

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
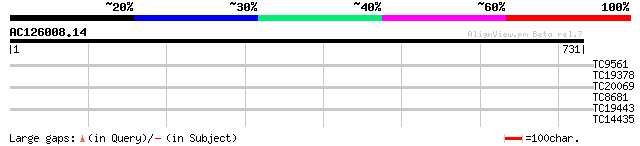
Query= AC126008.14 + phase: 0 /pseudo/partial
(731 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
TC9561 weakly similar to UP|Q7X9J1 (Q7X9J1) Eugenol O-methyltran... 31 0.80
TC19378 weakly similar to UP|O80967 (O80967) At2g39140 protein, ... 28 1.3
TC20069 similar to UP|Q9LKZ4 (Q9LKZ4) Receptor-like protein kina... 29 3.1
TC8681 similar to UP|Q8GVB7 (Q8GVB7) Aux/IAA protein, partial (32%) 28 6.8
TC19443 similar to UP|AAS04935 (AAS04935) NarJ, partial (6%) 28 6.8
TC14435 27 8.9
>TC9561 weakly similar to UP|Q7X9J1 (Q7X9J1) Eugenol O-methyltransferase,
partial (34%)
Length = 718
Score = 30.8 bits (68), Expect = 0.80
Identities = 13/25 (52%), Positives = 18/25 (72%)
Frame = -1
Query: 141 LQLIWYQARVLFRLRRIRCLHQSWV 165
LQ WYQ++V +RL+R LHQ W+
Sbjct: 328 LQGCWYQSQVQYRLQRA*HLHQLWL 254
>TC19378 weakly similar to UP|O80967 (O80967) At2g39140 protein, partial
(17%)
Length = 466
Score = 28.5 bits (62), Expect(2) = 1.3
Identities = 13/31 (41%), Positives = 19/31 (60%)
Frame = -3
Query: 441 RIRRNHLLCIVMLLRWVLVVCLCRIVRL*LM 471
R RR H I++LL +L+ C CR+ L L+
Sbjct: 464 RRRRRHRTSIILLLTLLLLCCWCRLALLVLL 372
Score = 20.0 bits (40), Expect(2) = 1.3
Identities = 5/14 (35%), Positives = 9/14 (63%)
Frame = -2
Query: 477 KCMRRIIRLMIWNW 490
+C R +R+ +W W
Sbjct: 249 RCRRYRVRVWVWVW 208
>TC20069 similar to UP|Q9LKZ4 (Q9LKZ4) Receptor-like protein kinase 3,
partial (14%)
Length = 517
Score = 28.9 bits (63), Expect = 3.1
Identities = 14/31 (45%), Positives = 19/31 (61%)
Frame = -1
Query: 602 QAIFWKRSRRVRRKIWNSWTE*RWLIKVKVE 632
+ + WKR RR RR+I+ SW W+ K VE
Sbjct: 112 ERVKWKRKRRSRRRIFASWF---WVWKKGVE 29
>TC8681 similar to UP|Q8GVB7 (Q8GVB7) Aux/IAA protein, partial (32%)
Length = 732
Score = 27.7 bits (60), Expect = 6.8
Identities = 17/40 (42%), Positives = 23/40 (57%), Gaps = 2/40 (5%)
Frame = +3
Query: 333 CVPSCLSVSFG*RK*VSL--VM*FLRVVFLWILPRLMQCC 370
C+PS LSV F R V++ FL V+LWIL + + C
Sbjct: 474 CLPSSLSVEFNWRDPVNIYTFWLFLCSVYLWILVAV*KIC 593
>TC19443 similar to UP|AAS04935 (AAS04935) NarJ, partial (6%)
Length = 533
Score = 27.7 bits (60), Expect = 6.8
Identities = 12/24 (50%), Positives = 15/24 (62%)
Frame = +2
Query: 440 CRIRRNHLLCIVMLLRWVLVVCLC 463
CR R HLLC+ + LR V + C C
Sbjct: 266 CRSERAHLLCLTVFLR-VALPCFC 334
>TC14435
Length = 866
Score = 27.3 bits (59), Expect = 8.9
Identities = 12/31 (38%), Positives = 18/31 (57%), Gaps = 3/31 (9%)
Frame = +2
Query: 488 WNWQPLCI---H*RFGGIICMVLSLRCSVTT 515
WNW+PLC H*+ +I +L L +T+
Sbjct: 413 WNWKPLCTVM*H*KKSYVILDILHLMVQITS 505
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.376 0.169 0.715
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 17,033,179
Number of Sequences: 28460
Number of extensions: 309056
Number of successful extensions: 6005
Number of sequences better than 10.0: 12
Number of HSP's better than 10.0 without gapping: 2491
Number of HSP's successfully gapped in prelim test: 249
Number of HSP's that attempted gapping in prelim test: 3299
Number of HSP's gapped (non-prelim): 3104
length of query: 731
length of database: 4,897,600
effective HSP length: 97
effective length of query: 634
effective length of database: 2,136,980
effective search space: 1354845320
effective search space used: 1354845320
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 13 ( 7.1 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 35 (21.5 bits)
S2: 59 (27.3 bits)
Medicago: description of AC126008.14


