

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC125480.3 + phase: 0
(476 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
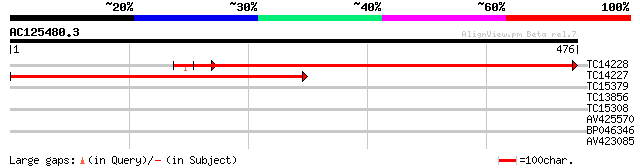
Score E
Sequences producing significant alignments: (bits) Value
TC14228 homologue to UP|Q9LLR4 (Q9LLR4) Sec61p, partial (67%) 627 e-180
TC14227 homologue to UP|Q9LLR4 (Q9LLR4) Sec61p, partial (53%) 482 e-137
TC15379 UP|Q9ZPL8 (Q9ZPL8) Protein phosphatase type 2C, complete 29 1.9
TC13856 28 4.3
TC15308 similar to UP|Q41725 (Q41725) TED3, partial (16%) 28 4.3
AV425570 28 4.3
BP046346 27 5.6
AV423085 27 9.5
>TC14228 homologue to UP|Q9LLR4 (Q9LLR4) Sec61p, partial (67%)
Length = 1268
Score = 627 bits (1616), Expect = e-180
Identities = 316/322 (98%), Positives = 320/322 (99%)
Frame = +2
Query: 155 IVQLFFAGIIVICLDELLQKGYGLGSGISLFIATNICENIIWKAFSPTTINSGRGAEFEG 214
++QL FAGIIVICLDELLQKGYGLGSGISLFIATNICENIIWKAFSPTTINSGRGAEFEG
Sbjct: 77 LLQLCFAGIIVICLDELLQKGYGLGSGISLFIATNICENIIWKAFSPTTINSGRGAEFEG 256
Query: 215 AVIALFHLLITRTDKVRALREAFYRQNLPNVTNLLATVLIFLIVIYFQGFRVVLPVRSKN 274
AVIALFHLLITR+DKVRALREAFYRQNLPNVTNLLATVLIFLIVIYFQGFRVVLPVRSKN
Sbjct: 257 AVIALFHLLITRSDKVRALREAFYRQNLPNVTNLLATVLIFLIVIYFQGFRVVLPVRSKN 436
Query: 275 ARGQQGSYPIKLFYTSNMPIILQSALVSNLYFISQLLHRKYSGNFIVNLLGKWKESEYGG 334
ARGQQGSYPIKLFYTSNMPIILQSALVSNLYFISQLLHRKYSGNF VNLLGKWK+SEYGG
Sbjct: 437 ARGQQGSYPIKLFYTSNMPIILQSALVSNLYFISQLLHRKYSGNFFVNLLGKWKDSEYGG 616
Query: 335 GHSIPVGGIAYYITAPSSLADMAANPFHALFYLVFMLSACALFSKTWIEVSGSSARDVAK 394
GHSIPVGGIAYYITAPSSLADMAANPFHALFYLVFMLSACALFSKTWIEVSGSSARDVAK
Sbjct: 617 GHSIPVGGIAYYITAPSSLADMAANPFHALFYLVFMLSACALFSKTWIEVSGSSARDVAK 796
Query: 395 QLKEQQMVMPGHRESNLQKELNRYIPTAAAFGGICIGALTVLADFMGAIGSGTGILLAVT 454
QLKEQQMVMPGHRESNLQKELNRYIPTAAAFGGICIGALTVLADFMGAIGSGTGILLAVT
Sbjct: 797 QLKEQQMVMPGHRESNLQKELNRYIPTAAAFGGICIGALTVLADFMGAIGSGTGILLAVT 976
Query: 455 IIYQYFETFEKERASELGFFGF 476
IIYQYFETFEKERASELGFFGF
Sbjct: 977 IIYQYFETFEKERASELGFFGF 1042
Score = 47.4 bits (111), Expect = 5e-06
Identities = 26/42 (61%), Positives = 31/42 (72%), Gaps = 6/42 (14%)
Frame = -1
Query: 138 GMYGSVGQL------GVGNAILIIVQLFFAGIIVICLDELLQ 173
G + ++ QL GVGNAILII+QL FAGIIVICL EL+Q
Sbjct: 128 GAHPNISQLYQQNTAGVGNAILIILQLCFAGIIVICLKELIQ 3
>TC14227 homologue to UP|Q9LLR4 (Q9LLR4) Sec61p, partial (53%)
Length = 880
Score = 482 bits (1240), Expect = e-137
Identities = 247/250 (98%), Positives = 249/250 (98%)
Frame = +2
Query: 1 MGGGFRVLHLVRPFLSFLPEVQTADRKVPFREKVIYTVISLFIFLVCSQLPLYGIHSTTG 60
MGGGFRVLHLVRPFLSFLPEVQTADRKVPFREKVIYTVISLFIFLVCSQLPLYGIHSTTG
Sbjct: 131 MGGGFRVLHLVRPFLSFLPEVQTADRKVPFREKVIYTVISLFIFLVCSQLPLYGIHSTTG 310
Query: 61 ADPFYWMRVILASNRGTVMELGITPIVTSGLVMQLLAGSKIIEVDNNVREDRALLNGAQK 120
ADPFYWMRVILASNRGTVMELGITPIVTSGLVMQLLAGSKIIEVDNNVREDRALLNGAQK
Sbjct: 311 ADPFYWMRVILASNRGTVMELGITPIVTSGLVMQLLAGSKIIEVDNNVREDRALLNGAQK 490
Query: 121 LLGILIAVGEAVAYVLSGMYGSVGQLGVGNAILIIVQLFFAGIIVICLDELLQKGYGLGS 180
LLGILIAVGEAVAYVLSGMYGSVGQLGVGNAILII+QL FAGIIVICLDELLQKGYGLGS
Sbjct: 491 LLGILIAVGEAVAYVLSGMYGSVGQLGVGNAILIILQLCFAGIIVICLDELLQKGYGLGS 670
Query: 181 GISLFIATNICENIIWKAFSPTTINSGRGAEFEGAVIALFHLLITRTDKVRALREAFYRQ 240
GISLFIATNICENIIWKAFSPTTINSGRGAEFEGAVIALFHLLITR+DKVRALREAFYRQ
Sbjct: 671 GISLFIATNICENIIWKAFSPTTINSGRGAEFEGAVIALFHLLITRSDKVRALREAFYRQ 850
Query: 241 NLPNVTNLLA 250
NLPNVTNLLA
Sbjct: 851 NLPNVTNLLA 880
>TC15379 UP|Q9ZPL8 (Q9ZPL8) Protein phosphatase type 2C, complete
Length = 1338
Score = 28.9 bits (63), Expect = 1.9
Identities = 16/52 (30%), Positives = 28/52 (53%), Gaps = 2/52 (3%)
Frame = +1
Query: 350 PSSLADMAANPFHALFYLVFMLSAC--ALFSKTWIEVSGSSARDVAKQLKEQ 399
PS ++ NPFH LF+L+ + LF I+V + R++ ++KE+
Sbjct: 106 PSQHTRLSLNPFHFLFFLLSLSLHLLHTLF*ILQIQVQAMTGREILHKMKEK 261
>TC13856
Length = 570
Score = 27.7 bits (60), Expect = 4.3
Identities = 14/38 (36%), Positives = 18/38 (46%)
Frame = -1
Query: 377 FSKTWIEVSGSSARDVAKQLKEQQMVMPGHRESNLQKE 414
F K W+ S + +A+QLK MV P H Q E
Sbjct: 231 FKKAWLSHKVYSLKTIARQLKRLCMVWPPHSGDLKQPE 118
>TC15308 similar to UP|Q41725 (Q41725) TED3, partial (16%)
Length = 1103
Score = 27.7 bits (60), Expect = 4.3
Identities = 26/114 (22%), Positives = 52/114 (44%), Gaps = 3/114 (2%)
Frame = -1
Query: 121 LLGILIAVGEAVAYVLSGMYGSVGQLGVGNAILIIVQLFFAGIIVICLDELLQKGYGLGS 180
L+ I+IA+ + V+ + AI++I + F+ ++V C+ ++ +G
Sbjct: 581 LIAIIIAIIVVIVVVVKFL-----------AIVVIAPVPFS-VVVFCVHFIVVVAAEIGL 438
Query: 181 GISLFIA---TNICENIIWKAFSPTTINSGRGAEFEGAVIALFHLLITRTDKVR 231
GIS F+A + E +I+K S + GR + A+ + ++ K R
Sbjct: 437 GISCFVALVFEVLIEKLIFKVCSGGSFLGGRILSRTRRICAI*AITLSHVKKFR 276
>AV425570
Length = 420
Score = 27.7 bits (60), Expect = 4.3
Identities = 33/116 (28%), Positives = 48/116 (40%), Gaps = 6/116 (5%)
Frame = +1
Query: 81 LGITPIVTSGLVMQLLAGSKIIEVDNNVREDRALLNGAQKLL------GILIAVGEAVAY 134
LGI P + + +V QLLA D RE A G +K+L + A+ +A+
Sbjct: 61 LGIVPFINAQIVFQLLAQVYPKLQDLQKREGEA---GRKKILQYTRYASVGFAIVQAIGQ 231
Query: 135 VLSGMYGSVGQLGVGNAILIIVQLFFAGIIVICLDELLQKGYGLGSGISLFIATNI 190
VL + V I ++ L + E + LG+G SL I TNI
Sbjct: 232 VLF-LRPYVNDFSTEWVISSVILLTLGSAFTTYIGERI-TDLKLGNGTSLLIFTNI 393
>BP046346
Length = 476
Score = 27.3 bits (59), Expect = 5.6
Identities = 15/43 (34%), Positives = 23/43 (52%)
Frame = +1
Query: 24 ADRKVPFREKVIYTVISLFIFLVCSQLPLYGIHSTTGADPFYW 66
A R +P ++KVIY I I + ++ + G HST D +W
Sbjct: 280 AQRALP-KKKVIYDFIVNKILRLVQEISIQGAHSTDKQDIGFW 405
>AV423085
Length = 411
Score = 26.6 bits (57), Expect = 9.5
Identities = 12/24 (50%), Positives = 16/24 (66%)
Frame = -3
Query: 171 LLQKGYGLGSGISLFIATNICENI 194
L QK Y LGS +SLF +N+ N+
Sbjct: 127 LAQKAYSLGSFLSLFGRSNLVANL 56
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.326 0.143 0.418
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 7,298,137
Number of Sequences: 28460
Number of extensions: 96252
Number of successful extensions: 654
Number of sequences better than 10.0: 16
Number of HSP's better than 10.0 without gapping: 651
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 653
length of query: 476
length of database: 4,897,600
effective HSP length: 94
effective length of query: 382
effective length of database: 2,222,360
effective search space: 848941520
effective search space used: 848941520
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.1 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.6 bits)
S2: 57 (26.6 bits)
Medicago: description of AC125480.3


