

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
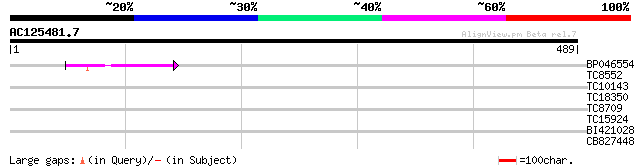
Query= AC125481.7 + phase: 0 /pseudo
(489 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
BP046554 81 4e-16
TC8552 32 0.23
TC10143 28 4.4
TC18350 similar to UP|O81911 (O81911) T7I23.17 protein, partial ... 27 5.8
TC8709 similar to UP|O49929 (O49929) Pore protein of 24 kDa (OEP... 27 7.5
TC15924 weakly similar to GB|BAC21577.1|24060129|AP005438 G-prot... 27 7.5
BI421028 27 7.5
CB827448 27 9.8
>BP046554
Length = 556
Score = 80.9 bits (198), Expect = 4e-16
Identities = 45/103 (43%), Positives = 62/103 (59%), Gaps = 6/103 (5%)
Frame = -3
Query: 49 EAMDMGETDASNVGQKI------VLQPSFAGGRHYMFNNCQDAMSICKKYGYPDLFVTIA 102
EA+D GET+ S+VG ++ ++ + M C I KK+GYPDLF+T
Sbjct: 389 EAIDKGETNPSSVGSELFCPHLLLVDIAICSIISRMQWPC-----IFKKFGYPDLFITFT 225
Query: 103 CNTNWREIQDFLKERNLKASDRPDIVCRVFKMKLDKLMDDFEK 145
CN+ W EIQ F++ RNL D P+I RVFKMKLD+L+ D +K
Sbjct: 224 CNSAWCEIQRFVQPRNLNVEDCPNICVRVFKMKLDRLISDLKK 96
>TC8552
Length = 535
Score = 32.0 bits (71), Expect = 0.23
Identities = 17/39 (43%), Positives = 22/39 (55%)
Frame = +2
Query: 282 CRYVSPCEAVWRIFAFDIHHK*PHVLKLLFHLHNEQGNI 320
C Y SP E W+I+ F + HVLKL+F+ E NI
Sbjct: 374 CDYSSPFEF*WQIYLFHV-----HVLKLVFNTWQESLNI 475
>TC10143
Length = 251
Score = 27.7 bits (60), Expect = 4.4
Identities = 14/36 (38%), Positives = 23/36 (63%)
Frame = -1
Query: 17 SVDCYTMIESQRLRYIRKNQKTIRCGILNGLHEAMD 52
S+ C+ +R R ++KN+K+ + G+LNG E MD
Sbjct: 194 SLVCFRGQNQERERKLKKNEKSKKKGLLNG-KEGMD 90
>TC18350 similar to UP|O81911 (O81911) T7I23.17 protein, partial (44%)
Length = 628
Score = 27.3 bits (59), Expect = 5.8
Identities = 14/47 (29%), Positives = 22/47 (46%)
Frame = +1
Query: 226 RYGGHVNVECCNKSNSIKYLFKYVNKVPDKALMQLSVDGDNRDKSKP 272
RY H C N + +KYL K P+ +++ L+ + D R P
Sbjct: 322 RYSRHELKGCINDAKCMKYLLINKVKFPEPSIIMLTEEEDPRGPKFP 462
>TC8709 similar to UP|O49929 (O49929) Pore protein of 24 kDa (OEP24),
complete
Length = 997
Score = 26.9 bits (58), Expect = 7.5
Identities = 17/52 (32%), Positives = 24/52 (45%), Gaps = 1/52 (1%)
Frame = +2
Query: 169 MDRGRCTKYYRKNYKGTTTIDEGYPRYKRR-DFGIHVDKQRVLLDNRYVVPY 219
+D G C YR +KG TT + Y K DF + +RV D+ + Y
Sbjct: 422 LDSGNCRLKYRYVHKGLTTFEPSYDVAKNTWDFAV---SRRVYGDDSFGASY 568
>TC15924 weakly similar to GB|BAC21577.1|24060129|AP005438 G-protein beta
family-like protein {Oryza sativa (japonica
cultivar-group);}, partial (11%)
Length = 1262
Score = 26.9 bits (58), Expect = 7.5
Identities = 10/18 (55%), Positives = 13/18 (71%)
Frame = +1
Query: 267 RDKSKPVDEIKQYYDCRY 284
R +SKP++ IK Y DC Y
Sbjct: 574 RKQSKPINSIKAYRDCLY 627
>BI421028
Length = 607
Score = 26.9 bits (58), Expect = 7.5
Identities = 15/53 (28%), Positives = 24/53 (44%)
Frame = +3
Query: 138 KLMDDFEKEECLAKLMQRHVPCGGARPFSPCMDRGRCTKYYRKNYKGTTTIDE 190
KL++ EKE + L +R V CG + C + +K + T+DE
Sbjct: 207 KLIESGEKERLMELLRERLVDCGWKDEM-----KAICRDFVKKKGRNNVTVDE 350
>CB827448
Length = 549
Score = 26.6 bits (57), Expect = 9.8
Identities = 16/59 (27%), Positives = 25/59 (42%), Gaps = 5/59 (8%)
Frame = +3
Query: 127 IVCRVFKMKLDKLMDDFEKEECLAKLMQRHVPCGG-----ARPFSPCMDRGRCTKYYRK 180
+VC+ K + ++ FEKEE A L + F+ C+ G Y+RK
Sbjct: 309 LVCQAQKASMAAAVEVFEKEELAASLAKYVADLSNKFTAERGAFTVCLSGGSLISYFRK 485
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.342 0.150 0.514
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 8,727,641
Number of Sequences: 28460
Number of extensions: 121238
Number of successful extensions: 741
Number of sequences better than 10.0: 16
Number of HSP's better than 10.0 without gapping: 741
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 741
length of query: 489
length of database: 4,897,600
effective HSP length: 94
effective length of query: 395
effective length of database: 2,222,360
effective search space: 877832200
effective search space used: 877832200
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 39 (22.0 bits)
S2: 57 (26.6 bits)
Medicago: description of AC125481.7


