

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
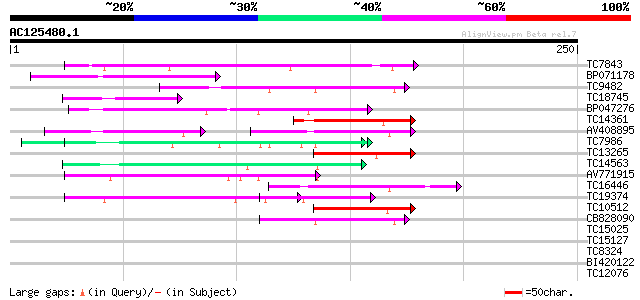
Query= AC125480.1 - phase: 0 /pseudo
(250 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
TC7843 weakly similar to GB|AAF03412.1|6090963|AF188712 mitochon... 55 9e-09
BP071178 50 4e-07
TC9482 similar to UP|Q9FI73 (Q9FI73) Mitochondrial carrier prote... 49 8e-07
TC18745 weakly similar to UP|ADT2_HUMAN (P05141) ADP,ATP carrier... 45 2e-05
BP047276 44 2e-05
TC14361 similar to PIR|T01729|T01729 mitochondrial solute carrie... 44 3e-05
AV408895 44 3e-05
TC7986 similar to UP|Q8SF02 (Q8SF02) Dicarboxylate/tricarboxylat... 43 4e-05
TC13265 similar to PIR|T01729|T01729 mitochondrial solute carrie... 42 8e-05
TC14563 similar to UP|Q9SSP4 (Q9SSP4) F3N23.2 protein (At1g72820... 42 1e-04
AV771915 42 1e-04
TC16446 similar to UP|BAC84529 (BAC84529) Mitochondrial aspartat... 41 2e-04
TC19374 similar to UP|Q8LNZ1 (Q8LNZ1) Mitochondrial uncoupling p... 40 4e-04
TC10512 similar to UP|Q9FI73 (Q9FI73) Mitochondrial carrier prot... 39 6e-04
CB828090 39 6e-04
TC15025 similar to UP|MCAT_ARATH (Q93XM7) Mitochondrial carnitin... 39 0.001
TC15127 weakly similar to GB|AAG29586.1|11094343|AF155813 mitoch... 38 0.001
TC8324 UP|Q8VWY1 (Q8VWY1) Mitochondrial phosphate transporter, c... 38 0.002
BI420122 37 0.003
TC12076 similar to GB|AAL69531.1|18491125|AY074833 At2g47490/T30... 36 0.007
>TC7843 weakly similar to GB|AAF03412.1|6090963|AF188712 mitochondrial
dicarboxylate carrier {Mus musculus;} , partial (22%)
Length = 1594
Score = 55.5 bits (132), Expect = 9e-09
Identities = 47/172 (27%), Positives = 76/172 (43%), Gaps = 16/172 (9%)
Frame = +3
Query: 25 PLDVVQTRLQVHDGRL-FSQYPRYNNTTHAIFTIARKEGLKGLYAG----------FPAG 73
P DV R+Q DGRL +Q Y++ AI +A++EG+ L+ G A
Sbjct: 699 PADVAMVRMQA-DGRLPAAQRRNYSSVVDAITRMAKQEGVTSLWRGSSLTVNRAMLVTAS 875
Query: 74 LLGATISWSLLVFYYGIVKDRHARSKVEKSGALMTVCLCANPAFVVKTR-LQLQTPLHHA 132
L + + ++ G+++D A + +NP V+KTR + ++
Sbjct: 876 QLASYDQFKEMILEKGLMRDGLGTHVTASFAAGFVAAVASNPVDVIKTRVMNMKVEPGAE 1055
Query: 133 RPYSGLYDAFRTIKREEGFSAFYRGIVPGFFLISQ----AAIQFIVYEQLRK 180
PY+G D R EG A Y+G +P IS+ + F+ EQ+RK
Sbjct: 1056PPYAGALDCALKTVRAEGPMALYKGFIP---TISRQGPFTVVLFVTLEQVRK 1202
>BP071178
Length = 408
Score = 50.1 bits (118), Expect = 4e-07
Identities = 26/84 (30%), Positives = 41/84 (47%)
Frame = +2
Query: 10 VASLTTGFAFITFQYPLDVVQTRLQVHDGRLFSQYPRYNNTTHAIFTIARKEGLKGLYAG 69
VA G + YPLD+ TRL GR ++ ++ H + TI K+G++G+Y G
Sbjct: 119 VAGAAAGCTSLVLVYPLDIAHTRLAADIGR--TEVRQFRGIHHFLVTIFHKDGIRGIYRG 292
Query: 70 FPAGLLGATISWSLLVFYYGIVKD 93
PA L G + L + +K+
Sbjct: 293 LPASLHGMVVHRGLYFGGFDTIKE 364
Score = 25.8 bits (55), Expect = 7.4
Identities = 15/59 (25%), Positives = 24/59 (40%)
Frame = +2
Query: 100 VEKSGALMTVCLCANPAFVVKTRLQLQTPLHHARPYSGLYDAFRTIKREEGFSAFYRGI 158
V + A T + P + TRL R + G++ TI ++G YRG+
Sbjct: 119 VAGAAAGCTSLVLVYPLDIAHTRLAADIGRTEVRQFRGIHHFLVTIFHKDGIRGIYRGL 295
>TC9482 similar to UP|Q9FI73 (Q9FI73) Mitochondrial carrier protein-like
(At5g48970), partial (42%)
Length = 529
Score = 48.9 bits (115), Expect = 8e-07
Identities = 36/130 (27%), Positives = 57/130 (43%), Gaps = 20/130 (15%)
Frame = +3
Query: 67 YAGFPAGLLGATISWSLLVFYYGIVKDRHARSKVEKSGALMTVCLCA-----------NP 115
YAG G W++ +Y R++ + E S + + LC +P
Sbjct: 63 YAGLQFGTYDTFKRWAMAWNHY-----RYSNTSAENSPSSFQLFLCGLAAGTCAKLVCHP 227
Query: 116 AFVVKTRLQLQTPLHHAR--------PYSGLYDAFRTIKREEGFSAFYRGIVPGFFLISQ 167
VVK R Q++ H R YS ++DA + I R EG++ Y+GI+P +
Sbjct: 228 LDVVKKRFQIEGLQRHPRYGARVERRAYSSMFDAIQRILRLEGWAGLYKGIIPSTVKAAP 407
Query: 168 A-AIQFIVYE 176
A A+ F+ YE
Sbjct: 408 AGAVTFVAYE 437
Score = 31.2 bits (69), Expect = 0.18
Identities = 22/76 (28%), Positives = 34/76 (43%), Gaps = 6/76 (7%)
Frame = +3
Query: 24 YPLDVVQTRLQVHDGRLFSQY------PRYNNTTHAIFTIARKEGLKGLYAGFPAGLLGA 77
+PLDVV+ R Q+ + +Y Y++ AI I R EG GLY G + A
Sbjct: 222 HPLDVVKKRFQIEGLQRHPRYGARVERRAYSSMFDAIQRILRLEGWAGLYKGIIPSTVKA 401
Query: 78 TISWSLLVFYYGIVKD 93
+ ++ Y + D
Sbjct: 402 APAGAVTFVAYELTSD 449
>TC18745 weakly similar to UP|ADT2_HUMAN (P05141) ADP,ATP carrier protein,
fibroblast isoform (ADP/ATP translocase 2) (Adenine
nucleotide translocator 2) (ANT 2), partial (9%)
Length = 834
Score = 44.7 bits (104), Expect = 2e-05
Identities = 24/53 (45%), Positives = 29/53 (54%)
Frame = +3
Query: 24 YPLDVVQTRLQVHDGRLFSQYPRYNNTTHAIFTIARKEGLKGLYAGFPAGLLG 76
YPLD+V+TRL ++ Y +HA TI R EG GLY G A LLG
Sbjct: 648 YPLDLVRTRLAAQRSTIY-----YRGISHAFSTICRDEGFLGLYKGLGATLLG 791
Score = 36.6 bits (83), Expect = 0.004
Identities = 24/63 (38%), Positives = 33/63 (52%), Gaps = 1/63 (1%)
Frame = +3
Query: 115 PAFVVKTRLQLQTPLHHARPYSGLYDAFRTIKREEGFSAFYRGIVPGFFLISQA-AIQFI 173
P +V+TRL Q + Y G+ AF TI R+EGF Y+G+ + + AI F
Sbjct: 651 PLDLVRTRLAAQRSTIY---YRGISHAFSTICRDEGFLGLYKGLGATLLGVGPSIAISFS 821
Query: 174 VYE 176
VYE
Sbjct: 822 VYE 830
>BP047276
Length = 516
Score = 44.3 bits (103), Expect = 2e-05
Identities = 40/147 (27%), Positives = 60/147 (40%), Gaps = 13/147 (8%)
Frame = +3
Query: 27 DVVQTRLQVHDGRLFSQYPRYNNTTHAIFTIARKEGLKGLYAGFPAGLLGATISWSLLV- 85
DV++TRLQ+ + Y H TI+R EG++ L+ G T ++L +
Sbjct: 84 DVIKTRLQL------DRSGNYKGIVHCGSTISRTEGVRALWKGLTPFATHLTFKYALRMG 245
Query: 86 ---FYYGIVKDRHARSKVEKSGALMT--------VCLCANPAFVVKTRLQLQTPLH-HAR 133
+ + KD K+ G L++ + P VVK +LQ Q L
Sbjct: 246 SNAVFQSMFKDSET-GKLSSHGRLLSGFGAGVLEAIVIVTPFEVVKIKLQQQRGLSPELL 422
Query: 134 PYSGLYDAFRTIKREEGFSAFYRGIVP 160
Y G RTI EEG + G+ P
Sbjct: 423 KYKGPVHCARTILHEEGIRGLWAGVSP 503
>TC14361 similar to PIR|T01729|T01729 mitochondrial solute carrier protein
homolog - Arabidopsis thaliana
{Arabidopsis thaliana;}, partial (20%)
Length = 591
Score = 43.9 bits (102), Expect = 3e-05
Identities = 22/55 (40%), Positives = 33/55 (60%), Gaps = 1/55 (1%)
Frame = +3
Query: 126 QTPLHHARPYSGLYDAFRTIKREEGFSAFYRGIVPGFF-LISQAAIQFIVYEQLR 179
+TPL Y+G+ DAFR R EGF A Y+G+VP ++ A+ F+ YE ++
Sbjct: 24 KTPLE----YTGMVDAFRKTVRYEGFGALYKGLVPNSVKVVPSIALAFVTYEMVK 176
>AV408895
Length = 429
Score = 43.5 bits (101), Expect = 3e-05
Identities = 27/73 (36%), Positives = 38/73 (51%), Gaps = 2/73 (2%)
Frame = +3
Query: 16 GFAFITFQYPLDVVQTRLQVHDGRLFSQYPRYNNTTHAIFTIARKEGLKGLYAGFPAGLL 75
G T YPLD+V+TRL + + Y HA+ TI+++EG+ GLY G LL
Sbjct: 213 GITAATSTYPLDLVRTRLAAQ-----TNFTYYRGIWHALQTISKEEGILGLYKGLGTTLL 377
Query: 76 --GATISWSLLVF 86
G I+ S V+
Sbjct: 378 TVGPNIAISFSVY 416
Score = 43.1 bits (100), Expect = 4e-05
Identities = 26/74 (35%), Positives = 37/74 (49%), Gaps = 1/74 (1%)
Frame = +3
Query: 107 MTVCLCANPAFVVKTRLQLQTPLHHARPYSGLYDAFRTIKREEGFSAFYRGIVPGFFLIS 166
+T P +V+TRL QT + Y G++ A +TI +EEG Y+G+ +
Sbjct: 216 ITAATSTYPLDLVRTRLAAQTNFTY---YRGIWHALQTISKEEGILGLYKGLGTTLLTVG 386
Query: 167 -QAAIQFIVYEQLR 179
AI F VYE LR
Sbjct: 387 PNIAISFSVYESLR 428
>TC7986 similar to UP|Q8SF02 (Q8SF02) Dicarboxylate/tricarboxylate carrier,
partial (96%)
Length = 1532
Score = 43.1 bits (100), Expect = 4e-05
Identities = 41/162 (25%), Positives = 63/162 (38%), Gaps = 10/162 (6%)
Frame = +3
Query: 6 WEYPVASLTTGFAFITFQYPLDVVQTRLQVHDGRLFSQYPRYNNTTHAIFTIARKEGLKG 65
++ + LT G T P D+ R+Q Q Y N HA++ I EG+
Sbjct: 507 YQKALCGLTAGAIGATVGSPADLALIRMQADATLPLHQRRHYTNAFHALYRIVADEGVLS 686
Query: 66 LYAGF-PAGLLGATISWSLLVFYYGIV---KDRHARSKVEKSGALMTV-----CLCANPA 116
L+ G P + ++ +L Y V KD + A +V C+ P
Sbjct: 687 LWKGAGPTVVRAMALNMGMLASYDQSVEFFKDSVGLGEASTVVAASSVSGFFAAACSLPF 866
Query: 117 FVVKTRLQLQTPLHHAR-PYSGLYDAFRTIKREEGFSAFYRG 157
VKT++Q P + PY+G D ++ G FY G
Sbjct: 867 DYVKTQIQKMQPDAEGKYPYTGSLDCAAKTLKQGGPFKFYTG 992
Score = 38.9 bits (89), Expect = 8e-04
Identities = 32/149 (21%), Positives = 58/149 (38%), Gaps = 13/149 (8%)
Frame = +3
Query: 25 PLDVVQTRLQVHDGRLFSQYPRYNNTTHAIFTIARKEGLKGLYAGFPAGLLGATISWSLL 84
P+D+++ R+Q+ G + + + +G+ Y G AGLL +
Sbjct: 291 PIDMIKVRIQLGQG----------SAVQVTSNMLKNDGVGAFYKGLSAGLLRQATYTTAR 440
Query: 85 VFYYGIVKDRHARSKVEKSGALMTVCLCA-----------NPAFVVKTRLQLQT--PLHH 131
+ + I+ + + K L LC +PA + R+Q PLH
Sbjct: 441 LGTFRILTSKAIEANDGKPLPLYQKALCGLTAGAIGATVGSPADLALIRMQADATLPLHQ 620
Query: 132 ARPYSGLYDAFRTIKREEGFSAFYRGIVP 160
R Y+ + A I +EG + ++G P
Sbjct: 621 RRHYTNAFHALYRIVADEGVLSLWKGAGP 707
>TC13265 similar to PIR|T01729|T01729 mitochondrial solute carrier protein
homolog - Arabidopsis thaliana
{Arabidopsis thaliana;}, partial (24%)
Length = 561
Score = 42.4 bits (98), Expect = 8e-05
Identities = 19/46 (41%), Positives = 29/46 (62%), Gaps = 1/46 (2%)
Frame = +2
Query: 135 YSGLYDAFRTIKREEGFSAFYRGIVP-GFFLISQAAIQFIVYEQLR 179
Y+G+ DAFR R EGF A Y+G+VP ++ A+ F+ YE ++
Sbjct: 89 YTGMVDAFRKTVRHEGFGALYQGLVPISVKVVPSLALAFVTYEVVK 226
>TC14563 similar to UP|Q9SSP4 (Q9SSP4) F3N23.2 protein (At1g72820/F3N23_2),
partial (55%)
Length = 1025
Score = 41.6 bits (96), Expect = 1e-04
Identities = 41/156 (26%), Positives = 57/156 (36%), Gaps = 22/156 (14%)
Frame = +2
Query: 24 YPLDVVQTRLQVHDGRLFSQYPRYNNTTHAIFTIARKEGLKGLYAGFPAGLLGATISWSL 83
YP V++TR QV + + FT+ R EG + LY GF L+G + +L
Sbjct: 521 YPAVVLKTRQQVAQTHV--------SCIKTAFTLIRGEGFRALYRGFGTSLMGTIPARAL 676
Query: 84 LVFYYGIVKDRHARSKVEKS----------------GALMTVCLCANPAFVVKTRLQLQT 127
+ + K + V A M L P VV RL +Q
Sbjct: 677 YMAALEVTKSNVGTATVRFGFAEPTAAAIASAAAGLSAAMAAQLVWTPVDVVSQRLMVQG 856
Query: 128 PLHHARP------YSGLYDAFRTIKREEGFSAFYRG 157
+ + P Y DAFR I +G YRG
Sbjct: 857 CCNSSNPKGSAVRYINGIDAFRKILNTDGPRGLYRG 964
Score = 27.3 bits (59), Expect = 2.5
Identities = 24/83 (28%), Positives = 35/83 (41%)
Frame = +2
Query: 115 PAFVVKTRLQLQTPLHHARPYSGLYDAFRTIKREEGFSAFYRGIVPGFFLISQAAIQFIV 174
PA V+KTR Q+ A+ + T+ R EGF A YRG G L+ + +
Sbjct: 524 PAVVLKTRQQV------AQTHVSCIKTAFTLIRGEGFRALYRGF--GTSLMGTIPARALY 679
Query: 175 YEQLRKTVVNLKTKGSKIQHQKP 197
L T N+ T + +P
Sbjct: 680 MAALEVTKSNVGTATVRFGFAEP 748
>AV771915
Length = 490
Score = 41.6 bits (96), Expect = 1e-04
Identities = 43/129 (33%), Positives = 55/129 (42%), Gaps = 16/129 (12%)
Frame = +1
Query: 25 PLDVVQTRLQVHDGRLFSQ---YPRYNNTTHAIFTIARKEGLKGLYAGFPAGLLGATISW 81
PLD + RLQ+ L P+Y + TIAR+EGL L+ G GL +
Sbjct: 103 PLDTAKVRLQLQKQALTGDGVALPKYKGM*GTVATIAREEGLASLWKGIVPGLHRQCVYG 282
Query: 82 SLLVFYYGIVKDRH-ARSKV------EKSGALMT----VCLCANPAFVVKTRLQLQTPLH 130
L V Y VK + R V +K A +T ANP +VK RLQ + L
Sbjct: 283 GLRVGLYEPVKTLYVGRDHVGDVPWSKKILAALTTGAVAIAVANPTDLVKVRLQAEGKLP 462
Query: 131 HARP--YSG 137
P YSG
Sbjct: 463 PGVPRRYSG 489
Score = 37.0 bits (84), Expect = 0.003
Identities = 24/56 (42%), Positives = 30/56 (52%), Gaps = 5/56 (8%)
Frame = +1
Query: 111 LCANPAFVVKTRLQLQ----TPLHHARP-YSGLYDAFRTIKREEGFSAFYRGIVPG 161
+C P K RLQLQ T A P Y G+ TI REEG ++ ++GIVPG
Sbjct: 91 VCTIPLDTAKVRLQLQKQALTGDGVALPKYKGM*GTVATIAREEGLASLWKGIVPG 258
Score = 28.1 bits (61), Expect = 1.5
Identities = 17/47 (36%), Positives = 26/47 (55%), Gaps = 1/47 (2%)
Frame = +1
Query: 1 MNELQWEYPV-ASLTTGFAFITFQYPLDVVQTRLQVHDGRLFSQYPR 46
+ ++ W + A+LTTG I P D+V+ RLQ +G+L PR
Sbjct: 340 VGDVPWSKKILAALTTGAVAIAVANPTDLVKVRLQA-EGKLPPGVPR 477
>TC16446 similar to UP|BAC84529 (BAC84529) Mitochondrial aspartate-glutamate
carrier protein-like, partial (43%)
Length = 668
Score = 40.8 bits (94), Expect = 2e-04
Identities = 30/86 (34%), Positives = 45/86 (51%), Gaps = 1/86 (1%)
Frame = +3
Query: 115 PAFVVKTRLQLQTPLHHARPYSGLYDAFRTIKREEGFSAFYRGIVPGFFLIS-QAAIQFI 173
P V+KTRL +Q A Y G+ D +TI +EEG AF +GI P I +I F
Sbjct: 165 PLDVIKTRLMVQGS---ANQYKGIVDCVQTIMKEEGPRAFLKGIGPRVLWIGIGGSIFFG 335
Query: 174 VYEQLRKTVVNLKTKGSKIQHQKPDQ 199
V E ++ +V + + + QH K ++
Sbjct: 336 VLESSKRFLV--ERRPTLAQHSKSER 407
Score = 27.7 bits (60), Expect = 1.9
Identities = 18/68 (26%), Positives = 32/68 (46%)
Frame = +3
Query: 25 PLDVVQTRLQVHDGRLFSQYPRYNNTTHAIFTIARKEGLKGLYAGFPAGLLGATISWSLL 84
PLDV++TRL V +Y + TI ++EG + G +L I S+
Sbjct: 165 PLDVIKTRLMVQ-----GSANQYKGIVDCVQTIMKEEGPRAFLKGIGPRVLWIGIGGSI- 326
Query: 85 VFYYGIVK 92
++G+++
Sbjct: 327 --FFGVLE 344
>TC19374 similar to UP|Q8LNZ1 (Q8LNZ1) Mitochondrial uncoupling protein,
partial (49%)
Length = 556
Score = 40.0 bits (92), Expect = 4e-04
Identities = 23/56 (41%), Positives = 28/56 (49%), Gaps = 5/56 (8%)
Frame = +3
Query: 111 LCANPAFVVKTRLQLQTP-----LHHARPYSGLYDAFRTIKREEGFSAFYRGIVPG 161
+C P K RLQLQ + Y G+ TI REEG SA ++GIVPG
Sbjct: 165 VCTIPLDTAKVRLQLQKQGIAGDVASLPKYKGMLGTIATIAREEGASALWKGIVPG 332
Score = 38.9 bits (89), Expect = 8e-04
Identities = 35/119 (29%), Positives = 50/119 (41%), Gaps = 14/119 (11%)
Frame = +3
Query: 25 PLDVVQTRLQVHDGRL---FSQYPRYNNTTHAIFTIARKEGLKGLYAGFPAGLLGATISW 81
PLD + RLQ+ + + P+Y I TIAR+EG L+ G GL +
Sbjct: 177 PLDTAKVRLQLQKQGIAGDVASLPKYKGMLGTIATIAREEGASALWKGIVPGLHRQCLYG 356
Query: 82 SLLVFYYGIVKDRHARS----------KVEKSGALMTVCL-CANPAFVVKTRLQLQTPL 129
L + Y VK + S K+ + V + ANP +VK RLQ + L
Sbjct: 357 GLRIGLYEPVKALYVGSDHVGDVPLSKKILAAFTTGAVAITVANPTDLVKVRLQAEGKL 533
Score = 28.9 bits (63), Expect = 0.87
Identities = 16/37 (43%), Positives = 22/37 (59%)
Frame = +3
Query: 10 VASLTTGFAFITFQYPLDVVQTRLQVHDGRLFSQYPR 46
+A+ TTG IT P D+V+ RLQ +G+L PR
Sbjct: 444 LAAFTTGAVAITVANPTDLVKVRLQA-EGKLAPGVPR 551
>TC10512 similar to UP|Q9FI73 (Q9FI73) Mitochondrial carrier protein-like
(At5g48970), partial (43%)
Length = 563
Score = 39.3 bits (90), Expect = 6e-04
Identities = 19/46 (41%), Positives = 28/46 (60%), Gaps = 1/46 (2%)
Frame = +1
Query: 135 YSGLYDAFRTIKREEGFSAFYRGIVPGFFLI-SQAAIQFIVYEQLR 179
Y+G++ A + I REEG F+RG VP ++ AIQF V +L+
Sbjct: 292 YTGMFQASKDILREEGVQGFWRGNVPALLMVMPYTAIQFTVLHKLK 429
>CB828090
Length = 427
Score = 39.3 bits (90), Expect = 6e-04
Identities = 23/75 (30%), Positives = 37/75 (48%), Gaps = 9/75 (12%)
Frame = +2
Query: 111 LCANPAFVVKTRLQLQTPLHHAR--------PYSGLYDAFRTIKREEGFSAFYRGIVPGF 162
L +P VVK R Q++ H R Y ++DA + I + EG++ Y+G+ P
Sbjct: 23 LVCHPLDVVKKRFQIEGLQRHPRYGARVEHRAYKNMFDAMKRIIQMEGWAGLYKGLFPST 202
Query: 163 FLISQA-AIQFIVYE 176
+ A A+ F+ YE
Sbjct: 203 VKAAPAGAVTFVAYE 247
Score = 30.4 bits (67), Expect = 0.30
Identities = 23/71 (32%), Positives = 32/71 (44%), Gaps = 7/71 (9%)
Frame = +2
Query: 24 YPLDVVQTRLQVHDGRLFSQYPR------YNNTTHAIFTIARKEGLKGLYAG-FPAGLLG 76
+PLDVV+ R Q+ + +Y Y N A+ I + EG GLY G FP+ +
Sbjct: 32 HPLDVVKKRFQIEGLQRHPRYGARVEHRAYKNMFDAMKRIIQMEGWAGLYKGLFPSTVKA 211
Query: 77 ATISWSLLVFY 87
A V Y
Sbjct: 212 APAGAVTFVAY 244
>TC15025 similar to UP|MCAT_ARATH (Q93XM7) Mitochondrial
carnitine/acylcarnitine carrier-like protein (A BOUT DE
SOUFFLE) (Carnitine/acylcarnitine translocase-like
protein) (CAC-like protein), partial (97%)
Length = 1256
Score = 38.5 bits (88), Expect = 0.001
Identities = 23/61 (37%), Positives = 32/61 (51%)
Frame = +2
Query: 10 VASLTTGFAFITFQYPLDVVQTRLQVHDGRLFSQYPRYNNTTHAIFTIARKEGLKGLYAG 69
VA G +F YP DVV++ +QV D + P+++ + A I EG KGLY G
Sbjct: 803 VAGGLAGASFWFLVYPTDVVKSVIQVDD----YKNPKFSGSLDAFRKIQATEGFKGLYKG 970
Query: 70 F 70
F
Sbjct: 971 F 973
Score = 36.6 bits (83), Expect = 0.004
Identities = 24/69 (34%), Positives = 33/69 (47%), Gaps = 1/69 (1%)
Frame = +2
Query: 115 PAFVVKTRLQLQTPLHHARPYSGLYDAFRTIKREEGFSAFYRGIVPGFF-LISQAAIQFI 173
P VVK+ +Q+ + +SG DAFR I+ EGF Y+G P I A F+
Sbjct: 848 PTDVVKSVIQVDD--YKNPKFSGSLDAFRKIQATEGFKGLYKGFGPAMARSIPANAACFL 1021
Query: 174 VYEQLRKTV 182
YE R +
Sbjct: 1022AYEMTRSAL 1048
>TC15127 weakly similar to GB|AAG29586.1|11094343|AF155813 mitochondrial
uncoupling protein 5 short form {Mus musculus;}, partial
(24%)
Length = 650
Score = 38.1 bits (87), Expect = 0.001
Identities = 27/76 (35%), Positives = 42/76 (54%), Gaps = 6/76 (7%)
Frame = +3
Query: 111 LCANPAFVVKTR-LQLQTPLHHARPYSGLYD-AFRTIKREEGFSAFYRGIVPGFFLISQ- 167
+ +NP V+KTR + ++ A PY+G D A +T+K E G A Y+G +P IS+
Sbjct: 189 VASNPVDVIKTRVMNMKVDAGEAPPYNGALDCALKTVKAE-GPLALYKGFIP---TISRQ 356
Query: 168 ---AAIQFIVYEQLRK 180
+ F+ EQ+RK
Sbjct: 357 GPFTVVLFVTLEQVRK 404
Score = 32.0 bits (71), Expect = 0.10
Identities = 22/73 (30%), Positives = 31/73 (42%), Gaps = 3/73 (4%)
Frame = +3
Query: 1 MNELQWEYPVASLTTGFAFITFQYPLDVVQTR---LQVHDGRLFSQYPRYNNTTHAIFTI 57
MN+ + AS GF P+DV++TR ++V G + P YN
Sbjct: 129 MNDGIGTHVAASFAAGFVAAVASNPVDVIKTRVMNMKVDAG----EAPPYNGALDCALKT 296
Query: 58 ARKEGLKGLYAGF 70
+ EG LY GF
Sbjct: 297 VKAEGPLALYKGF 335
>TC8324 UP|Q8VWY1 (Q8VWY1) Mitochondrial phosphate transporter, complete
Length = 1605
Score = 37.7 bits (86), Expect = 0.002
Identities = 37/164 (22%), Positives = 68/164 (40%), Gaps = 16/164 (9%)
Frame = +3
Query: 13 LTTGFAFITFQYPLDVVQTRLQVHDGRLFSQYPRYNNTTHAIFTIARKEGLKGLYAGFPA 72
L+ G +T PLD+V+ +Q+ +Y + + + +++G+KG + G+
Sbjct: 354 LSCGLTHMTVT-PLDLVKCNMQIDP-------TKYKSISSGFGVLLKEQGVKGFFRGWVP 509
Query: 73 GLLGATISWSLLVFYYGIVKDRHA-----------RSKVEKSGA-----LMTVCLCANPA 116
LLG + + +Y K ++ ++ + +G+ + V LC P
Sbjct: 510 TLLGYSAQGACKFGFYEFFKKYYSDIAGPEYATKYKTLIYLAGSASAEVIADVALC--PF 683
Query: 117 FVVKTRLQLQTPLHHARPYSGLYDAFRTIKREEGFSAFYRGIVP 160
VK R+Q Q GL D + EG Y+G+VP
Sbjct: 684 EAVKVRVQTQPGFAR-----GLSDGLPKFVKAEGTLGLYKGLVP 800
Score = 30.4 bits (67), Expect = 0.30
Identities = 20/71 (28%), Positives = 31/71 (43%), Gaps = 1/71 (1%)
Frame = +3
Query: 111 LCANPAFVVKTRLQLQTPLHHARPYSGLYDAFRTIKREEGFSAFYRGIVPGFFLIS-QAA 169
+ P +VK +Q+ Y + F + +E+G F+RG VP S Q A
Sbjct: 375 MTVTPLDLVKCNMQIDPT-----KYKSISSGFGVLLKEQGVKGFFRGWVPTLLGYSAQGA 539
Query: 170 IQFIVYEQLRK 180
+F YE +K
Sbjct: 540 CKFGFYEFFKK 572
>BI420122
Length = 590
Score = 37.0 bits (84), Expect = 0.003
Identities = 32/124 (25%), Positives = 49/124 (38%), Gaps = 23/124 (18%)
Frame = +1
Query: 24 YPLDVVQTRLQVHDGRLFSQYPRYNNTTHAIFT----IARKEGLKGLYAGFPAGLLGATI 79
YPL V TR Q + P NN F + + EG + LY G L+G
Sbjct: 202 YPLQTVNTRQQT------DRDPNKNNKNLGTFQLMCQVVKHEGWERLYGGLTPSLVGTAA 363
Query: 80 SWSLLVFYYGIVKDRHARSKVEKSG----------------ALMTVC---LCANPAFVVK 120
S + ++Y I ++R +E+ A ++ C L NP ++V
Sbjct: 364 SQGVYYYFYQIFRNRAEAGVLEQKRLGIGDGSVGMMSSLFVAALSGCVNVLLTNPIWLVV 543
Query: 121 TRLQ 124
TR+Q
Sbjct: 544 TRMQ 555
>TC12076 similar to GB|AAL69531.1|18491125|AY074833 At2g47490/T30B22.21
{Arabidopsis thaliana;}, partial (20%)
Length = 506
Score = 35.8 bits (81), Expect = 0.007
Identities = 26/72 (36%), Positives = 37/72 (51%), Gaps = 7/72 (9%)
Frame = +2
Query: 11 ASLTTGFAFITFQYPLDVVQTRLQVHDGRLFSQYPRYNNTTHA-------IFTIARKEGL 63
A + G TF PLDV++TRLQV +L + N +T A + I +KEGL
Sbjct: 302 AGASAGVIAATFVCPLDVIKTRLQVGLPQLAN-----NGSTKAGRVIVGSLEQIFQKEGL 466
Query: 64 KGLYAGFPAGLL 75
+G+Y G +L
Sbjct: 467 RGMYRGLAPTVL 502
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.327 0.143 0.437
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 4,362,829
Number of Sequences: 28460
Number of extensions: 56843
Number of successful extensions: 445
Number of sequences better than 10.0: 72
Number of HSP's better than 10.0 without gapping: 413
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 435
length of query: 250
length of database: 4,897,600
effective HSP length: 88
effective length of query: 162
effective length of database: 2,393,120
effective search space: 387685440
effective search space used: 387685440
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.1 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.7 bits)
S2: 54 (25.4 bits)
Medicago: description of AC125480.1


