaps = 3/30 (10%)
Frame = +1
Query: 163 WSNEPTLLCYECDSCKAGVLEDIR---RNW 189
W N+P LLC+ C S + +R R W
Sbjct: 4 WDNDPMLLCFNCHSRRLDYCRTLRLIGRRW 93
>AV776874
Length = 588
Score = 28.1 bits (61), Expect = 1.4
Identities = 14/46 (30%), Positives = 26/46 (56%), Gaps = 9/46 (19%)
Frame = -2
Query: 31 WFHWCLFSCCMCTMVVLG--DYVIANSSTI-------RFDYIWFWC 67
+FH C++ C+ ++V D V+A SS++ R D++ F+C
Sbjct: 155 FFHCCVYILCVHDLIVDSCLDLVVATSSSLLYFNCV*RLDFVVFYC 18
>TC15308 similar to UP|Q41725 (Q41725) TED3, partial (16%)
Length = 1103
Score = 28.1 bits (61), Expect = 1.4
Identities = 12/49 (24%), Positives = 20/49 (40%)
Frame = -3
Query: 7 YNLCKFSPNTTSCNRFYCTCDIISWFHWCLFSCCMCTMVVLGDYVIANS 55
+N C + C+ +C+C + C CC C GD + +S
Sbjct: 561 HNCCYCCCSKVPCHCCHCSCTLQCCSFLCTLHCCSCCGNWSGDILFCSS 415
>AV774500
Length = 513
Score = 27.3 bits (59), Expect = 2.4
Identities = 20/47 (42%), Positives = 25/47 (52%), Gaps = 1/47 (2%)
Frame = -3
Query: 169 LLCYECDSCKAGVLEDIRRNWHKLSVLTVTMLILLIGI-YSIGCCAF 214
LLC C + + RN KLSVL ML L++ I YS GC +F
Sbjct: 271 LLCISQILCITYMYAQVLRNGRKLSVL---ML*LVLTIQYSFGCHSF 140
>BP032030
Length = 454
Score = 27.3 bits (59), Expect = 2.4
Identities = 12/44 (27%), Positives = 19/44 (42%)
Frame = +2
Query: 141 PPTACNYNMEAVMMTQDSDCYKWSNEPTLLCYECDSCKAGVLED 184
PP + E + + S+ ++W + LC C K G L D
Sbjct: 299 PPPVLPQSSETLTIVHGSEAFQWKSTILSLCPACRKNKPGGLSD 430
>TC10107 similar to UP|Q9MB24 (Q9MB24) S1-3 protein (Fragment), partial
(65%)
Length = 575
Score = 26.9 bits (58), Expect = 3.1
Identities = 14/42 (33%), Positives = 21/42 (49%)
Frame = +2
Query: 22 FYCTCDIISWFHWCLFSCCMCTMVVLGDYVIANSSTIRFDYI 63
F C+C +W H C+F C+ M VL +T+ + YI
Sbjct: 461 FRCSC---TWLHICIFL*CIIYMYVL--------NTLHYGYI 553
>AV766560
Length = 398
Score = 26.9 bits (58), Expect = 3.1
Identities = 11/26 (42%), Positives = 17/26 (65%)
Frame = -3
Query: 20 NRFYCTCDIISWFHWCLFSCCMCTMV 45
++F+CTC I S F +C CC T++
Sbjct: 114 HKFHCTCKISS*F*FC--KCCTSTLL 43
>TC18216 similar to GB|CAA28991.1|34041|HSKER65A keratin type II (AA1-215)
{Homo sapiens;} , partial (16%)
Length = 501
Score = 26.9 bits (58), Expect = 3.1
Identities = 14/48 (29%), Positives = 18/48 (37%)
Frame = +1
Query: 31 WFHWCLFSCCMCTMVVLGDYVIANSSTIRFDYIWFWCD**RWWCRSAW 78
W+ WC SCC S + + WFWC WW +W
Sbjct: 49 WWVWC--SCC---------------SH*WWIWWWFWCCSFSWWIWRSW 141
>TC17021 homologue to UP|Q9SSP4 (Q9SSP4) F3N23.2 protein
(At1g72820/F3N23_2), partial (22%)
Length = 488
Score = 26.6 bits (57), Expect = 4.1
Identities = 10/17 (58%), Positives = 11/17 (63%)
Frame = +2
Query: 32 FHWCLFSCCMCTMVVLG 48
FH CLFS C+C LG
Sbjct: 347 FHSCLFSSCICFFFSLG 397
>BP060559
Length = 386
Score = 26.6 bits (57), Expect = 4.1
Identities = 15/40 (37%), Positives = 21/40 (52%)
Frame = -2
Query: 7 YNLCKFSPNTTSCNRFYCTCDIISWFHWCLFSCCMCTMVV 46
Y KF N+ SC +Y F +C+FSCC T+V+
Sbjct: 289 YGQGKFWLNSRSC--WYSVN-----FRFCIFSCCTSTLVL 191
>TC15614 similar to PIR|T00553|T00553 disease resistance response protein
homolog F12L6.9 - Arabidopsis thaliana
{Arabidopsis thaliana;}, partial (23%)
Length = 475
Score = 26.6 bits (57), Expect = 4.1
Identities = 9/20 (45%), Positives = 13/20 (65%)
Frame = +2
Query: 34 WCLFSCCMCTMVVLGDYVIA 53
WC SCCM +++G + IA
Sbjct: 74 WCSQSCCMVESIMMGWWTIA 133
>TC15341 similar to UP|Q9M0H8 (Q9M0H8) Predicted proline-rich protein,
partial (33%)
Length = 1002
Score = 26.2 bits (56), Expect = 5.4
Identities = 9/19 (47%), Positives = 9/19 (47%)
Frame = -2
Query: 24 CTCDIISWFHWCLFSCCMC 42
CTC W H C CC C
Sbjct: 929 CTC*GNCWGHCCCCCCCCC 873
>TC16190 weakly similar to UP|Q9LQ75 (Q9LQ75) T1N6.22 protein, partial (41%)
Length = 1221
Score = 26.2 bits (56), Expect = 5.4
Identities = 13/35 (37%), Positives = 19/35 (54%), Gaps = 5/35 (14%)
Frame = -2
Query: 7 YNLCKFSPNTTSCNRFYCTCD-----IISWFHWCL 36
+ LCK+S ++ CN CT +I+WF W L
Sbjct: 1094 FTLCKYSCSSNVCN---CTVAGDIGLLITWFLWLL 999
>TC9607 similar to UP|Q8S9J2 (Q8S9J2) At2g33840/T1B8.14, partial (24%)
Length = 615
Score = 26.2 bits (56), Expect = 5.4
Identities = 11/12 (91%), Positives = 11/12 (91%)
Frame = +3
Query: 189 WHKLSVLTVTML 200
WHKLSVLTVT L
Sbjct: 546 WHKLSVLTVTNL 581
>BP041085
Length = 449
Score = 26.2 bits (56), Expect = 5.4
Identities = 11/32 (34%), Positives = 16/32 (49%), Gaps = 4/32 (12%)
Frame = +1
Query: 12 FSPNTTSCNRF----YCTCDIISWFHWCLFSC 39
F TT+ NR+ C C +I+ HW + C
Sbjct: 316 FHAPTTNYNRY*LFVICRCWVINMIHWVMLFC 411
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.343 0.149 0.575
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 6,197,195
Number of Sequences: 28460
Number of extensions: 112938
Number of successful extensions: 1637
Number of sequences better than 10.0: 55
Number of HSP's better than 10.0 without gapping: 1605
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 1632
length of query: 241
length of database: 4,897,600
effective HSP length: 88
effective length of query: 153
effective length of database: 2,393,120
effective search space: 366147360
effective search space used: 366147360
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 38 (21.6 bits)
S2: 54 (25.4 bits)
Medicago: description of AC126015.14


