

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC126006.11 + phase: 0
(325 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
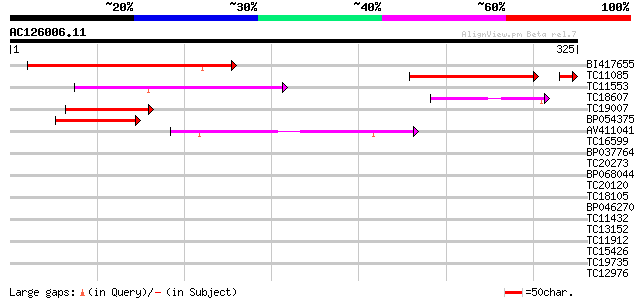
BI417655 187 2e-48
TC11085 similar to GB|AAP13430.1|30023794|BT006322 At3g54390 {Ar... 115 3e-27
TC11553 weakly similar to UP|BAD10712 (BAD10712) Proline-rich pr... 61 3e-10
TC18607 similar to UP|O80512 (O80512) At2g44730 protein, partial... 45 2e-05
TC19007 similar to UP|Q8W3M8 (Q8W3M8) 6b-interacting protein 1, ... 45 2e-05
BP054375 41 3e-04
AV411041 40 5e-04
TC16599 similar to UP|Q8W239 (Q8W239) GT-2 factor (Fragment), pa... 33 0.086
BP037764 30 0.43
TC20273 similar to UP|REMO_SOLTU (P93788) Remorin (pp34), partia... 30 0.43
BP068044 28 1.6
TC20120 weakly similar to UP|Q946Y3 (Q946Y3) SWITCH1 splice vari... 28 2.1
TC18105 UP|Q93WR8 (Q93WR8) MEK map kinase kinsae, partial (5%) 27 3.6
BP046270 27 4.7
TC11432 weakly similar to UP|Q9LQY3 (Q9LQY3) T24P13.7, partial (... 27 4.7
TC13152 similar to UP|Q84XX6 (Q84XX6) Leunig (Fragment), partial... 27 4.7
TC11912 27 6.1
TC15426 weakly similar to UP|HS7E_DROME (P29845) Heat shock 70 k... 27 6.1
TC19735 similar to UP|Q8NJM5 (Q8NJM5) Probable ATP-dependent RNA... 27 6.1
TC12976 similar to UP|O02448 (O02448) Lox22-otx (Fragment), part... 27 6.1
>BI417655
Length = 509
Score = 187 bits (475), Expect = 2e-48
Identities = 95/123 (77%), Positives = 106/123 (85%), Gaps = 3/123 (2%)
Frame = +2
Query: 11 IQQNQESPRK-LQNNIIGSGNERLKRDEWSEGAVTTLLEAYESKWVLRNRAKLKGQDWED 69
I +ESPRK + G G +RLKRDEWSEGAV TLLEAYE+KWVLRNRAKLKGQDWE+
Sbjct: 65 ITAKEESPRKAIGGGGSGGGGDRLKRDEWSEGAVCTLLEAYEAKWVLRNRAKLKGQDWEE 244
Query: 70 VAKHVSSRSNSTKSPKTQTQCKNKIESMKKRYRSESASSD--VSSWPLYSRLDLLLRGTG 127
VAKHVSSR+N+TKSPKTQTQCKNKIESMKKRYRSESA++ SSWPLYSRLD+LLRGTG
Sbjct: 245 VAKHVSSRANNTKSPKTQTQCKNKIESMKKRYRSESATATDAASSWPLYSRLDVLLRGTG 424
Query: 128 QLS 130
+S
Sbjct: 425 PVS 433
>TC11085 similar to GB|AAP13430.1|30023794|BT006322 At3g54390 {Arabidopsis
thaliana;}, partial (25%)
Length = 585
Score = 115 bits (289), Expect(2) = 3e-27
Identities = 58/74 (78%), Positives = 70/74 (94%)
Frame = +3
Query: 230 NNNKRQKKETMGIAESIRWVAEVVVRSEQTRMETLKEIEKMRVEAEAKRGEMEIKRTEII 289
N+++R++KE IAESIRW+AEVVVRSE TRM+T+KEIEKMRVEAEAKRGEM++KRTEII
Sbjct: 72 NHDQRRRKEDSEIAESIRWLAEVVVRSEPTRMDTMKEIEKMRVEAEAKRGEMDLKRTEII 251
Query: 290 ANTQLEIARIFANV 303
A+TQLEIARIFA+V
Sbjct: 252 AHTQLEIARIFASV 293
Score = 22.3 bits (46), Expect(2) = 3e-27
Identities = 10/10 (100%), Positives = 10/10 (100%)
Frame = +1
Query: 316 VDSSLRIGRS 325
VDSSLRIGRS
Sbjct: 304 VDSSLRIGRS 333
>TC11553 weakly similar to UP|BAD10712 (BAD10712) Proline-rich protein
family-like, partial (27%)
Length = 493
Score = 60.8 bits (146), Expect = 3e-10
Identities = 34/124 (27%), Positives = 59/124 (47%), Gaps = 2/124 (1%)
Frame = -1
Query: 38 WSEGAVTTLLEAYESKWVLRNRAKLKGQDWEDVAKHVSSRS--NSTKSPKTQTQCKNKIE 95
W+ L+ AY+ KW R L+ WE+VA V++R + K+ QC++KI+
Sbjct: 421 WTHQETVNLIRAYQEKWYSLKRGPLRSNQWEEVAVVVAARCGYDFAHPSKSALQCRHKID 242
Query: 96 SMKKRYRSESASSDVSSWPLYSRLDLLLRGTGQLSTTTLNNNQAGLVLLEPSSLVMSHAQ 155
+++R+R+E W +S +D L RG +S L + G+ E + +
Sbjct: 241 KLRQRHRAEKHRPATRPWQYFSLMDDLERGPLPISAHPLASLPPGVDEDEEDDEIEDGGE 62
Query: 156 EEVH 159
+VH
Sbjct: 61 GDVH 50
>TC18607 similar to UP|O80512 (O80512) At2g44730 protein, partial (16%)
Length = 516
Score = 45.1 bits (105), Expect = 2e-05
Identities = 28/71 (39%), Positives = 39/71 (54%), Gaps = 3/71 (4%)
Frame = +2
Query: 242 IAESIRWVAEVVVRSEQTRMETLKEIEKMRVEAEAKRGEMEIKRTEIIANTQLEIARIFA 301
+ +I+ + + VR EQ +ME KEIE MR+E E+KRTE+I +Q I FA
Sbjct: 23 VVSAIKVLGDGFVRMEQMKMEMAKEIEMMRIET-------EMKRTEMILESQQRIVESFA 181
Query: 302 NV---NNKDKD 309
NNK K+
Sbjct: 182 RAISENNKKKN 214
>TC19007 similar to UP|Q8W3M8 (Q8W3M8) 6b-interacting protein 1, partial
(19%)
Length = 427
Score = 44.7 bits (104), Expect = 2e-05
Identities = 20/50 (40%), Positives = 34/50 (68%)
Frame = +1
Query: 33 LKRDEWSEGAVTTLLEAYESKWVLRNRAKLKGQDWEDVAKHVSSRSNSTK 82
++ D WSE A +TL++A+ S+++ NR L+ +DW+DVA V++ TK
Sbjct: 253 VREDCWSEEASSTLVDAWGSRYLELNRGNLRQKDWQDVADAVNALHGHTK 402
>BP054375
Length = 418
Score = 40.8 bits (94), Expect = 3e-04
Identities = 19/49 (38%), Positives = 30/49 (60%)
Frame = +1
Query: 27 GSGNERLKRDEWSEGAVTTLLEAYESKWVLRNRAKLKGQDWEDVAKHVS 75
G G + D WSEGA L+EA+ +++ +R LK + W++VA+ VS
Sbjct: 268 GGGGGGGREDCWSEGATAVLIEAWGERYLELSRGNLKQKHWKEVAEIVS 414
>AV411041
Length = 428
Score = 40.0 bits (92), Expect = 5e-04
Identities = 36/154 (23%), Positives = 64/154 (41%), Gaps = 12/154 (7%)
Frame = +1
Query: 93 KIESMKKRYRSESAS------SDVSSWPLYSRLDLLLRGTGQLSTTTLNNNQAGLVLLEP 146
+I+++KK+Y+SE A S WP Y RLD L+ T ++ N+ Q L L P
Sbjct: 1 RIDTVKKKYKSEKAKISAAGGGTTSKWPFYDRLDQLIGPTAKIPAGNSNSQQQKLPLGIP 180
Query: 147 SSLVMSHAQEEVHQPLVPPPITTAQNSHGSNGVDKLIKED-EIETKSSDHIANKNPLETD 205
+ ++ NS + + +ED E++ + + P+++D
Sbjct: 181 VGVRSGGG------------ASSQYNSQKQHQQQQEQEEDVELKNQKMQYRRRPPPVDSD 324
Query: 206 SS-----TPALYSEKDEQRSKKRRMRTENNNNKR 234
SS +P R +++R R N+ R
Sbjct: 325 SSEREVVSPVSSDSYPPGRYERKRARVMNSRGGR 426
>TC16599 similar to UP|Q8W239 (Q8W239) GT-2 factor (Fragment), partial (37%)
Length = 693
Score = 32.7 bits (73), Expect = 0.086
Identities = 23/97 (23%), Positives = 42/97 (42%)
Frame = +1
Query: 26 IGSGNERLKRDEWSEGAVTTLLEAYESKWVLRNRAKLKGQDWEDVAKHVSSRSNSTKSPK 85
IGSG R R E LL+ + + LKG WE+V++ ++ + K
Sbjct: 181 IGSGGSRWPRQE-----TLALLKIRSDMDAVFRDSSLKGPLWEEVSRKLAGLGYHRSAKK 345
Query: 86 TQTQCKNKIESMKKRYRSESASSDVSSWPLYSRLDLL 122
+ + +N + K+ S SA + ++ + +L L
Sbjct: 346 CKEKFENVYKYHKRTKESRSAKPEGKTYRFFDQLQAL 456
>BP037764
Length = 519
Score = 30.4 bits (67), Expect = 0.43
Identities = 28/78 (35%), Positives = 37/78 (46%), Gaps = 1/78 (1%)
Frame = -1
Query: 169 TAQNSHGSNGVDKLIKEDEIETKSSDHIANKNPLETDSSTPALYS-EKDEQRSKKRRMRT 227
T QN +GS LI ET+S ET +ST + E E+ SKK++M+T
Sbjct: 252 TEQNQNGSG----LISSV*FETQSL--------WETPNST*RVEEGEGGERESKKKKMKT 109
Query: 228 ENNNNKRQKKETMGIAES 245
E K Q E G+A S
Sbjct: 108 ETEIPKPQPFEAKGVAAS 55
>TC20273 similar to UP|REMO_SOLTU (P93788) Remorin (pp34), partial (64%)
Length = 569
Score = 30.4 bits (67), Expect = 0.43
Identities = 37/181 (20%), Positives = 67/181 (36%), Gaps = 31/181 (17%)
Frame = +1
Query: 141 LVLLEPSSLV-MSHAQEEVHQPLVPPPITTAQNSHGSNGVDKLIKEDEIETKSSDHIANK 199
L+ +E S L M+ Q++ P P+ K + ++K+ + K
Sbjct: 25 LL*IEHSKLASMAELQKKAESDTTPAPVAAVPPPSNDAVAKKAPEVAADDSKALVVVPEK 204
Query: 200 NPLETDSSTP--------ALYSEKDEQRSKKRRMRTENNNNKRQKKETMGIAESIRW--- 248
P+ + + AL + E+R + E+ +K + K +++ + W
Sbjct: 205 TPVPENKPSSKGSLDRDVALAELEKEKRLSYVKAWEESEKSKTENKAQKNLSDVVAWENS 384
Query: 249 ---VAEVVVRSEQTRMETLK----------------EIEKMRVEAEAKRGEMEIKRTEII 289
E +R + R+E K E E+ R EAKRGE +K E+
Sbjct: 385 KKAALEAQLRKIEERLEKKKAEYGEKMKNKIALVHKEAEERRAMIEAKRGEDLLKAEELA 564
Query: 290 A 290
A
Sbjct: 565 A 567
>BP068044
Length = 266
Score = 28.5 bits (62), Expect = 1.6
Identities = 13/37 (35%), Positives = 24/37 (64%)
Frame = +1
Query: 214 EKDEQRSKKRRMRTENNNNKRQKKETMGIAESIRWVA 250
+K EQ +KK + T+ + KRQKKE ++I++++
Sbjct: 85 KKFEQGTKKTAIATDPXHRKRQKKERTCYHQTIKYIS 195
>TC20120 weakly similar to UP|Q946Y3 (Q946Y3) SWITCH1 splice variant S,
partial (9%)
Length = 557
Score = 28.1 bits (61), Expect = 2.1
Identities = 25/101 (24%), Positives = 39/101 (37%), Gaps = 1/101 (0%)
Frame = -3
Query: 167 ITTAQNSHGSNGVDKLIKEDEIETKSSDHIANKNPLETDSSTPALYSEKDEQRSKKRRMR 226
I AQN G + + E+E + +++ P+ P K K R ++
Sbjct: 396 IFIAQNGEGGSEPPDISNEEE--SNGEENLLKLEPI----FYPEGRKRKSRSHGKLREVK 235
Query: 227 TEN-NNNKRQKKETMGIAESIRWVAEVVVRSEQTRMETLKE 266
E K++T RW AE +EQ+ E LKE
Sbjct: 234 EEPCGRQSTSKRKTKKHESKERWSAERYSLAEQSMWEVLKE 112
>TC18105 UP|Q93WR8 (Q93WR8) MEK map kinase kinsae, partial (5%)
Length = 550
Score = 27.3 bits (59), Expect = 3.6
Identities = 11/45 (24%), Positives = 23/45 (50%)
Frame = +2
Query: 151 MSHAQEEVHQPLVPPPITTAQNSHGSNGVDKLIKEDEIETKSSDH 195
+ H + E+ +PPP T+A S G+ + + ++ + + DH
Sbjct: 416 LRHKRNEMRPIQLPPPSTSATGSPGAAANNNINRDQRPQRRRKDH 550
>BP046270
Length = 549
Score = 26.9 bits (58), Expect = 4.7
Identities = 12/30 (40%), Positives = 19/30 (63%), Gaps = 3/30 (10%)
Frame = +3
Query: 51 ESKWVLRNR---AKLKGQDWEDVAKHVSSR 77
+S W L++ A + GQ W++V KH+ SR
Sbjct: 174 DSCWKLKSFFCFASVSGQQWKEVLKHLESR 263
>TC11432 weakly similar to UP|Q9LQY3 (Q9LQY3) T24P13.7, partial (89%)
Length = 858
Score = 26.9 bits (58), Expect = 4.7
Identities = 14/37 (37%), Positives = 21/37 (55%)
Frame = -1
Query: 105 SASSDVSSWPLYSRLDLLLRGTGQLSTTTLNNNQAGL 141
S+SS S+WP + LDL L T + ++ N AG+
Sbjct: 399 SSSSITSTWPFLATLDLSLAATPVFHSKSIVNVIAGV 289
>TC13152 similar to UP|Q84XX6 (Q84XX6) Leunig (Fragment), partial (60%)
Length = 453
Score = 26.9 bits (58), Expect = 4.7
Identities = 14/38 (36%), Positives = 22/38 (57%)
Frame = -1
Query: 252 VVVRSEQTRMETLKEIEKMRVEAEAKRGEMEIKRTEII 289
V VRSE + T + + + E A+RG + +KRT +I
Sbjct: 207 VKVRSEYPGLSTSQTLTVLSNEDVARRGMLGLKRTSVI 94
>TC11912
Length = 505
Score = 26.6 bits (57), Expect = 6.1
Identities = 13/27 (48%), Positives = 17/27 (62%)
Frame = +2
Query: 133 TLNNNQAGLVLLEPSSLVMSHAQEEVH 159
TL +N + L PSSL S +QE+VH
Sbjct: 101 TLASNTTAISSLPPSSLPSSSSQEKVH 181
>TC15426 weakly similar to UP|HS7E_DROME (P29845) Heat shock 70 kDa protein
cognate 5, partial (3%)
Length = 1216
Score = 26.6 bits (57), Expect = 6.1
Identities = 18/62 (29%), Positives = 30/62 (48%), Gaps = 2/62 (3%)
Frame = +2
Query: 53 KWVLR-NRAKLKGQDWEDVAKHVSSRSN-STKSPKTQTQCKNKIESMKKRYRSESASSDV 110
K V+R + LKG D +D+A +++ +N K+ K K K KK+ + D+
Sbjct: 635 KAVMRLSMTSLKGADAKDIASSITAIANEKIKAEKEANAGKKKTGGKKKQLNVDKPDDDI 814
Query: 111 SS 112
S
Sbjct: 815 VS 820
>TC19735 similar to UP|Q8NJM5 (Q8NJM5) Probable ATP-dependent RNA helicase,
partial (5%)
Length = 565
Score = 26.6 bits (57), Expect = 6.1
Identities = 21/100 (21%), Positives = 42/100 (42%), Gaps = 3/100 (3%)
Frame = +3
Query: 142 VLLEPSSLVMSHAQEEVHQPLVPPPITTAQNS---HGSNGVDKLIKEDEIETKSSDHIAN 198
VL P + E V ++ P Q+S N K K+++ + K +AN
Sbjct: 54 VLPSPLNEAYETVAEAVQNLIMAEPEDNLQSSPMVETENNKKKKEKKNKHDKKRPRDVAN 233
Query: 199 KNPLETDSSTPALYSEKDEQRSKKRRMRTENNNNKRQKKE 238
+ E + S++D + SKK++ + + +K++
Sbjct: 234 EEHAEEEEEQIGNNSDRDGESSKKKKKKKHKVEVEERKEK 353
>TC12976 similar to UP|O02448 (O02448) Lox22-otx (Fragment), partial (3%)
Length = 610
Score = 26.6 bits (57), Expect = 6.1
Identities = 18/76 (23%), Positives = 34/76 (44%)
Frame = +2
Query: 173 SHGSNGVDKLIKEDEIETKSSDHIANKNPLETDSSTPALYSEKDEQRSKKRRMRTENNNN 232
S G++ DK + S + +P T + TPAL E+ ++ KK+R +
Sbjct: 275 SLGNSWNDKFAAAEAASFSSGTRGLSSSP--TPTPTPALMEER-KREGKKKRHGLKRKKR 445
Query: 233 KRQKKETMGIAESIRW 248
KR + ++ ++ W
Sbjct: 446 KRMMRRSIRRRTTVTW 493
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.306 0.123 0.328
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 4,361,424
Number of Sequences: 28460
Number of extensions: 52210
Number of successful extensions: 249
Number of sequences better than 10.0: 40
Number of HSP's better than 10.0 without gapping: 246
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 248
length of query: 325
length of database: 4,897,600
effective HSP length: 90
effective length of query: 235
effective length of database: 2,336,200
effective search space: 549007000
effective search space used: 549007000
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.1 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 43 (22.0 bits)
S2: 55 (25.8 bits)
Medicago: description of AC126006.11


