

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC126014.1 + phase: 0
(405 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
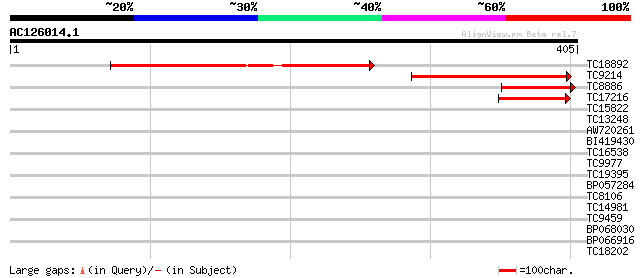
Score E
Sequences producing significant alignments: (bits) Value
TC18892 similar to UP|Q9M9U5 (Q9M9U5) F6A14.15 protein (At1g1874... 253 5e-68
TC9214 161 2e-40
TC8886 similar to UP|Q9M9U5 (Q9M9U5) F6A14.15 protein (At1g18740... 91 5e-19
TC17216 57 6e-09
TC15822 similar to GB|AAC06246.1|2981175|AF053700 deltex {Homo s... 35 0.017
TC13248 29 1.6
AW720261 29 1.6
BI419430 28 2.1
TC16538 25 3.9
TC9977 similar to UP|Q9FNI3 (Q9FNI3) Similarity to RAV2-like DNA... 27 4.7
TC19395 27 4.7
BP057284 27 4.7
TC8106 weakly similar to UP|Q9XIV1 (Q9XIV1) mRNA expressed in cu... 27 6.1
TC14981 similar to UP|AAQ62873 (AAQ62873) At1g76970, partial (11%) 27 6.1
TC9459 similar to AAQ23982 (AAQ23982) Transcription factor Rap21... 27 8.0
BP068030 27 8.0
BP066916 27 8.0
TC18202 weakly similar to UP|Q7PQB7 (Q7PQB7) ENSANGP00000017078 ... 27 8.0
>TC18892 similar to UP|Q9M9U5 (Q9M9U5) F6A14.15 protein
(At1g18740/F6A14_15), partial (35%)
Length = 547
Score = 253 bits (645), Expect = 5e-68
Identities = 129/189 (68%), Positives = 156/189 (82%), Gaps = 1/189 (0%)
Frame = +2
Query: 73 LDCFLICQEEFKAILHTHKGQVVRPPLDRMVSEYFERSVKALDVCNAIRDGVEQIRVWQK 132
LD F+ CQEEF+ ILH HK VV+ PLDRMV ++FERSVKALDVCNAIRDG+EQIR W+K
Sbjct: 2 LDSFICCQEEFRMILHNHKALVVKSPLDRMVGDFFERSVKALDVCNAIRDGIEQIRQWKK 181
Query: 133 LLEIVLYALDHQRSIGEGQFRRAKKALIDLSISMLDDGKESNA-SVAHRNRSFGRSNGGR 191
LLEIVL ALD +++IGEGQFRRAKKAL+DL+I MLDD K+SNA S+AHRNRSFGR+N
Sbjct: 182 LLEIVLCALDQKKNIGEGQFRRAKKALVDLAIGMLDD-KDSNANSIAHRNRSFGRNN--- 349
Query: 192 DRDHHQHGNSHSNNNINTYQHRSMGQFRSLSWSVSRTWSAARQLQAIGNNINPPRANDLM 251
H+Q NSH N++ + GQFRSLSWSVSR WSAARQLQAIGNN++PP+A++++
Sbjct: 350 ---HNQIHNSHHNHHHLLGSSSTFGQFRSLSWSVSRNWSAARQLQAIGNNLSPPKASEIV 520
Query: 252 ATNGLAMSV 260
ATNGLA+ V
Sbjct: 521 ATNGLALVV 547
>TC9214
Length = 587
Score = 161 bits (407), Expect = 2e-40
Identities = 75/114 (65%), Positives = 94/114 (81%)
Frame = +2
Query: 288 HFSIPRSYTWAIPLLLLHERIMEESKKRDRKNACGLLREIQQIEKCVRVMSDLVDSAHFP 347
HFS+PR + WA P+ LH+RI+EES+KR+RKN+CGL+REIQQIEKC R M++L DS FP
Sbjct: 2 HFSVPRQFLWAAPMTSLHDRILEESRKRERKNSCGLMREIQQIEKCARAMNELGDSVQFP 181
Query: 348 LTEEKEGEVRQKVHEVSKVCDALKDGLDPLERQVREVFHRIVRSKTEGLDSLGR 401
L+E+KE EVR +V +V VC+ LK+GLDPLE QVREVF RIV S+ +GLDSLGR
Sbjct: 182 LSEKKEEEVRHRVQDVVSVCEDLKEGLDPLECQVREVFRRIVCSRMDGLDSLGR 343
>TC8886 similar to UP|Q9M9U5 (Q9M9U5) F6A14.15 protein
(At1g18740/F6A14_15), partial (11%)
Length = 604
Score = 90.5 bits (223), Expect = 5e-19
Identities = 42/53 (79%), Positives = 48/53 (90%)
Frame = +2
Query: 352 KEGEVRQKVHEVSKVCDALKDGLDPLERQVREVFHRIVRSKTEGLDSLGRPNN 404
++G R++ E S+VCDALKDGLDPLERQVREVFHRIVRS+TEGLDSLGRPNN
Sbjct: 2 RKGXFRKRXQEXSQVCDALKDGLDPLERQVREVFHRIVRSRTEGLDSLGRPNN 160
>TC17216
Length = 567
Score = 57.0 bits (136), Expect = 6e-09
Identities = 28/51 (54%), Positives = 37/51 (71%)
Frame = +2
Query: 350 EEKEGEVRQKVHEVSKVCDALKDGLDPLERQVREVFHRIVRSKTEGLDSLG 400
EE++ EV Q + + VC+A + GLD LERQVREVF +I+ +TEGLD LG
Sbjct: 2 EEQKMEVEQDLKVLMDVCEAFRGGLDLLERQVREVFRKIMTCRTEGLDYLG 154
>TC15822 similar to GB|AAC06246.1|2981175|AF053700 deltex {Homo sapiens;} ,
partial (3%)
Length = 1104
Score = 35.4 bits (80), Expect = 0.017
Identities = 25/72 (34%), Positives = 42/72 (57%)
Frame = -3
Query: 293 RSYTWAIPLLLLHERIMEESKKRDRKNACGLLREIQQIEKCVRVMSDLVDSAHFPLTEEK 352
R++ AIPLLL +++ + + K + RE+++IE+ R S+L+ A P+ E
Sbjct: 406 RNFVRAIPLLLANKKKVGFHDFTEHKFSACHFRELREIERVERAHSELLVVAG-PIVERG 230
Query: 353 EGEVRQKVHEVS 364
EGEV + HEV+
Sbjct: 229 EGEVIE--HEVA 200
>TC13248
Length = 553
Score = 28.9 bits (63), Expect = 1.6
Identities = 14/31 (45%), Positives = 20/31 (64%)
Frame = -3
Query: 294 SYTWAIPLLLLHERIMEESKKRDRKNACGLL 324
+YTWAI +H I +ES KR+RK+ L+
Sbjct: 326 TYTWAIKFQWIH-NISKESSKRERKHQIHLI 237
>AW720261
Length = 515
Score = 28.9 bits (63), Expect = 1.6
Identities = 16/62 (25%), Positives = 33/62 (52%), Gaps = 4/62 (6%)
Frame = -1
Query: 143 HQRSIGEGQFRRAKKALI----DLSISMLDDGKESNASVAHRNRSFGRSNGGRDRDHHQH 198
HQ+ IG+ + + A++ +S+S D +++ H + S+ +NGGR+ + +
Sbjct: 263 HQK-IGKRELNS*ENAVVVAANGISMSCKDPPPHPVSTLQHHSHSYKLNNGGREEERRRR 87
Query: 199 GN 200
GN
Sbjct: 86 GN 81
>BI419430
Length = 586
Score = 28.5 bits (62), Expect = 2.1
Identities = 16/48 (33%), Positives = 26/48 (53%), Gaps = 9/48 (18%)
Frame = +3
Query: 97 PPLDRMVSEYFERSVKALD----VCNAIRD-----GVEQIRVWQKLLE 135
PP+ VSE+F S ++L +C+ +R G ++ +WQ LLE
Sbjct: 333 PPVSLSVSEFFASSFQSLKERLCICSVLRGFCCMLGTDESMLWQCLLE 476
>TC16538
Length = 693
Score = 25.0 bits (53), Expect(2) = 3.9
Identities = 14/48 (29%), Positives = 23/48 (47%)
Frame = +1
Query: 149 EGQFRRAKKALIDLSISMLDDGKESNASVAHRNRSFGRSNGGRDRDHH 196
+G K+ IDL+ D+ +N ++A + RS NG +HH
Sbjct: 1 QGNGDTVKRGPIDLNAYTEDENVGTNLNLAVQGRSTTPENGSVRIEHH 144
Score = 20.8 bits (42), Expect(2) = 3.9
Identities = 10/29 (34%), Positives = 13/29 (44%)
Frame = +2
Query: 207 INTYQHRSMGQFRSLSWSVSRTWSAARQL 235
+ +YQH S+ L R W AA L
Sbjct: 146 LRSYQHHSLQHRVQLENMDPRDWDAAEAL 232
>TC9977 similar to UP|Q9FNI3 (Q9FNI3) Similarity to RAV2-like DNA-binding
protein, partial (5%)
Length = 468
Score = 27.3 bits (59), Expect = 4.7
Identities = 13/30 (43%), Positives = 18/30 (59%), Gaps = 3/30 (10%)
Frame = +3
Query: 188 NGGRDRDHHQH---GNSHSNNNINTYQHRS 214
N G D +HH + NSHSN+N + H+S
Sbjct: 252 NQGTDTNHHFYQYSSNSHSNSNPHMVPHQS 341
>TC19395
Length = 509
Score = 27.3 bits (59), Expect = 4.7
Identities = 10/25 (40%), Positives = 16/25 (64%)
Frame = -1
Query: 120 IRDGVEQIRVWQKLLEIVLYALDHQ 144
++ +EQ+ +WQ LL +VL HQ
Sbjct: 266 VKPKIEQLIIWQCLLSVVLRTTSHQ 192
>BP057284
Length = 555
Score = 27.3 bits (59), Expect = 4.7
Identities = 15/67 (22%), Positives = 33/67 (48%)
Frame = +2
Query: 110 SVKALDVCNAIRDGVEQIRVWQKLLEIVLYALDHQRSIGEGQFRRAKKALIDLSISMLDD 169
S AL I +E ++ +++ + +DHQ S+G+ ++ ++ ++ +
Sbjct: 68 STTALSPAQDITFRLESVKCLASIIKSMGEWMDHQMSLGDSYLAKSPESCSITESNLTLN 247
Query: 170 GKESNAS 176
G+E NAS
Sbjct: 248 GEEGNAS 268
>TC8106 weakly similar to UP|Q9XIV1 (Q9XIV1) mRNA expressed in cucumber
hypocotyls (Arabinogalactan protein), partial (32%)
Length = 760
Score = 26.9 bits (58), Expect = 6.1
Identities = 15/37 (40%), Positives = 17/37 (45%)
Frame = -2
Query: 178 AHRNRSFGRSNGGRDRDHHQHGNSHSNNNINTYQHRS 214
AHRNRS R N R H NS ++ HRS
Sbjct: 183 AHRNRSRWRGNRNWCRWWRAHRNSRRRCGCRSFGHRS 73
>TC14981 similar to UP|AAQ62873 (AAQ62873) At1g76970, partial (11%)
Length = 677
Score = 26.9 bits (58), Expect = 6.1
Identities = 13/36 (36%), Positives = 21/36 (58%)
Frame = +1
Query: 191 RDRDHHQHGNSHSNNNINTYQHRSMGQFRSLSWSVS 226
R + QHG SHS+N ++ +GQ ++LS + S
Sbjct: 142 RQQFFEQHGGSHSSNGSSSSSDGLVGQTQNLSLNSS 249
>TC9459 similar to AAQ23982 (AAQ23982) Transcription factor Rap212, partial
(12%)
Length = 434
Score = 26.6 bits (57), Expect = 8.0
Identities = 14/61 (22%), Positives = 28/61 (44%)
Frame = +2
Query: 171 KESNASVAHRNRSFGRSNGGRDRDHHQHGNSHSNNNINTYQHRSMGQFRSLSWSVSRTWS 230
KE N + + F + +++ + NS+SN+N Q R+ ++R + W+
Sbjct: 206 KECNIAGCLGCKFFPEEESNNNNNNNNNSNSNSNSNKKKQQKRAKKKYRGVRQRPWGKWA 385
Query: 231 A 231
A
Sbjct: 386 A 388
>BP068030
Length = 413
Score = 26.6 bits (57), Expect = 8.0
Identities = 11/16 (68%), Positives = 14/16 (86%)
Frame = -3
Query: 381 VREVFHRIVRSKTEGL 396
+REV HR+VRS+TE L
Sbjct: 411 IREVVHRLVRSRTEFL 364
>BP066916
Length = 529
Score = 26.6 bits (57), Expect = 8.0
Identities = 13/39 (33%), Positives = 22/39 (56%), Gaps = 6/39 (15%)
Frame = -3
Query: 272 WALVAAIPCQDRGLNVHF------SIPRSYTWAIPLLLL 304
+A ++ P + +G+NVHF +IP TWA L++
Sbjct: 134 FAKISVNPLELKGVNVHFTEQI*SNIPNGVTWATKHLMV 18
>TC18202 weakly similar to UP|Q7PQB7 (Q7PQB7) ENSANGP00000017078 (Fragment),
partial (13%)
Length = 507
Score = 26.6 bits (57), Expect = 8.0
Identities = 10/13 (76%), Positives = 13/13 (99%)
Frame = +3
Query: 380 QVREVFHRIVRSK 392
Q+REVFHR+VRS+
Sbjct: 81 QIREVFHRLVRSQ 119
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.320 0.134 0.395
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 7,208,860
Number of Sequences: 28460
Number of extensions: 101454
Number of successful extensions: 614
Number of sequences better than 10.0: 36
Number of HSP's better than 10.0 without gapping: 609
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 613
length of query: 405
length of database: 4,897,600
effective HSP length: 92
effective length of query: 313
effective length of database: 2,279,280
effective search space: 713414640
effective search space used: 713414640
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 56 (26.2 bits)
Medicago: description of AC126014.1


