

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC126010.6 + phase: 0
(518 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
Score E
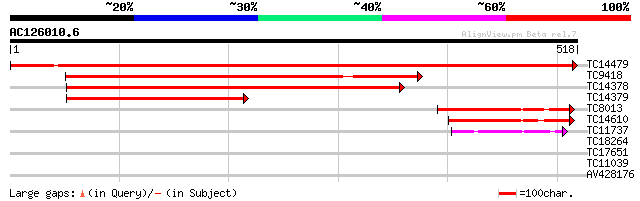
Sequences producing significant alignments: (bits) Value
TC14479 homologue to UP|GLYM_PEA (P34899) Serine hydroxymethyltr... 955 0.0
TC9418 similar to UP|Q94JQ3 (Q94JQ3) AT4g32520/F8B4_220 (Serine... 438 e-124
TC14378 similar to UP|O23254 (O23254) Serine hydroxymethyltransf... 410 e-115
TC14379 homologue to UP|O23254 (O23254) Serine hydroxymethyltran... 237 3e-63
TC8013 homologue to UP|Q8LBY1 (Q8LBY1) Hydroxymethyltransferase,... 96 2e-20
TC14610 similar to UP|O23254 (O23254) Serine hydroxymethyltransf... 92 2e-19
TC11737 48 3e-06
TC18264 similar to UP|Q9SP93 (Q9SP93) Thiol protease, partial (4%) 29 1.6
TC17651 weakly similar to UP|Q8KBL5 (Q8KBL5) Oxidoreductase, sho... 28 2.8
TC11039 weakly similar to UP|O65074 (O65074) YGL010w-like protei... 28 4.7
AV428176 27 6.2
>TC14479 homologue to UP|GLYM_PEA (P34899) Serine hydroxymethyltransferase,
mitochondrial precursor (Serine methylase) (Glycine
hydroxymethyltransferase) (SHMT) , complete
Length = 1874
Score = 955 bits (2468), Expect = 0.0
Identities = 476/518 (91%), Positives = 501/518 (95%)
Frame = +2
Query: 1 MAMAMALRRLSSSINKSSRPLFSASSVYYKSSLPDEAVYDKENSRVSWPKQLNSSLEEID 60
MAMAMALRRLSSSI+K RPLF+ASSVYYKSSLPDEAVY+K + V+WPKQLN+ LE +D
Sbjct: 17 MAMAMALRRLSSSIDKPLRPLFNASSVYYKSSLPDEAVYEK--TTVTWPKQLNAPLEVVD 190
Query: 61 PEIADIIELEKARQWKGLELIPSENFTSLSVMQAVGSIMTNKYSEGYPGARYYGGNEYID 120
PEIADIIE EKARQWKGLELIPSENFTSLSVMQAVGS+MTNKYSEGYPGARYYGGNE+ID
Sbjct: 191 PEIADIIEHEKARQWKGLELIPSENFTSLSVMQAVGSVMTNKYSEGYPGARYYGGNEFID 370
Query: 121 MAETLCQKRALEAFRLDPAKWGVNVQPLSGSPSNFHVYTALLKPHDRIMALDLPHGGHLS 180
MAETLCQKRALEAFRLDPAKWGVNVQPLSGSPSNFHVYTALLKPHDRIMALDLPHGGHLS
Sbjct: 371 MAETLCQKRALEAFRLDPAKWGVNVQPLSGSPSNFHVYTALLKPHDRIMALDLPHGGHLS 550
Query: 181 HGYQTDTKKISAVSIFFETMPYRLDESTGYIDYDQLEKSATLFRPKLIVAGASAYARLYD 240
HGYQTDTKKISAVSIFFETMPYRLDESTGYIDYDQLE SA LFRPKLIVAGASAYARLYD
Sbjct: 551 HGYQTDTKKISAVSIFFETMPYRLDESTGYIDYDQLEASAKLFRPKLIVAGASAYARLYD 730
Query: 241 YARIRKVCDKQKAVMLADMAHISGLVAAGVIPSPFDYADVVTTTTHKSLRGPRGAMIFFR 300
Y RIRKVCDKQKAV+LADMAHISGLVAAGVIPSPFDYADVVTTTTHKSLRGPRGAMIFFR
Sbjct: 731 YDRIRKVCDKQKAVLLADMAHISGLVAAGVIPSPFDYADVVTTTTHKSLRGPRGAMIFFR 910
Query: 301 KGLKEVNKQGKEVFYDYEDKINQAVFPGLQGGPHNHTITGLAVALKQATTPEYKAYQEQV 360
KG+KE+NKQGKEV YDYEDKINQAVFPGLQGGPHNHTITGLAVALKQATTPEYKAYQEQV
Sbjct: 911 KGVKEINKQGKEVLYDYEDKINQAVFPGLQGGPHNHTITGLAVALKQATTPEYKAYQEQV 1090
Query: 361 LSNCAKFAQALSEKGYELVSGGTENHLVLVNLKNKGIDGSRVEKVLEAVHIAANKNTVPG 420
LSNC+KFAQALSE+GYELVSGGTENHLVLVNLKNKGIDGSRVEKVLE+VHIAANKNTVPG
Sbjct: 1091LSNCSKFAQALSERGYELVSGGTENHLVLVNLKNKGIDGSRVEKVLESVHIAANKNTVPG 1270
Query: 421 DVSAMVPGGIRMGTPALTSRGFVEEDFVKVAEYFDASVNLALKIKAESKGTKLKDFVETL 480
DVSAMVPGGIRMGTPALTSRGFVEEDFVKVAE+FDA+VNLALK KAESKGTKLKDF+ T+
Sbjct: 1271DVSAMVPGGIRMGTPALTSRGFVEEDFVKVAEFFDAAVNLALKAKAESKGTKLKDFLATI 1450
Query: 481 QSSSYVQSEISKLRHDVEEFAKQFPTIGFEKSSMKYNK 518
Q SSY Q+EI+KLRHDVEE+AKQFPTIGFEK++MKY +
Sbjct: 1451QESSYFQTEIAKLRHDVEEYAKQFPTIGFEKATMKYKE 1564
>TC9418 similar to UP|Q94JQ3 (Q94JQ3) AT4g32520/F8B4_220 (Serine
hydroxymethyltransferase) (Serine methylase) (Glycine
hydroxymethyltransferase) (SHMT) , partial (60%)
Length = 1231
Score = 438 bits (1127), Expect = e-124
Identities = 217/326 (66%), Positives = 253/326 (77%)
Frame = +3
Query: 52 LNSSLEEIDPEIADIIELEKARQWKGLELIPSENFTSLSVMQAVGSIMTNKYSEGYPGAR 111
L+S L E+DPE+ II EK RQ+K LELI SENFTS +VM+AVGS +TNKYSEG PG R
Sbjct: 276 LDSGLSEVDPEVYAIIGKEKDRQFKSLELIASENFTSRAVMEAVGSCLTNKYSEGLPGKR 455
Query: 112 YYGGNEYIDMAETLCQKRALEAFRLDPAKWGVNVQPLSGSPSNFHVYTALLKPHDRIMAL 171
YYGGNE+ID E CQ+RAL AF +D KWGVNVQPLSGSP+NF VYTALLKPHDRIM L
Sbjct: 456 YYGGNEFIDELEIKCQERALAAFHVDGNKWGVNVQPLSGSPANFAVYTALLKPHDRIMGL 635
Query: 172 DLPHGGHLSHGYQTDTKKISAVSIFFETMPYRLDESTGYIDYDQLEKSATLFRPKLIVAG 231
DLPHGGHLSHG+ T +++SA SI+FE+MPYRLDESTG+IDYD LEK+A LFRPKLI+AG
Sbjct: 636 DLPHGGHLSHGFMTPKRRVSATSIYFESMPYRLDESTGFIDYDMLEKTAILFRPKLIIAG 815
Query: 232 ASAYARLYDYARIRKVCDKQKAVMLADMAHISGLVAAGVIPSPFDYADVVTTTTHKSLRG 291
ASAY R DY R+RK+ D+ A ++ DMAHISGLVAA V+ +PFD+ D+VTTTTHKSLRG
Sbjct: 816 ASAYPRDIDYPRMRKIADQVGAFLMMDMAHISGLVAASVLANPFDFCDIVTTTTHKSLRG 995
Query: 292 PRGAMIFFRKGLKEVNKQGKEVFYDYEDKINQAVFPGLQGGPHNHTITGLAVALKQATTP 351
PRG MIFF+K D E IN AVFPGLQGGPHNHTI GLAV L+ A +P
Sbjct: 996 PRGGMIFFKKDTVH--------GVDLEPAINNAVFPGLQGGPHNHTIGGLAVCLEHAQSP 1151
Query: 352 EYKAYQEQVLSNCAKFAQALSEKGYE 377
E+K YQ QV++NC A L E GY+
Sbjct: 1152EFKVYQNQVVANCRALANRLVELGYK 1229
>TC14378 similar to UP|O23254 (O23254) Serine hydroxymethyltransferase
(Serine methylase) (Glycine hydroxymethyltransferase)
(SHMT) , partial (68%)
Length = 1023
Score = 410 bits (1055), Expect = e-115
Identities = 203/310 (65%), Positives = 243/310 (77%), Gaps = 2/310 (0%)
Frame = +3
Query: 53 NSSLEEIDPEIADIIELEKARQWKGLELIPSENFTSLSVMQAVGSIMTNKYSEGYPGARY 112
N+ L E+D EI D+IE EK RQ +G+ELI SENFTS +V++A+GS +TNKYSEG PG RY
Sbjct: 90 NTPLIEVDGEIHDLIEKEKRRQCRGIELIASENFTSFAVIEALGSALTNKYSEGMPGNRY 269
Query: 113 YGGNEYIDMAETLCQKRALEAFRLDPAKWGVNVQPLSGSPSNFHVYTALLKPHDRIMALD 172
YGGNE+ID E LC+ RAL+AF LD AKWGVNVQP SGSP+NF YTA+L PHDRIM LD
Sbjct: 270 YGGNEFIDQIENLCRSRALQAFHLDAAKWGVNVQPYSGSPANFAAYTAVLNPHDRIMGLD 449
Query: 173 LPHGGHLSHGYQTD-TKKISAVSIFFETMPYRLDESTGYIDYDQLEKSATLFRPKLIVAG 231
LP GGHL+HGY T KKISA SI+FE++PY+++ +TGYIDYD+LE+ A FRPKLI+ G
Sbjct: 450 LPSGGHLTHGYYTSGGKKISATSIYFESLPYKVNSTTGYIDYDRLEEKALDFRPKLIICG 629
Query: 232 ASAYARLYDYARIRKVCDKQKAVMLADMAHISGLVAAGVIPSPFDYADVVTTTTHKSLRG 291
SAY R +DY R R+V DK A++L DMAHISGLVAA + PF+Y D+VTTTTHKSLRG
Sbjct: 630 GSAYPRDWDYRRFREVADKCGALLLCDMAHISGLVAAQEVNDPFEYCDIVTTTTHKSLRG 809
Query: 292 PRGAMIFFRKGLKEVNK-QGKEVFYDYEDKINQAVFPGLQGGPHNHTITGLAVALKQATT 350
PR MIF+RKG K K Q + Y++EDKIN AVFP LQGGPHNH I LAVALKQ+ T
Sbjct: 810 PRAGMIFYRKGPKPAKKGQPEGAVYEFEDKINFAVFPSLQGGPHNHQIGALAVALKQSMT 989
Query: 351 PEYKAYQEQV 360
P +KAY +QV
Sbjct: 990 PGFKAYAKQV 1019
>TC14379 homologue to UP|O23254 (O23254) Serine hydroxymethyltransferase
(Serine methylase) (Glycine hydroxymethyltransferase)
(SHMT) , partial (37%)
Length = 605
Score = 237 bits (605), Expect = 3e-63
Identities = 113/167 (67%), Positives = 137/167 (81%), Gaps = 1/167 (0%)
Frame = +1
Query: 53 NSSLEEIDPEIADIIELEKARQWKGLELIPSENFTSLSVMQAVGSIMTNKYSEGYPGARY 112
N+ L +DPEI D+IE EK RQ +G+ELI SENFTS +V+QA+GS +TNKYSEG PG RY
Sbjct: 103 NTPLSTVDPEIHDLIEHEKRRQCRGIELIASENFTSFAVIQALGSALTNKYSEGMPGNRY 282
Query: 113 YGGNEYIDMAETLCQKRALEAFRLDPAKWGVNVQPLSGSPSNFHVYTALLKPHDRIMALD 172
YGGNE+ID E LC+ RAL+AF LD A WGVNVQP SGSP+NF YTA+L+PHDRIM LD
Sbjct: 283 YGGNEFIDQIENLCRSRALQAFHLDAAAWGVNVQPYSGSPANFAAYTAVLQPHDRIMGLD 462
Query: 173 LPHGGHLSHGYQTD-TKKISAVSIFFETMPYRLDESTGYIDYDQLEK 218
LP GGHL+HGY T KKISA SI+FE++PY+++ STGYIDYD+LE+
Sbjct: 463 LPSGGHLTHGYYTSGGKKISATSIYFESLPYKVNFSTGYIDYDRLEE 603
>TC8013 homologue to UP|Q8LBY1 (Q8LBY1) Hydroxymethyltransferase, partial
(26%)
Length = 690
Score = 95.5 bits (236), Expect = 2e-20
Identities = 53/125 (42%), Positives = 77/125 (61%)
Frame = +2
Query: 392 LKNKGIDGSRVEKVLEAVHIAANKNTVPGDVSAMVPGGIRMGTPALTSRGFVEEDFVKVA 451
L+ G+ ++VEK+ + I NKN V GD SA+ PGG+R+G PA+TSRG VE+DF ++
Sbjct: 2 LRPLGLTXNKVEKLCDLCSITVNKNAVFGDSSALAPGGVRVGAPAMTSRGLVEKDFEQIG 181
Query: 452 EYFDASVNLALKIKAESKGTKLKDFVETLQSSSYVQSEISKLRHDVEEFAKQFPTIGFEK 511
E+ +V L L I+ E G LKDF + L ++ ++ L+ DVE+F+ F GF
Sbjct: 182 EFLHRAVTLTLDIQKE-YGKLLKDFNKGLVNNKALED----LKADVEKFSSSFDMPGFLM 346
Query: 512 SSMKY 516
S MKY
Sbjct: 347 SEMKY 361
>TC14610 similar to UP|O23254 (O23254) Serine hydroxymethyltransferase
(Serine methylase) (Glycine hydroxymethyltransferase)
(SHMT) , partial (24%)
Length = 553
Score = 92.0 bits (227), Expect = 2e-19
Identities = 51/115 (44%), Positives = 74/115 (64%)
Frame = +3
Query: 402 VEKVLEAVHIAANKNTVPGDVSAMVPGGIRMGTPALTSRGFVEEDFVKVAEYFDASVNLA 461
VEK+ + +I NKN V GD SA+ PGG+R+G PA+TSRG VE+DF ++ E+ +V+L
Sbjct: 3 VEKLCDLCNITVNKNAVFGDSSALAPGGVRVGAPAMTSRGLVEKDFEQIGEFLHRAVSLT 182
Query: 462 LKIKAESKGTKLKDFVETLQSSSYVQSEISKLRHDVEEFAKQFPTIGFEKSSMKY 516
L+I+ E G LKDF + L ++ I +L+ DVE+F+ F GF S +KY
Sbjct: 183 LEIQNE-YGKLLKDFNKGLVNN----KAIEELKADVEKFSASFDMPGFLVSELKY 332
>TC11737
Length = 640
Score = 48.1 bits (113), Expect = 3e-06
Identities = 31/106 (29%), Positives = 54/106 (50%)
Frame = +3
Query: 404 KVLEAVHIAANKNTVPGDVSAMVPGGIRMGTPALTSRGFVEEDFVKVAEYFDASVNLALK 463
KV E HI N+ + G +S+ G +R+G+PA+T+RG +E+DF +A++ + S +
Sbjct: 6 KVCETFHITINQCAIYGFISS---GQVRIGSPAMTTRGCLEDDFETIADFLEGS*DYQAL 176
Query: 464 IKAESKGTKLKDFVETLQSSSYVQSEISKLRHDVEEFAKQFPTIGF 509
+ + G + F ++ +S L E F+ QF GF
Sbjct: 177 YRGNT-GNHARSFSRVFRTIKIFRSSEIVLS---ETFSSQFAMPGF 302
>TC18264 similar to UP|Q9SP93 (Q9SP93) Thiol protease, partial (4%)
Length = 570
Score = 29.3 bits (64), Expect = 1.6
Identities = 35/143 (24%), Positives = 59/143 (40%), Gaps = 8/143 (5%)
Frame = -3
Query: 12 SSINKSSRPLFSASSVYYKSSLPDEAVYDKENSRVSWPKQLNSSL---EEIDPEIADIIE 68
+++ RP +S KS +P E K+ + K ++ L EE+ P I + I
Sbjct: 463 AAVTNRFRPRLFSSDETPKSPVPQEPEEIKDVDNKEFKKMIDQYLKGDEEVLPLIQEAIL 284
Query: 69 LEKARQWKGLELIPSENFTSLSVMQAVGSIMTNKYSEGYPGARYYGGNEYID---MAETL 125
+ R+ G + M + + ++ E + A + +E ID A +
Sbjct: 283 I---RRLSGKHDDTDDEMMDELRMGPLDDVSDREFEEDFEEA--HQTDEEIDDLYNARDV 119
Query: 126 CQKRAL--EAFRLDPAKWGVNVQ 146
QKR + E F +DP KW VQ
Sbjct: 118 VQKRMVHDEYFNMDPKKWDEMVQ 50
>TC17651 weakly similar to UP|Q8KBL5 (Q8KBL5) Oxidoreductase, short-chain
dehydrogenase/reductase family, partial (12%)
Length = 599
Score = 28.5 bits (62), Expect = 2.8
Identities = 13/32 (40%), Positives = 20/32 (61%)
Frame = +2
Query: 138 PAKWGVNVQPLSGSPSNFHVYTALLKPHDRIM 169
P +W + PLS PS FHV+ +LL+ H ++
Sbjct: 20 PHQW---LPPLSQPPSTFHVHCSLLRLHHLLL 106
>TC11039 weakly similar to UP|O65074 (O65074) YGL010w-like protein, partial
(79%)
Length = 733
Score = 27.7 bits (60), Expect = 4.7
Identities = 13/28 (46%), Positives = 19/28 (67%)
Frame = +1
Query: 11 SSSINKSSRPLFSASSVYYKSSLPDEAV 38
SSS +SS PL S+S+ ++ SSLP +
Sbjct: 112 SSSFGQSSSPLSSSSTSHHLSSLPSSPI 195
>AV428176
Length = 416
Score = 27.3 bits (59), Expect = 6.2
Identities = 13/44 (29%), Positives = 22/44 (49%)
Frame = +3
Query: 303 LKEVNKQGKEVFYDYEDKINQAVFPGLQGGPHNHTITGLAVALK 346
LK ++K G +D +DK + V PG + P + + G + K
Sbjct: 132 LKCLDKMGVAKLHDLDDKKEEVVLPGFRFHPTDEELVGFYLRRK 263
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.315 0.132 0.373
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 7,258,870
Number of Sequences: 28460
Number of extensions: 86073
Number of successful extensions: 360
Number of sequences better than 10.0: 22
Number of HSP's better than 10.0 without gapping: 353
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 353
length of query: 518
length of database: 4,897,600
effective HSP length: 94
effective length of query: 424
effective length of database: 2,222,360
effective search space: 942280640
effective search space used: 942280640
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.6 bits)
S2: 57 (26.6 bits)
Medicago: description of AC126010.6


