

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC126014.9 + phase: 0
(233 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
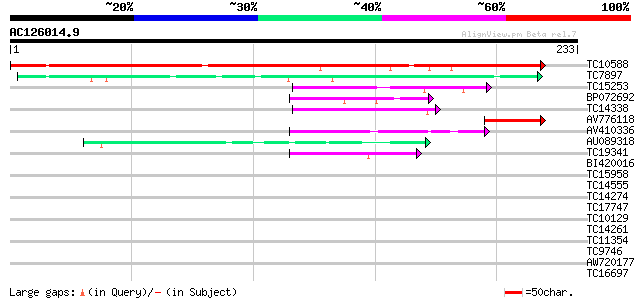
Score E
Sequences producing significant alignments: (bits) Value
TC10588 similar to UP|Q9LPU4 (Q9LPU4) T22I11.8, partial (36%) 322 3e-89
TC7897 weakly similar to UP|ENL2_ARATH (Q9T076) Early nodulin-li... 50 3e-07
TC15253 weakly similar to UP|Q9ZT16 (Q9ZT16) Arabinogalactan-pro... 45 1e-05
BP072692 43 5e-05
TC14338 similar to UP|Q9MAH5 (Q9MAH5) F12M16.16, partial (33%) 41 2e-04
AV776118 40 3e-04
AV410336 40 5e-04
AU089318 40 5e-04
TC19341 similar to UP|Q9CAD1 (Q9CAD1) GMP synthase, partial (28%) 39 8e-04
BI420016 39 0.001
TC15958 similar to GB|AAP37838.1|30725632|BT008479 At4g24220 {Ar... 38 0.001
TC14555 similar to UP|Q43558 (Q43558) Proline rich protein precu... 38 0.002
TC14274 weakly similar to GB|AAF79559.1|8778551|AC022464 F22G5.3... 38 0.002
TC17747 homologue to UP|Q8WML4 (Q8WML4) MUC1 protein precursor, ... 38 0.002
TC10129 weakly similar to UP|Q7T9Y7 (Q7T9Y7) Odv-e66, partial (10%) 37 0.002
TC14261 weakly similar to UP|Q05929 (Q05929) EDGP precursor (Fra... 37 0.003
TC11354 similar to UP|CAE54480 (CAE54480) Alpha glucosidase II ... 37 0.003
TC9746 37 0.003
AW720177 37 0.004
TC16697 37 0.004
>TC10588 similar to UP|Q9LPU4 (Q9LPU4) T22I11.8, partial (36%)
Length = 976
Score = 322 bits (825), Expect = 3e-89
Identities = 168/230 (73%), Positives = 186/230 (80%), Gaps = 10/230 (4%)
Frame = +1
Query: 1 MSLPIFFYFLILSLFFKLSHSTTILVDGSSEWKNPTVSIGDSITFKHKQNYNLYIFKNQK 60
MS PIFF+FL+LSLF KLSHSTTI+VDG SEWKNPTV IGDSI FKHKQN++LYIFKNQ
Sbjct: 22 MSFPIFFFFLLLSLF-KLSHSTTIVVDGFSEWKNPTVHIGDSIIFKHKQNHSLYIFKNQN 198
Query: 61 AFNLCNFTQANLLTDPSTTSCSYTWHPSRVGFFYFTFSNDSLKACQDSQKLAIKVTPTKA 120
AFN+CN TQA LLT+P+TT SYTWHPSR GFFYFTF+N SLK CQ+SQKLAIKV+ A
Sbjct: 199 AFNICNLTQATLLTNPTTT--SYTWHPSRTGFFYFTFNNGSLKTCQNSQKLAIKVSSAAA 372
Query: 121 SAPEAS------SPMPTTPGPSSGGDIQSSPSFPWPFHPHQ-GSSPGPAPTPEASSPI-T 172
+ P+ S +P+ TTP PSSGGD+ SSPSFPWPF PHQ +SPGPAP ASSP T
Sbjct: 373 APPQGSEVTPEPTPVVTTPAPSSGGDVPSSPSFPWPFRPHQAAASPGPAPA--ASSPAET 546
Query: 173 VPLVPYKG--SGDGMPFINSNPAVPLPTGEVDSATIHPLATSGHQGQVPI 220
VPLVP KG + MPFINSNPAVPLPTGEVDSATI PL TSGHQGQV I
Sbjct: 547 VPLVPGKGGSTSTSMPFINSNPAVPLPTGEVDSATIRPLPTSGHQGQVMI 696
>TC7897 weakly similar to UP|ENL2_ARATH (Q9T076) Early nodulin-like protein
2 precursor (Phytocyanin-like protein), partial (31%)
Length = 1482
Score = 50.1 bits (118), Expect = 3e-07
Identities = 64/258 (24%), Positives = 86/258 (32%), Gaps = 42/258 (16%)
Frame = +3
Query: 4 PIFFYFLILSLFFKLSHSTTILVDGSSEW-KNPTVS-----------IGDSITFKHKQNY 51
P+ FL+ SL S + V GS W NP+ S I D+I FK+ +
Sbjct: 156 PLCLLFLLFSLL-PSSQAHKFYVGGSKGWIPNPSESYTLWAGRNRFQIKDTIVFKYTKGS 332
Query: 52 NLYIFKNQKAFNLCNFTQANLLTDPSTTSCSYTWHPSRVGFFYFTFSNDSLKACQDSQKL 111
+ + ++ F+ CN +AN + +T+ R G FYF D C+ QK+
Sbjct: 333 DSVLEVKKEDFDNCN--KANPIKKFEDGDTEFTF--DRSGPFYFISGKDG--NCEKGQKM 494
Query: 112 AI------------------------KVTPTKASAPEASSPMPT------TPGPSSGGDI 141
+ K +P S P A P P P P+
Sbjct: 495 ILVVISPRGTQPPPSPPPKSSPSPSPKASPPSGSPPTAGEPSPAGTPPSVLPSPAGAPVP 674
Query: 142 QSSPSFPWPFHPHQGSSPGPAPTPEASSPITVPLVPYKGSGDGMPFINSNPAVPLPTGEV 201
PS P SP P P A+SP P P P P G
Sbjct: 675 AGGPSAGTPPSASTSPSPSPVSAPPAASPAPGAESPSSSPTSNSPAGGPTATSPSPAG-- 848
Query: 202 DSATIHPLATSGHQGQVP 219
DS P A S G P
Sbjct: 849 DSPAGGPPAPSPTTGDTP 902
>TC15253 weakly similar to UP|Q9ZT16 (Q9ZT16) Arabinogalactan-protein
(ATAGP4) (AT5G10430/F12B17_220), partial (42%)
Length = 575
Score = 45.1 bits (105), Expect = 1e-05
Identities = 32/84 (38%), Positives = 41/84 (48%), Gaps = 2/84 (2%)
Frame = +3
Query: 117 PTKASAPEASSPMPTTPGPSSGGDIQSSPSFPWPFHPHQGSSPGPAPTPEASS-PITVPL 175
P A+ P A++P PTT P++ +SP P P +SP APTP ASS P +P
Sbjct: 219 PPPATPPPAATPAPTTTPPAATPAPSASPPAPTPT-----ASPTGAPTPSASSPPAPIPS 383
Query: 176 VPYKGSGDGM-PFINSNPAVPLPT 198
P G G P NS P P+
Sbjct: 384 GPASGPGPAAGPGPNSADTPPPPS 455
Score = 30.8 bits (68), Expect = 0.21
Identities = 20/62 (32%), Positives = 29/62 (46%), Gaps = 1/62 (1%)
Frame = +3
Query: 116 TPTKASAPEASS-PMPTTPGPSSGGDIQSSPSFPWPFHPHQGSSPGPAPTPEASSPITVP 174
+PT A P ASS P P GP+SG + P P+ +P P P A+ + P
Sbjct: 327 SPTGAPTPSASSPPAPIPSGPASGPGPAAGPG------PNSADTP---PPPSAAFSLNQP 479
Query: 175 LV 176
++
Sbjct: 480 II 485
>BP072692
Length = 422
Score = 42.7 bits (99), Expect = 5e-05
Identities = 26/61 (42%), Positives = 35/61 (56%), Gaps = 2/61 (3%)
Frame = +2
Query: 116 TPTKASAPEASSPMPTTPGPS-SGGDIQSSPSFPW-PFHPHQGSSPGPAPTPEASSPITV 173
+PT ++AP SSP +TP PS S + P FP+ P HP SS PTP +S+P +
Sbjct: 233 SPTPSAAPPLSSPPSSTPSPSPSTSSSKVGPDFPFSPTHPPSRSS--QPPTPPSSTPTSD 406
Query: 174 P 174
P
Sbjct: 407 P 409
>TC14338 similar to UP|Q9MAH5 (Q9MAH5) F12M16.16, partial (33%)
Length = 1751
Score = 40.8 bits (94), Expect = 2e-04
Identities = 24/68 (35%), Positives = 30/68 (43%), Gaps = 7/68 (10%)
Frame = +3
Query: 117 PTKASAPEASSPMPTTPGPSSGGDIQSSPSFPWPFHPHQGSSPGPAPTPEASSP------ 170
P+ + P P P P S + PS P+P P SSP PAP+P A P
Sbjct: 81 PSSLAPSPLQPPPPLPPSPPSLSSAATVPSPPYPSPPAATSSPPPAPSPPAPPPPPSTIL 260
Query: 171 -ITVPLVP 177
+T P VP
Sbjct: 261 TLTGPTVP 284
Score = 26.2 bits (56), Expect = 5.2
Identities = 19/56 (33%), Positives = 22/56 (38%), Gaps = 3/56 (5%)
Frame = +3
Query: 153 PHQGSSPGPAPTPEASSPITVPLVPY---KGSGDGMPFINSNPAVPLPTGEVDSAT 205
P Q P P P SS TVP PY + P S PA P P + + T
Sbjct: 102 PLQPPPPLPPSPPSLSSAATVPSPPYPSPPAATSSPPPAPSPPAPPPPPSTILTLT 269
>AV776118
Length = 334
Score = 40.0 bits (92), Expect = 3e-04
Identities = 19/25 (76%), Positives = 19/25 (76%)
Frame = -1
Query: 196 LPTGEVDSATIHPLATSGHQGQVPI 220
LPTGEVDSATI PL S H GQV I
Sbjct: 334 LPTGEVDSATIRPLPPSAHHGQVMI 260
>AV410336
Length = 358
Score = 39.7 bits (91), Expect = 5e-04
Identities = 28/82 (34%), Positives = 37/82 (44%)
Frame = +1
Query: 116 TPTKASAPEASSPMPTTPGPSSGGDIQSSPSFPWPFHPHQGSSPGPAPTPEASSPITVPL 175
T T ++AP +P P TP P S +P+ P PH G + PA T +S T P
Sbjct: 40 TSTSSTAPPPHAPPPPTPNPKSPSSSSPTPTSP---SPHSGRNSSPATTTFTTSTST-PT 207
Query: 176 VPYKGSGDGMPFINSNPAVPLP 197
P S +P +S A LP
Sbjct: 208 PPPPSS---LPAASSTAASLLP 264
>AU089318
Length = 641
Score = 39.7 bits (91), Expect = 5e-04
Identities = 37/144 (25%), Positives = 58/144 (39%), Gaps = 1/144 (0%)
Frame = +3
Query: 31 EWKNPT-VSIGDSITFKHKQNYNLYIFKNQKAFNLCNFTQANLLTDPSTTSCSYTWHPSR 89
+W P +GD+I F++ ++ + + CN + S T H
Sbjct: 162 KWAAPXNFQVGDTIIFEYNAQFHNVMRVTHAMYXTCNASSPIATFSTGNDSIKITNHGHH 341
Query: 90 VGFFYFTFSNDSLKACQDSQKLAIKVTPTKASAPEASSPMPTTPGPSSGGDIQSSPSFPW 149
FF+ CQ QK+ I V SA A++P P TP PSS ++P
Sbjct: 342 --FFFCGVPGH----CQAGQKVDINV--ISVSAATAAAPTP-TPAPSSPSSSLATP---- 482
Query: 150 PFHPHQGSSPGPAPTPEASSPITV 173
S+ PAP+P ++P+ V
Sbjct: 483 -------SANVPAPSPNNATPLIV 533
>TC19341 similar to UP|Q9CAD1 (Q9CAD1) GMP synthase, partial (28%)
Length = 484
Score = 38.9 bits (89), Expect = 8e-04
Identities = 22/55 (40%), Positives = 27/55 (49%), Gaps = 1/55 (1%)
Frame = +1
Query: 116 TPTKASAPEASSPMPTTPGPSSGGDIQSSPS-FPWPFHPHQGSSPGPAPTPEASS 169
+P+ + AP S P P + PSS +PS P P SS GPAPT SS
Sbjct: 43 SPSASPAPPPSPPSPNSTPPSSSSPAAPTPSTLPTPLPSLTASSTGPAPTASLSS 207
>BI420016
Length = 559
Score = 38.5 bits (88), Expect = 0.001
Identities = 22/67 (32%), Positives = 32/67 (46%), Gaps = 1/67 (1%)
Frame = +2
Query: 116 TPTKASAPEASSP-MPTTPGPSSGGDIQSSPSFPWPFHPHQGSSPGPAPTPEASSPITVP 174
TP+ S+P + P P +P P++ S PS P P P +P P P+ S+ + P
Sbjct: 236 TPSPPSSPTLTPPPSPPSPTPNAPLSPSSPPSPPTPPSPPPSPAPNPTPSATGSTTPSSP 415
Query: 175 LVPYKGS 181
L P S
Sbjct: 416 LTPTSAS 436
Score = 35.4 bits (80), Expect = 0.009
Identities = 30/97 (30%), Positives = 35/97 (35%), Gaps = 1/97 (1%)
Frame = +2
Query: 117 PTKASAPEASSPMPTTPGPSSGGDIQSSPSFPWPFHPHQGSSPGPAPTPEASSPITVPLV 176
P AS P S TP PS P P P P SSP PTP + P P
Sbjct: 221 PVPASTPSPPSSPTLTPPPS--------PPSPTPNAPLSPSSPPSPPTPPSPPPSPAPNP 376
Query: 177 PYKGSGDGMPFINSNPAVPLPTG-EVDSATIHPLATS 212
+G P S+P P S+ P +TS
Sbjct: 377 TPSATGSTTP---SSPLTPTSASLSSPSSLFSPASTS 478
>TC15958 similar to GB|AAP37838.1|30725632|BT008479 At4g24220 {Arabidopsis
thaliana;}, partial (52%)
Length = 800
Score = 38.1 bits (87), Expect = 0.001
Identities = 28/89 (31%), Positives = 41/89 (45%), Gaps = 12/89 (13%)
Frame = +3
Query: 118 TKASAPEASSPMPTTPGPSSGGDIQSSPSFPWPFHPHQGSSPGPAPTP------------ 165
T+++ A+SP+PTTP PSS SSP+ P +SPGP TP
Sbjct: 384 TQSTTSSATSPIPTTPKPSS----PSSPT-----SPTSSTSPGPTATPNRRIARSTAPCS 536
Query: 166 EASSPITVPLVPYKGSGDGMPFINSNPAV 194
E SS + P P + P +++ A+
Sbjct: 537 ETSSAPSSPTRPISAT*RSKPAVSTTSAL 623
>TC14555 similar to UP|Q43558 (Q43558) Proline rich protein precursor,
partial (43%)
Length = 1038
Score = 37.7 bits (86), Expect = 0.002
Identities = 33/99 (33%), Positives = 41/99 (41%), Gaps = 3/99 (3%)
Frame = +2
Query: 116 TPTKASAPEASSPMPTTPGPSSGGDIQSSPSFPWPFHPHQGSSPGPAPTP---EASSPIT 172
+PT + AP P PTTP P + QSSP P SSP PA TP ++S P
Sbjct: 200 SPTTSPAP----PTPTTPQPPAA---QSSPPPAQSSPPPVQSSPPPASTPPPAQSSPPPV 358
Query: 173 VPLVPYKGSGDGMPFINSNPAVPLPTGEVDSATIHPLAT 211
P + + P PA P P + P AT
Sbjct: 359 SSPPPVQSTPPPAPASTPPPASPPPFSPPPATPPPPAAT 475
Score = 37.0 bits (84), Expect = 0.003
Identities = 26/92 (28%), Positives = 32/92 (34%)
Frame = +2
Query: 106 QDSQKLAIKVTPTKASAPEASSPMPTTPGPSSGGDIQSSPSFPWPFHPHQGSSPGPAPTP 165
Q S A P ++S P SSP P P P+ P PF P P P P
Sbjct: 299 QSSPPPASTPPPAQSSPPPVSSPPPVQSTPPPAPASTPPPASPPPFSP----PPATPPPP 466
Query: 166 EASSPITVPLVPYKGSGDGMPFINSNPAVPLP 197
A+ P + VP + S P P
Sbjct: 467 AATPPPALTPVPATSPAPAPAKVKSKSPAPAP 562
>TC14274 weakly similar to GB|AAF79559.1|8778551|AC022464 F22G5.35
{Arabidopsis thaliana;} , partial (16%)
Length = 1208
Score = 37.7 bits (86), Expect = 0.002
Identities = 30/96 (31%), Positives = 40/96 (41%), Gaps = 6/96 (6%)
Frame = +3
Query: 116 TPTKASAPEA-SSPMPTTPG--PSSGGDIQSSPSFP---WPFHPHQGSSPGPAPTPEASS 169
T + S P A S P PTTP P SG + S S PF P + S+P P P SS
Sbjct: 147 TSSSGSTPTAVSPPNPTTPATPPPSGTSVSPSLSLTPSLTPFSPSRSSTPNPPIPPNPSS 326
Query: 170 PITVPLVPYKGSGDGMPFINSNPAVPLPTGEVDSAT 205
P + P + P ++P+ P +T
Sbjct: 327 PPSASSSPTSTTSMIPPASANSPSSAPPAAPTARST 434
>TC17747 homologue to UP|Q8WML4 (Q8WML4) MUC1 protein precursor, partial
(4%)
Length = 507
Score = 37.7 bits (86), Expect = 0.002
Identities = 32/109 (29%), Positives = 48/109 (43%), Gaps = 11/109 (10%)
Frame = +2
Query: 116 TPTKASAPEASSPMPTTPGPSSGGDIQSSPSFPWPFHPHQGSSPGPAPTPEASSPITVPL 175
+P+ S+P + SP P P P+S + SSP+ P P GSSP P+P +S L
Sbjct: 95 SPSTTSSP-SPSPSPFVPAPTSS-PVASSPAAPTPQSWSPGSSPAPSPMGSSS------L 250
Query: 176 VPYKG-----------SGDGMPFINSNPAVPLPTGEVDSATIHPLATSG 213
+ ++G S G + V + G V + T+ LA G
Sbjct: 251 ISHEGVEGGRFTDSDSSSSGSGEMTGRKKVAIVIGVVVAVTVVVLAGMG 397
>TC10129 weakly similar to UP|Q7T9Y7 (Q7T9Y7) Odv-e66, partial (10%)
Length = 562
Score = 37.4 bits (85), Expect = 0.002
Identities = 28/88 (31%), Positives = 40/88 (44%), Gaps = 3/88 (3%)
Frame = +1
Query: 116 TPTKASAPEASSPMPTTPGPS-SGGDIQSSPSFPWPFHPHQGSSPGPAPTPE-ASSPITV 173
TP+ + P + P TP PS S ++ S P P P + H +P P TP S P +
Sbjct: 46 TPSPGNQPPHTPIPPKTPSPSYSPPNVPSPPKAPSP-NNHPPYTPTPPKTPSPTSQPPYI 222
Query: 174 PLVPYKGSGDGM-PFINSNPAVPLPTGE 200
PL P S P + + P+ P P +
Sbjct: 223 PLPPKTPSPISQPPHVPTPPSTPSPISQ 306
Score = 32.0 bits (71), Expect = 0.094
Identities = 26/88 (29%), Positives = 36/88 (40%), Gaps = 3/88 (3%)
Frame = +1
Query: 116 TPTKASAPEASSPMPTTPGPSS-GGDIQSSPSFPWPF-HPHQGSSPGPAPTPEASSPITV 173
TP+ S P + P TP P+S + P+ P P P +P P+P + P +
Sbjct: 286 TPSPISQPPYTPTPPKTPSPTSQPPHTPTPPNTPSPISQPPYTPTPPKTPSP-TNQPPHI 462
Query: 174 PLVPYKGSGDGM-PFINSNPAVPLPTGE 200
P P S P S P P PT +
Sbjct: 463 PSPPNSSSPTSQPPNTPSPPKTPSPTSQ 546
Score = 26.6 bits (57), Expect = 4.0
Identities = 15/50 (30%), Positives = 20/50 (40%)
Frame = +1
Query: 116 TPTKASAPEASSPMPTTPGPSSGGDIQSSPSFPWPFHPHQGSSPGPAPTP 165
TPT P ++ P P P + SSP+ P P +P P P
Sbjct: 412 TPTPPKTPSPTNQPPHIPSPPN----SSSPTSQPPNTPSPPKTPSPTSQP 549
>TC14261 weakly similar to UP|Q05929 (Q05929) EDGP precursor (Fragment),
partial (84%)
Length = 1604
Score = 37.0 bits (84), Expect = 0.003
Identities = 27/95 (28%), Positives = 39/95 (40%), Gaps = 19/95 (20%)
Frame = +2
Query: 98 SNDSLKACQDSQKLAIKVTPTKASAPEASSPMPTTPGPSSGGDIQSSPSFPWP------- 150
SN S K+ ++ K ++ A+ S+ PTTP + + PS PWP
Sbjct: 170 SNTSPKSTKEHPKSQ*NLSSISAATSCGSTVKPTTPPQPTAQHAANPPSAPWPDPAAAAI 349
Query: 151 -FHP-----------HQGSSPGPAPTPEASSPITV 173
HP ++P P P P ASSP T+
Sbjct: 350 ASHPPSPAATTTLVASSRTTPSPEPPPAASSPKTL 454
>TC11354 similar to UP|CAE54480 (CAE54480) Alpha glucosidase II , partial
(6%)
Length = 583
Score = 37.0 bits (84), Expect = 0.003
Identities = 23/62 (37%), Positives = 31/62 (49%), Gaps = 3/62 (4%)
Frame = -1
Query: 116 TPTKASAPEASSPMPTT-PGPSSGGDIQSSPSFPWPFHPHQGSSP--GPAPTPEASSPIT 172
T T +SA ++ +P+T P P+S SSPS P P S P P P P ++SP
Sbjct: 226 TTTASSASPSTKTLPSTRPRPASESPTSSSPSSPPPSSGSSVSPPRISPPPPPPSNSPTA 47
Query: 173 VP 174
P
Sbjct: 46 TP 41
Score = 32.3 bits (72), Expect = 0.072
Identities = 34/122 (27%), Positives = 47/122 (37%), Gaps = 26/122 (21%)
Frame = -1
Query: 117 PTKASAPEASSPMPTTPGPSSGGDIQSSPSFPWPFH-PHQGSSPGPAPT----------- 164
P A P S P +P P++ SSP+ P P+ SSP P+ T
Sbjct: 367 PDPAPPPPPPSSPPMSPSPTATSPPTSSPNTHPPTPTPNP*SSPSPSTTTASSASPSTKT 188
Query: 165 ------------PEASSPITVPLVPYKGSGDGMPFINSNPAVP--LPTGEVDSATIHPLA 210
P +SSP + P P GS P I+ P P PT S+ +P +
Sbjct: 187 LPSTRPRPASESPTSSSPSSPP--PSSGSSVSPPRISPPPPPPSNSPTATPPSSATNPSS 14
Query: 211 TS 212
+S
Sbjct: 13 SS 8
Score = 32.0 bits (71), Expect = 0.094
Identities = 29/111 (26%), Positives = 41/111 (36%), Gaps = 1/111 (0%)
Frame = -1
Query: 88 SRVGFFYFTFSNDSLKACQDSQKLAIKVTPTKASAPEASSPMPTTPGPSSGGDIQSSPSF 147
S + FF+FT ++ + K + S +SS P++PG +
Sbjct: 535 SPLSFFHFTCVRIAMTLTR*GIKRCV-------SRSSSSSAPPSSPGRKKSSETA----- 392
Query: 148 PWPFHPHQGSSPGPA-PTPEASSPITVPLVPYKGSGDGMPFINSNPAVPLP 197
PH S P PA P P SSP P P N++P P P
Sbjct: 391 ---IRPHSASEPDPAPPPPPPSSPPMSPSPTATSPPTSSP--NTHPPTPTP 254
>TC9746
Length = 637
Score = 37.0 bits (84), Expect = 0.003
Identities = 37/106 (34%), Positives = 41/106 (37%), Gaps = 7/106 (6%)
Frame = +2
Query: 118 TKASAPEASSPMPTTP--GPSSGGDIQSSP-----SFPWPFHPHQGSSPGPAPTPEASSP 170
T S P +SSP TT SS QSSP P P HPH SP P P AS
Sbjct: 341 TLTSPPHSSSPHSTTSPLASSSASPCQSSP*SPSPPSPSPKHPH---SPSPRSPPPAS-- 505
Query: 171 ITVPLVPYKGSGDGMPFINSNPAVPLPTGEVDSATIHPLATSGHQG 216
P PY F +S P V P+ S A S +G
Sbjct: 506 ---PSPPY--------FSSSAPPVSCPSSTATSLCQSSAAFSSPRG 610
>AW720177
Length = 505
Score = 36.6 bits (83), Expect = 0.004
Identities = 32/113 (28%), Positives = 42/113 (36%), Gaps = 7/113 (6%)
Frame = +1
Query: 114 KVTPTKASAPEASSPMPTTPGPSSGGDIQSSPSFPWPFHPHQGSSPGPAPTPEASSP--- 170
K P+K S P S +P I S PSFP P P S P+P S+P
Sbjct: 145 KSPPSKTSKPSGHSSESNSPTSKPTSPI-SPPSFPPPPPPTHSLSTSSTPSPPLSTPPSP 321
Query: 171 ----ITVPLVPYKGSGDGMPFINSNPAVPLPTGEVDSATIHPLATSGHQGQVP 219
T P P + P S P+ P T +AT + + S +P
Sbjct: 322 SRNSATPP--PTSLAXSSKPKATSTPSSPPSTATSPTATYYSVVDSSSTPPLP 474
>TC16697
Length = 635
Score = 36.6 bits (83), Expect = 0.004
Identities = 25/96 (26%), Positives = 39/96 (40%), Gaps = 3/96 (3%)
Frame = -1
Query: 123 PEASSPMPTTPGPSSGGDIQSSPSFPWPFHPHQGSSPGPA---PTPEASSPITVPLVPYK 179
P P P++P P + + +P+ P + Q P P+ PT A P L P
Sbjct: 572 PPNHPPPPSSPSPPAPSQRKPAPATAPPSNSSQSPPPNPSSTDPTSPAPQPHPSHLSPAS 393
Query: 180 GSGDGMPFINSNPAVPLPTGEVDSATIHPLATSGHQ 215
S G P + +P+ P + S + P+ S Q
Sbjct: 392 SSSQGSPVPSPSPSPP-----ITSPSSPPIGDSSRQ 300
Score = 35.4 bits (80), Expect = 0.009
Identities = 25/72 (34%), Positives = 30/72 (40%), Gaps = 3/72 (4%)
Frame = -1
Query: 118 TKASAPEASSPMPTTPGPSSGGDIQSSPSFPWPFH--PHQGSSPG-PAPTPEASSPITVP 174
+++ P SS PT+P P P P H P SS G P P+P S PIT P
Sbjct: 479 SQSPPPNPSSTDPTSPAPQ-----------PHPSHLSPASSSSQGSPVPSPSPSPPITSP 333
Query: 175 LVPYKGSGDGMP 186
P G P
Sbjct: 332 SSPPIGDSSRQP 297
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.317 0.133 0.418
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 5,494,497
Number of Sequences: 28460
Number of extensions: 102564
Number of successful extensions: 1758
Number of sequences better than 10.0: 353
Number of HSP's better than 10.0 without gapping: 1422
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 1619
length of query: 233
length of database: 4,897,600
effective HSP length: 87
effective length of query: 146
effective length of database: 2,421,580
effective search space: 353550680
effective search space used: 353550680
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.6 bits)
S2: 54 (25.4 bits)
Medicago: description of AC126014.9


