

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC126015.11 + phase: 0
(371 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
Score E
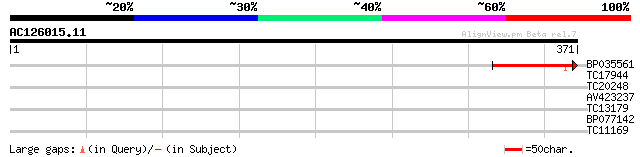
Sequences producing significant alignments: (bits) Value
BP035561 94 4e-20
TC17944 38 0.002
TC20248 similar to UP|Q9ZVA9 (Q9ZVA9) F9K20.4 protein, partial (8%) 36 0.012
AV423237 28 2.4
TC13179 similar to UP|Q9ZVA9 (Q9ZVA9) F9K20.4 protein, partial (5%) 28 3.2
BP077142 26 9.3
TC11169 similar to UP|Q9SL72 (Q9SL72) At2g20090 protein, partial... 26 9.3
>BP035561
Length = 495
Score = 94.0 bits (232), Expect = 4e-20
Identities = 46/57 (80%), Positives = 51/57 (88%), Gaps = 2/57 (3%)
Frame = -3
Query: 317 RPSKPVPGDCGYHLHAHRLFLSHPITNEIIEMIAPLPSILQTHEEA--IAQAQQTTA 371
RP+KPVPGDCGY+LHAH+L LSHPITNEII++IAPLPSIL T EEA IA AQQTTA
Sbjct: 490 RPTKPVPGDCGYYLHAHKLILSHPITNEIIDIIAPLPSILMTAEEAKEIAVAQQTTA 320
>TC17944
Length = 537
Score = 38.1 bits (87), Expect = 0.002
Identities = 22/29 (75%), Positives = 24/29 (81%), Gaps = 2/29 (6%)
Frame = +3
Query: 345 IIEMIAPLPSILQTHEEA--IAQAQQTTA 371
II++IAPLPSIL T EEA IA AQQTTA
Sbjct: 120 IIDIIAPLPSILMTAEEAKEIAVAQQTTA 206
>TC20248 similar to UP|Q9ZVA9 (Q9ZVA9) F9K20.4 protein, partial (8%)
Length = 496
Score = 35.8 bits (81), Expect = 0.012
Identities = 34/117 (29%), Positives = 52/117 (44%)
Frame = +3
Query: 109 KPSGLQVLPGGLFQQRTVLTQLQWETNNKSTIEAHQKPHPVPVHRLGRGTSGILLCAKTK 168
KP G+ V GG+ +R++ + S ++ P VHRL R SGIL+ +T+
Sbjct: 3 KPPGMPV-QGGIDIKRSL------DVVAASCLKYEYSEPPRLVHRLDRDCSGILVMGRTQ 161
Query: 169 LARARLASYFADGTSRIEGKGRHTNQELGKIAKIYRALVSGIINNDQVTINQPIGMV 225
+ L S F + TS N+ L + YRALV G + + +G V
Sbjct: 162 TSATVLHSIFREKTSSASDDLGTENRVL---QRKYRALVIGCPRRPRGLVTASLGKV 323
>AV423237
Length = 310
Score = 28.1 bits (61), Expect = 2.4
Identities = 12/29 (41%), Positives = 19/29 (65%)
Frame = +1
Query: 275 RIHLSFIGHPLLGDPLYAVGGQPKCLDCD 303
R+HL F+G ++G+ Y VG C+DC+
Sbjct: 91 RVHLLFMGLIIMGNLCYLVG----CIDCE 165
>TC13179 similar to UP|Q9ZVA9 (Q9ZVA9) F9K20.4 protein, partial (5%)
Length = 533
Score = 27.7 bits (60), Expect = 3.2
Identities = 14/40 (35%), Positives = 21/40 (52%), Gaps = 1/40 (2%)
Frame = +2
Query: 268 SGRPHQIRIHLS-FIGHPLLGDPLYAVGGQPKCLDCDFLD 306
+GR HQ+R+H + +G P++GD Y K D D
Sbjct: 11 TGRKHQLRVHCAEVLGTPIVGDYKYVWQAHRKWSHFDLSD 130
>BP077142
Length = 385
Score = 26.2 bits (56), Expect = 9.3
Identities = 9/18 (50%), Positives = 13/18 (72%)
Frame = +3
Query: 301 DCDFLDESFAEDGGYTRP 318
DC+ +E AE+GG T+P
Sbjct: 144 DCNLFEEKHAENGGKTKP 197
>TC11169 similar to UP|Q9SL72 (Q9SL72) At2g20090 protein, partial (14%)
Length = 650
Score = 26.2 bits (56), Expect = 9.3
Identities = 9/15 (60%), Positives = 12/15 (80%)
Frame = +1
Query: 327 GYHLHAHRLFLSHPI 341
G+HLH H+L LS P+
Sbjct: 118 GHHLHNHKLLLSFPL 162
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.318 0.137 0.409
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 6,270,484
Number of Sequences: 28460
Number of extensions: 87480
Number of successful extensions: 439
Number of sequences better than 10.0: 15
Number of HSP's better than 10.0 without gapping: 436
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 439
length of query: 371
length of database: 4,897,600
effective HSP length: 92
effective length of query: 279
effective length of database: 2,279,280
effective search space: 635919120
effective search space used: 635919120
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 56 (26.2 bits)
Medicago: description of AC126015.11


