

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC126007.11 + phase: 0
(525 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
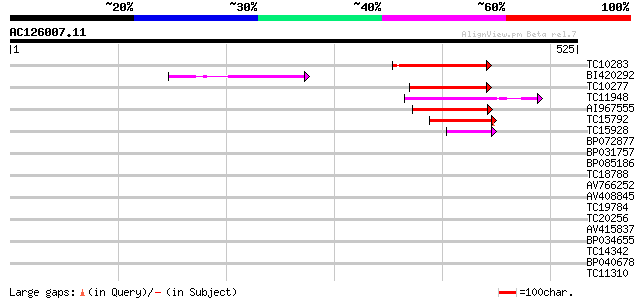
Score E
Sequences producing significant alignments: (bits) Value
TC10283 weakly similar to UP|AAR14274 (AAR14274) Predicted prote... 110 6e-25
BI420292 109 1e-24
TC10277 weakly similar to GB|AAP68251.1|31711790|BT008812 At2g36... 84 4e-17
TC11948 73 1e-13
AI967555 70 6e-13
TC15792 weakly similar to UP|AAR14274 (AAR14274) Predicted prote... 69 2e-12
TC15928 42 2e-04
BP072877 37 0.010
BP031757 36 0.014
BP085186 36 0.014
TC18788 weakly similar to UP|Q9SSA6 (Q9SSA6) F4P13.1 protein, pa... 35 0.030
AV766252 35 0.039
AV408845 32 0.20
TC19784 weakly similar to UP|Q8S9L3 (Q8S9L3) AT4g38220/F20D10_34... 32 0.33
TC20256 similar to UP|CYB_LAMGT (Q94T68) Cytochrome b, partial (6%) 30 0.74
AV415837 30 0.74
BP034655 30 1.3
TC14342 weakly similar to UP|Q8LEJ6 (Q8LEJ6) Arabinogalactan pro... 29 2.2
BP040678 29 2.2
TC11310 similar to UP|BAD11331 (BAD11331) BRI1-KD interacting pr... 29 2.2
>TC10283 weakly similar to UP|AAR14274 (AAR14274) Predicted protein, partial
(10%)
Length = 678
Score = 110 bits (275), Expect = 6e-25
Identities = 53/92 (57%), Positives = 72/92 (77%)
Frame = +2
Query: 355 RKIKDKYYFGGMYCAVLIVNQVVVSAGIFRVFGKEVAELSLIATKAEYQKQGFFKCLLSC 414
+K+KD + F GMYCAVL +N +VV A +FRVFG+EVAEL L+AT+ +YQ++G+F+ L SC
Sbjct: 2 KKVKD-HDFRGMYCAVLTINPMVVCACLFRVFGQEVAELPLVATRKDYQRKGYFRSLFSC 178
Query: 415 IENVLKELKVERLVLPAAHEAESMWIDKFGFT 446
IE +L LKV+ VLPA EA+S+W KFGF+
Sbjct: 179 IERMLGCLKVKNFVLPATREAKSLWTTKFGFS 274
>BI420292
Length = 553
Score = 109 bits (273), Expect = 1e-24
Identities = 54/130 (41%), Positives = 71/130 (54%)
Frame = +2
Query: 148 PKEKWYCEYCRNKLQKDKNVEHKENVVTTQKIIESDPSEQIAKICTLSVKHKEVEHSSCA 207
P WYC+YC N+ Q++K+V+ + ++E D E CA
Sbjct: 230 PSGLWYCKYCENETQENKDVDQGRSF-----LVEHD------------------EGGQCA 340
Query: 208 LCSERHFNNGEFSPWTVMICDQCEKDYHVGCLKDHNMANLKKVPKHYWFCGVDCYDIHMK 267
LC F F P TVMICDQCEK+YHV CLKDHN NL+K+P+ WFC DC IH
Sbjct: 341 LCGGHDFIKDVFGPRTVMICDQCEKEYHVECLKDHNKQNLEKLPEGKWFCCSDCDHIHNV 520
Query: 268 LKNFMARGDV 277
L ++ARG++
Sbjct: 521 LVKYVARGEM 550
>TC10277 weakly similar to GB|AAP68251.1|31711790|BT008812 At2g36720
{Arabidopsis thaliana;}, partial (11%)
Length = 600
Score = 84.3 bits (207), Expect = 4e-17
Identities = 41/76 (53%), Positives = 54/76 (70%)
Frame = +3
Query: 371 LIVNQVVVSAGIFRVFGKEVAELSLIATKAEYQKQGFFKCLLSCIENVLKELKVERLVLP 430
L+VN V+SA + R+FG +VAEL L+AT +G+F L SCIE +L + V+ LVLP
Sbjct: 3 LVVNSSVLSAXMLRIFGSDVAELPLVATSNGNHGKGYFPTLFSCIERLLAFMNVKSLVLP 182
Query: 431 AAHEAESMWIDKFGFT 446
AA EAES+W DKFGF+
Sbjct: 183 AAEEAESIWTDKFGFS 230
>TC11948
Length = 591
Score = 73.2 bits (178), Expect = 1e-13
Identities = 41/128 (32%), Positives = 69/128 (53%)
Frame = +1
Query: 366 MYCAVLIVNQVVVSAGIFRVFGKEVAELSLIATKAEYQKQGFFKCLLSCIENVLKELKVE 425
M CAVL+++ V S +FR+ G +VAE+ L A+ +++ Q F+CL I N L LKV
Sbjct: 4 MLCAVLLLDGTVASTALFRLIGPKVAEMPLAASSFDFEGQAVFRCLFDMIVNQLTSLKVT 183
Query: 426 RLVLPAAHEAESMWIDKFGFTEPNQGLGRRYYRRSWSFHLNKGVEQNTHLTYVHALYLMY 485
L++P+ + W D++GF+ ++ + +YR S+ ++T LM+
Sbjct: 184 TLIIPSCEITKEFWRDRYGFSIVSEE-EKLFYRESY------------NMTEFGETVLMH 324
Query: 486 RKEIDSCF 493
+K DS F
Sbjct: 325 KKL*DSAF 348
>AI967555
Length = 442
Score = 70.5 bits (171), Expect = 6e-13
Identities = 33/74 (44%), Positives = 47/74 (62%)
Frame = +3
Query: 374 NQVVVSAGIFRVFGKEVAELSLIATKAEYQKQGFFKCLLSCIENVLKELKVERLVLPAAH 433
N V+S R+FG++VAE+ L+AT+ +Y++ G + L+ +E LKEL VERLVLPA
Sbjct: 120 NDEVISVATIRIFGQKVAEMPLVATRIQYRRHGMCRVLMDELEKKLKELGVERLVLPAVP 299
Query: 434 EAESMWIDKFGFTE 447
W + FGF E
Sbjct: 300 GVVETWTNSFGFVE 341
>TC15792 weakly similar to UP|AAR14274 (AAR14274) Predicted protein, partial
(7%)
Length = 567
Score = 68.6 bits (166), Expect = 2e-12
Identities = 34/64 (53%), Positives = 46/64 (71%), Gaps = 2/64 (3%)
Frame = +3
Query: 389 EVAELSLIATKAEYQKQGFFKCLLSCIENVLKELKVERLVLPAAHEAESMWIDKFGFT-- 446
++A L +AT +Y+ +G+F L SCIE +L L V+ LVLPAA EAES+WI KFGF+
Sbjct: 3 DIAVLPFLATSNKYRGKGYFPTLFSCIERLLSFLSVKNLVLPAAEEAESIWIHKFGFSKM 182
Query: 447 EPNQ 450
EP+Q
Sbjct: 183 EPDQ 194
>TC15928
Length = 521
Score = 42.0 bits (97), Expect = 2e-04
Identities = 20/46 (43%), Positives = 27/46 (58%)
Frame = +2
Query: 405 QGFFKCLLSCIENVLKELKVERLVLPAAHEAESMWIDKFGFTEPNQ 450
QG + L+ IE+ L LKV+ LV+P+ E MW KF F E N+
Sbjct: 5 QGICRSLMKEIESFLCSLKVKTLVIPSVPETVEMWTQKFNFNEVNE 142
>BP072877
Length = 433
Score = 36.6 bits (83), Expect = 0.010
Identities = 18/43 (41%), Positives = 26/43 (59%)
Frame = -3
Query: 62 TSNSHASPTTINYRDQCLHKLVFQENVLEDGAAVGYFVYEEVS 104
TS ++ S I + Q +HKL+F+EN L DGA V Y + +S
Sbjct: 140 TSTTNKSEWRIAKKYQRMHKLIFEENGLPDGAEVAYMHVDRIS 12
>BP031757
Length = 507
Score = 36.2 bits (82), Expect = 0.014
Identities = 15/38 (39%), Positives = 23/38 (60%)
Frame = +2
Query: 19 KDGDDVEKKMKNKKKKKNKNKKESNLRSNESKNLQPSP 56
KDG+D E++ + KKKKK++ KKE + + P P
Sbjct: 182 KDGEDYEEEEEKKKKKKSRKKKEEEEEEEQEEPESPQP 295
>BP085186
Length = 388
Score = 36.2 bits (82), Expect = 0.014
Identities = 18/52 (34%), Positives = 26/52 (49%)
Frame = -2
Query: 14 VEAVAKDGDDVEKKMKNKKKKKNKNKKESNLRSNESKNLQPSPKHTTQTSNS 65
+ + K+ +KK K KKKKK K KK+ N R K +PK + + S
Sbjct: 282 ITRMGKEMKKKKKKKKKKKKKKKKKKKKQNQRKKRMKRKMSTPKKEKEKTPS 127
>TC18788 weakly similar to UP|Q9SSA6 (Q9SSA6) F4P13.1 protein, partial (7%)
Length = 951
Score = 35.0 bits (79), Expect = 0.030
Identities = 15/56 (26%), Positives = 28/56 (49%)
Frame = +1
Query: 223 TVMICDQCEKDYHVGCLKDHNMANLKKVPKHYWFCGVDCYDIHMKLKNFMARGDVL 278
+V++CD C+ +YH CL L ++P+ W+C C D +N + ++
Sbjct: 148 SVLLCDTCDAEYHTYCLN----PPLARIPEGNWYC-PSCVDGKHATQNVIEHSQII 300
>AV766252
Length = 585
Score = 34.7 bits (78), Expect = 0.039
Identities = 29/105 (27%), Positives = 48/105 (45%), Gaps = 5/105 (4%)
Frame = +2
Query: 29 KNKKKKKNKNKKESNLRSNESKNLQPSPKHTTQTSNSHASPTTIN-YRDQCLHKLVFQEN 87
+ KKK +NKN+KE L + + + K +T+N AS + + ++ L K VF
Sbjct: 113 QRKKK*ENKNRKE*QLNKRKKRKAK*QRKGGKKTNNKVASKISFS*WKRTQLLKSVF*LL 292
Query: 88 VLEDGAAVGYFVYEEVSPSKFEAHAGWA----SRRKPWPLIIEFL 128
+ ED V YF S S + W+ S+ P ++ F+
Sbjct: 293 LSEDAGDVQYFFTANKSFSLSRGPSSWSTI*LSQSMPLTMLFTFI 427
>AV408845
Length = 429
Score = 32.3 bits (72), Expect = 0.20
Identities = 14/26 (53%), Positives = 20/26 (76%)
Frame = -1
Query: 19 KDGDDVEKKMKNKKKKKNKNKKESNL 44
+ G +VEK+ K KKKKK K KK+++L
Sbjct: 162 RKG*EVEKRKKKKKKKKKKKKKKASL 85
>TC19784 weakly similar to UP|Q8S9L3 (Q8S9L3) AT4g38220/F20D10_340, partial
(21%)
Length = 415
Score = 31.6 bits (70), Expect = 0.33
Identities = 16/38 (42%), Positives = 22/38 (57%)
Frame = +2
Query: 31 KKKKKNKNKKESNLRSNESKNLQPSPKHTTQTSNSHAS 68
KKKKK K K+ S + SN SK+ + H T+ S +S
Sbjct: 110 KKKKKKKKKQSSLVSSNTSKSKLTNQTHVTKKHQSFSS 223
>TC20256 similar to UP|CYB_LAMGT (Q94T68) Cytochrome b, partial (6%)
Length = 543
Score = 30.4 bits (67), Expect = 0.74
Identities = 14/34 (41%), Positives = 19/34 (55%)
Frame = -1
Query: 20 DGDDVEKKMKNKKKKKNKNKKESNLRSNESKNLQ 53
D + +K KN KKKK K K ESN + +S +
Sbjct: 540 DSKEDDKTDKNNKKKKKKKKAESNAKKPKSDRFE 439
>AV415837
Length = 422
Score = 30.4 bits (67), Expect = 0.74
Identities = 20/50 (40%), Positives = 30/50 (60%), Gaps = 3/50 (6%)
Frame = -1
Query: 26 KKMKNKKKKKN--KNKKESNLRSNESKNLQPS-PKHTTQTSNSHASPTTI 72
K+ KN KK KN + K +++NLQ + PK+TT++ +S ASP I
Sbjct: 371 KQSKNTKKGKNA*ECMKHYMRHKEKTQNLQRNDPKNTTKSGSSAASPGAI 222
>BP034655
Length = 517
Score = 29.6 bits (65), Expect = 1.3
Identities = 14/34 (41%), Positives = 19/34 (55%)
Frame = +2
Query: 25 EKKMKNKKKKKNKNKKESNLRSNESKNLQPSPKH 58
+KK K KKKKK K KK++ ++K P H
Sbjct: 170 KKKKKKKKKKKKKKKKKTKRMMKKTKMKTTFPLH 271
>TC14342 weakly similar to UP|Q8LEJ6 (Q8LEJ6) Arabinogalactan protein-like,
partial (27%)
Length = 724
Score = 28.9 bits (63), Expect = 2.2
Identities = 21/63 (33%), Positives = 28/63 (44%), Gaps = 3/63 (4%)
Frame = -3
Query: 26 KKMKNKKKKKNKNKKESNLRSNESKNLQPSPKHTTQTSNSHASPTTINYRD---QCLHKL 82
K K K K KNK K E N + S NL P ++ ++N T +RD Q +H
Sbjct: 446 KNNKIKIKLKNKRK*EKNETAMPS*NLHPQTRNNKGSNNCTKVGDTDTHRDWKYQLMHPT 267
Query: 83 VFQ 85
Q
Sbjct: 266 TTQ 258
>BP040678
Length = 587
Score = 28.9 bits (63), Expect = 2.2
Identities = 14/33 (42%), Positives = 18/33 (54%), Gaps = 4/33 (12%)
Frame = +1
Query: 199 KEVEHSSCALCSER--HFNNGEFSPWTV--MIC 227
KE + C C R HF+N FSPW + +IC
Sbjct: 394 KE*HNKKCPCCETRRTHFSNSTFSPWCLGSVIC 492
>TC11310 similar to UP|BAD11331 (BAD11331) BRI1-KD interacting protein 102
(Fragment), partial (13%)
Length = 667
Score = 28.9 bits (63), Expect = 2.2
Identities = 17/55 (30%), Positives = 24/55 (42%)
Frame = +1
Query: 10 DNIVVEAVAKDGDDVEKKMKNKKKKKNKNKKESNLRSNESKNLQPSPKHTTQTSN 64
D +V E K + EKK K KKKKK++ E E + KH + +
Sbjct: 34 DVVVDEGKVKVEVEGEKKEKKKKKKKDQENGEVASSDEEKAEKEKKKKHKVKVED 198
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.319 0.135 0.406
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 9,964,308
Number of Sequences: 28460
Number of extensions: 149614
Number of successful extensions: 1046
Number of sequences better than 10.0: 66
Number of HSP's better than 10.0 without gapping: 1008
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 1036
length of query: 525
length of database: 4,897,600
effective HSP length: 94
effective length of query: 431
effective length of database: 2,222,360
effective search space: 957837160
effective search space used: 957837160
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 57 (26.6 bits)
Medicago: description of AC126007.11


