

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
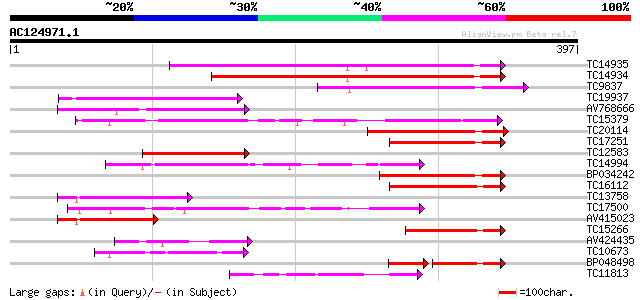
Query= AC124971.1 + phase: 0
(397 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
TC14935 similar to UP|Q9LSN8 (Q9LSN8) Protein phosphatase 2C-lik... 149 1e-36
TC14934 similar to UP|Q9LSN8 (Q9LSN8) Protein phosphatase 2C-lik... 148 2e-36
TC9837 similar to UP|Q9M3V0 (Q9M3V0) Protein phosphatase 2C (PP2... 95 2e-20
TC19937 weakly similar to GB|AAL87371.1|19548015|AY081718 AT4g33... 92 2e-19
AV768666 90 6e-19
TC15379 UP|Q9ZPL8 (Q9ZPL8) Protein phosphatase type 2C, complete 89 2e-18
TC20114 similar to UP|Q9LZ09 (Q9LZ09) Protein phosphatase-like p... 81 3e-16
TC17251 similar to UP|Q9M3V0 (Q9M3V0) Protein phosphatase 2C (PP... 73 7e-14
TC12583 similar to UP|Q9M3V0 (Q9M3V0) Protein phosphatase 2C (PP... 70 6e-13
TC14994 similar to UP|O82469 (O82469) Protein phosphatase-2C , p... 70 8e-13
BP034242 65 3e-11
TC16112 similar to GB|AAL87371.1|19548015|AY081718 AT4g33920/F17... 65 3e-11
TC13758 similar to GB|AAL87371.1|19548015|AY081718 AT4g33920/F17... 64 4e-11
TC17500 UP|Q9ZPL9 (Q9ZPL9) Nodule-enhanced protein phosphatase t... 60 5e-10
AV415023 57 7e-09
TC15266 similar to UP|Q9M3V0 (Q9M3V0) Protein phosphatase 2C (PP... 54 3e-08
AV424435 52 1e-07
TC10673 homologue to UP|Q8S8Z0 (Q8S8Z0) Protein phosphatase 2C, ... 51 4e-07
BP048498 35 5e-07
TC11813 similar to UP|Q8LJ68 (Q8LJ68) Protein phosphatase 2C-lik... 50 5e-07
>TC14935 similar to UP|Q9LSN8 (Q9LSN8) Protein phosphatase 2C-like protein,
partial (56%)
Length = 1089
Score = 149 bits (375), Expect = 1e-36
Identities = 89/241 (36%), Positives = 141/241 (57%), Gaps = 6/241 (2%)
Frame = +1
Query: 113 MEQAVDFIEKGFKEYVTNNIDDDGRVGSVGSCCLFGIIWGRTLFVANVGDSRAILGSSKG 172
+E+A E+G+ V+N+ + ++ + GSCCL G+I+ +TLFVAN GDSR +LG G
Sbjct: 7 IERAFLQTEEGYTALVSNSWNSRPQIANAGSCCLVGVIFQQTLFVANAGDSRVVLGKKVG 186
Query: 173 FFKRPHVVQLTVDHHVSHAAAREEIRNHITNDPFVLCKNRGSLRVKSLIEITRSIGDAYL 232
+QL+ +H+ + A REE+R NDP ++ G +VK +I+++RSIGD Y+
Sbjct: 187 NTDGVAAIQLSTEHNANLEAIREELRELHPNDPQIVVLKYGVWKVKGIIQVSRSIGDVYM 366
Query: 233 KWS--DPHPSFETFSRYE---ANVISEKPFTDRRDIDESDKFLIFASHGFWKLMTNSEAA 287
K + + P F E +++ P + +D FLIFAS G W+ ++N +A
Sbjct: 367 KDARFNREPLAAKFRLPEPMNMPIMTANPTILSHSLQPNDLFLIFASDGLWEHLSNEKAV 546
Query: 288 DIVYNNSQDGISKRLVRAAL-EKAINDIITYCNLQNLKAGNGLLGRRHYYDDVTVIVIFL 346
DIV +N + G +K LV+AAL E A + Y +L+ + RRH++DD++VIV+FL
Sbjct: 547 DIVNSNPRAGSAKILVKAALHEAASKREMRYSDLRKIDK----KVRRHFHDDISVIVLFL 714
Query: 347 N 347
N
Sbjct: 715 N 717
>TC14934 similar to UP|Q9LSN8 (Q9LSN8) Protein phosphatase 2C-like protein,
partial (57%)
Length = 932
Score = 148 bits (373), Expect = 2e-36
Identities = 82/212 (38%), Positives = 130/212 (60%), Gaps = 6/212 (2%)
Frame = +1
Query: 142 GSCCLFGIIWGRTLFVANVGDSRAILGSSKGFFKRPHVVQLTVDHHVSHAAAREEIRNHI 201
G+CCL G+I+ +TLFVAN+GDSR +LG G +QL+ +H+ + R+E++
Sbjct: 10 GTCCLVGVIFQQTLFVANLGDSRCVLGKKVGNTGGVAAIQLSTEHNANLEDIRQELKELH 189
Query: 202 TNDPFVLCKNRGSLRVKSLIEITRSIGDAYLKWS----DP-HPSFETFSRYEANVISEKP 256
+DP ++ G RVK +I+++RSIGD+Y+K + +P +P F ++S P
Sbjct: 190 PDDPQIVVLKHGVWRVKGIIQVSRSIGDSYMKHAQFNREPINPKFRLPEPMNMPIMSANP 369
Query: 257 FTDRRDIDESDKFLIFASHGFWKLMTNSEAADIVYNNSQDGISKRLVRAALEKAIND-II 315
+ SD FLIFAS G W+ ++N +A DIV++N + G +KRLV+AAL++A +
Sbjct: 370 TILSHPLQPSDSFLIFASDGLWEHLSNEQAVDIVHSNPRAGSAKRLVKAALQEAARKREM 549
Query: 316 TYCNLQNLKAGNGLLGRRHYYDDVTVIVIFLN 347
Y +L+ + RRH++DD+TVIV+FLN
Sbjct: 550 RYSDLRKIDK----KVRRHFHDDITVIVLFLN 633
>TC9837 similar to UP|Q9M3V0 (Q9M3V0) Protein phosphatase 2C (PP2C) ,
partial (43%)
Length = 727
Score = 95.1 bits (235), Expect = 2e-20
Identities = 53/154 (34%), Positives = 91/154 (58%), Gaps = 6/154 (3%)
Frame = +3
Query: 216 RVKSLIEITRSIGDAYLKWSD-----PHPSFETFSRYEANVISEKPFTDRRDIDESDKFL 270
RVK +I+I+RSIGD YLK ++ + F ++ ++S +P + D+F+
Sbjct: 15 RVKGIIQISRSIGDVYLKKAEFNREPLYAKFRLREPFKRPILSSEPSITVHQLQPHDQFI 194
Query: 271 IFASHGFWKLMTNSEAADIVYNNSQDGISKRLVRAALEKAIND-IITYCNLQNLKAGNGL 329
IFAS G W+ ++N EA DIV N+ + G ++RLV+ AL++A + Y +L+ + G
Sbjct: 195 IFASDGLWEHLSNQEAVDIVQNSPRSGSARRLVKTALQEAAKKREMRYSDLRRIDRG--- 365
Query: 330 LGRRHYYDDVTVIVIFLNKRSDTPTTEPEFQSTT 363
RRH++DD+TVIV++L+ + + +F S +
Sbjct: 366 -VRRHFHDDITVIVVYLDSNLMSRASTVKFPSVS 464
>TC19937 weakly similar to GB|AAL87371.1|19548015|AY081718
AT4g33920/F17I5_110 {Arabidopsis thaliana;}, partial
(30%)
Length = 593
Score = 91.7 bits (226), Expect = 2e-19
Identities = 53/129 (41%), Positives = 75/129 (58%)
Frame = +2
Query: 35 NLVWCEDRKENDYHCSIATSQSNTVMEDFYQVEFGKNSLFVGVYDGHKGLDAARFIRVCL 94
+L W D K +H SIA +Q+N+ +ED QV + FVGVYDGH G +A+RFIR L
Sbjct: 212 SLQWHMDSKP--HHYSIAVAQANSSLEDQGQVYISPSETFVGVYDGHGGPEASRFIRNNL 385
Query: 95 FPELSRLVTENKVVSEDIMEQAVDFIEKGFKEYVTNNIDDDGRVGSVGSCCLFGIIWGRT 154
F L + +E +SE ++++A E+ F V + ++ SVGSCCL I
Sbjct: 386 FSFLHKFASEEGGLSEQVIKKAFSATEEEFLHLVKQSWTKRPQIASVGSCCLLXAISKGV 565
Query: 155 LFVANVGDS 163
L+VAN+GDS
Sbjct: 566 LYVANLGDS 592
>AV768666
Length = 463
Score = 90.1 bits (222), Expect = 6e-19
Identities = 53/144 (36%), Positives = 80/144 (54%), Gaps = 9/144 (6%)
Frame = +1
Query: 34 DNLVWCEDR-KENDYHCSIATSQSNTVMEDFYQVEFGKNSL--------FVGVYDGHKGL 84
D L+W +D K + S+A Q+N ++ED +E G S FVGVYDGH G
Sbjct: 7 DGLLWYKDSGKHLNGEFSMAVVQANNLLEDQSYLESGSLSTSDSGPYGTFVGVYDGHGGP 186
Query: 85 DAARFIRVCLFPELSRLVTENKVVSEDIMEQAVDFIEKGFKEYVTNNIDDDGRVGSVGSC 144
+ +R+I L R + + + +S D++ +A+ E GF V ++ +VGSC
Sbjct: 187 ETSRYIN----EHLVRHLKKQQSMSVDVIRKAIQATEDGFMSLVAKQWSMKPQIAAVGSC 354
Query: 145 CLFGIIWGRTLFVANVGDSRAILG 168
CL G+I TL++AN+GDSRA+LG
Sbjct: 355 CLIGVICNGTLYIANLGDSRAVLG 426
>TC15379 UP|Q9ZPL8 (Q9ZPL8) Protein phosphatase type 2C, complete
Length = 1338
Score = 88.6 bits (218), Expect = 2e-18
Identities = 87/312 (27%), Positives = 142/312 (44%), Gaps = 13/312 (4%)
Frame = +1
Query: 47 YHCSIATSQSNTVMEDFYQVEF---GKNSL-FVGVYDGHKGLDAARFIRVCLFPELSRLV 102
YH + +S+ MED+ +F N L ++DGH G + +++ LF
Sbjct: 331 YH--LVKGRSDHAMEDYVVAQFKTVDNNELGLFAIFDGHSGHNVPDYLQSNLF------- 483
Query: 103 TENKVVSEDIMEQAVDFIEKGFKEYVTNNIDDDGRVGSVGSCCLFGIIWG-RTLFVANVG 161
+N + D + V+ ++K + + + ++ G +G GS + I+ + L VAN+G
Sbjct: 484 -DNILKEPDFWTKPVEAVKKAYVDTDSTILEKSGELGRGGSTAVTAILINCQKLVVANLG 660
Query: 162 DSRAILGSSKGFFKRPHVVQLTVDHHVSHAAAREEIRNH---ITNDPFVLCKNRGSL-RV 217
DSRA+L K + L+VDH A E+IRN ++N P G + RV
Sbjct: 661 DSRAVL------CKNGEAIPLSVDH--EPATESEDIRNRGGFVSNFP-------GDVPRV 795
Query: 218 KSLIEITRSIGDAYLK---WSDPHPSFETFSRYEANVISEKPFTDRRDIDESDKFLIFAS 274
+ ++R+ GD LK S+PH + E ID+ +F+I AS
Sbjct: 796 DGQLAVSRAFGDKSLKKHLSSEPHVTVEL-------------------IDDDAEFIILAS 918
Query: 275 HGFWKLMTNSEAADIVYN-NSQDGISKRLVRAALEKAINDIITYCNLQNLKAGNGLLGRR 333
G WK+M+N EA D + N +K L AL++ +D I+ C + L+ + G
Sbjct: 919 DGLWKVMSNQEAVDAIRNVKDARSAAKNLTEEALKRNSSDDIS-CVVVRLQ*SHNSRGSF 1095
Query: 334 HYYDDVTVIVIF 345
YY +VI F
Sbjct: 1096FYYKLGSVI*YF 1131
>TC20114 similar to UP|Q9LZ09 (Q9LZ09) Protein phosphatase-like protein,
partial (33%)
Length = 587
Score = 81.3 bits (199), Expect = 3e-16
Identities = 43/100 (43%), Positives = 67/100 (67%), Gaps = 1/100 (1%)
Frame = +1
Query: 251 VISEKPFTDRRDIDESDKFLIFASHGFWKLMTNSEAADIVYNNSQDGISKRLVRAAL-EK 309
++S +P + +D+FLIFAS G W+ ++N EA +IV NN ++GI++RLV+AAL E
Sbjct: 43 ILSWEPSITTHKLHSNDQFLIFASDGLWEQLSNQEAVNIVSNNPRNGIARRLVKAALREA 222
Query: 310 AINDIITYCNLQNLKAGNGLLGRRHYYDDVTVIVIFLNKR 349
A + +LQ ++ G RRH++DD+TVIV+FLN +
Sbjct: 223 AKKREMRVSDLQKIEQG----VRRHFHDDITVIVVFLNPK 330
>TC17251 similar to UP|Q9M3V0 (Q9M3V0) Protein phosphatase 2C (PP2C) ,
partial (30%)
Length = 715
Score = 73.2 bits (178), Expect = 7e-14
Identities = 38/82 (46%), Positives = 55/82 (66%), Gaps = 1/82 (1%)
Frame = +3
Query: 267 DKFLIFASHGFWKLMTNSEAADIVYNNSQDGISKRLVRAALEKAIND-IITYCNLQNLKA 325
D+F+IFAS G W+ TN +A DIV N+ GI+KRLV+ AL++A + Y +L +
Sbjct: 6 DQFIIFASDGLWEHFTNQDAVDIVQNHPHHGIAKRLVKMALQEAAKKREMRYSDLNKIDR 185
Query: 326 GNGLLGRRHYYDDVTVIVIFLN 347
G RRH++DD+TVIV+FL+
Sbjct: 186 G----VRRHFHDDITVIVVFLD 239
>TC12583 similar to UP|Q9M3V0 (Q9M3V0) Protein phosphatase 2C (PP2C) ,
partial (23%)
Length = 277
Score = 70.1 bits (170), Expect = 6e-13
Identities = 33/75 (44%), Positives = 49/75 (65%)
Frame = +2
Query: 94 LFPELSRLVTENKVVSEDIMEQAVDFIEKGFKEYVTNNIDDDGRVGSVGSCCLFGIIWGR 153
LF L R +E+K +S +++ +A E+GF VT + ++ SVGSCCL G+I G
Sbjct: 8 LFQHLKRFASEHKSMSVEVIRKAYQATEEGFLSVVTKLWPMNAQIASVGSCCLVGVICGG 187
Query: 154 TLFVANVGDSRAILG 168
L++AN+GDSRA+LG
Sbjct: 188 VLYIANLGDSRAVLG 232
>TC14994 similar to UP|O82469 (O82469) Protein phosphatase-2C , partial
(89%)
Length = 1672
Score = 69.7 bits (169), Expect = 8e-13
Identities = 63/230 (27%), Positives = 104/230 (44%), Gaps = 7/230 (3%)
Frame = +3
Query: 68 FGKNSLFVGVYDGHKGLDAARFIR----VCLFPELSRLVTENKVVSEDIMEQAVDFIEKG 123
F S F GV+DGH G +AA +IR F ++S + V + +++ + + K
Sbjct: 594 FPNPSAFYGVFDGHGGPEAAAYIRKNVIKFFFEDVS--FPQTSEVDKVFLQEVENSLRKA 767
Query: 124 FKEYVTNNIDDDGRVGSVGSCCLFGIIWGRTLFVANVGDSRAILGSSKGFFKRPHVVQLT 183
F + DD S G+ L I+GR L VAN GD RA+L S KG + ++
Sbjct: 768 FLLADSALADDCSVNTSSGTTALTAFIFGRLLMVANAGDCRAVL-SRKG-----EAIDMS 929
Query: 184 VDHHVSHAAAR---EEIRNHITNDPFVLCKNRGSLRVKSLIEITRSIGDAYLKWSDPHPS 240
DH + + R E++ +I + + ++ +TR++GD +K PS
Sbjct: 930 EDHRPIYPSERRRVEDLGGYIEDG-----------YLNGVLSVTRALGDWDMKLPKGAPS 1076
Query: 241 FETFSRYEANVISEKPFTDRRDIDESDKFLIFASHGFWKLMTNSEAADIV 290
+I+E F + + E D+FLI G W +M++ A +V
Sbjct: 1077---------PLIAEPEFR-QMVLTEDDEFLIIGCDGIWDVMSSQHAVSLV 1196
>BP034242
Length = 539
Score = 64.7 bits (156), Expect = 3e-11
Identities = 37/89 (41%), Positives = 58/89 (64%), Gaps = 1/89 (1%)
Frame = -1
Query: 260 RRDIDESDKFLIFASHGFWKLMTNSEAADIVYNNSQDGISKRLVRAALEKAI-NDIITYC 318
+R + D FLIFAS G W +T+ A +I+ + + GI+KRLVRAALE+A + Y
Sbjct: 527 KRKLKVEDLFLIFASDGLWDHLTDEAAVEIISRSPRTGIAKRLVRAALEEAAKKNGKRYE 348
Query: 319 NLQNLKAGNGLLGRRHYYDDVTVIVIFLN 347
+L+ ++ G RR ++DD+TVIV++L+
Sbjct: 347 DLKRIEKG----VRRLFHDDITVIVLYLD 273
>TC16112 similar to GB|AAL87371.1|19548015|AY081718 AT4g33920/F17I5_110
{Arabidopsis thaliana;}, partial (21%)
Length = 612
Score = 64.7 bits (156), Expect = 3e-11
Identities = 35/82 (42%), Positives = 54/82 (65%), Gaps = 1/82 (1%)
Frame = +3
Query: 267 DKFLIFASHGFWKLMTNSEAADIVYNNSQDGISKRLVRAALEKAIND-IITYCNLQNLKA 325
D FLIFAS G W +++ A ++V+ + GI+KRLVRAAL++A + Y ++ +
Sbjct: 3 DLFLIFASDGLWDQLSDEAAVEMVFKYPRAGIAKRLVRAALQEAAKKREMRYNDIMKIDK 182
Query: 326 GNGLLGRRHYYDDVTVIVIFLN 347
G RRH++DD+TVIVI+L+
Sbjct: 183 GI----RRHFHDDITVIVIYLD 236
>TC13758 similar to GB|AAL87371.1|19548015|AY081718 AT4g33920/F17I5_110
{Arabidopsis thaliana;}, partial (28%)
Length = 466
Score = 63.9 bits (154), Expect = 4e-11
Identities = 38/98 (38%), Positives = 57/98 (57%), Gaps = 3/98 (3%)
Frame = +2
Query: 34 DNLVWCEDRKEN---DYHCSIATSQSNTVMEDFYQVEFGKNSLFVGVYDGHKGLDAARFI 90
D L+W D K + D+ SIA +Q+N+ +ED QV ++ +VGVYDGH G A+RF+
Sbjct: 164 DGLLWHTDLKPHASGDF--SIAVAQANSSLEDQSQVFTSPSATYVGVYDGHGGPQASRFV 337
Query: 91 RVCLFPELSRLVTENKVVSEDIMEQAVDFIEKGFKEYV 128
LFP L + TE +S D++++A E+ F V
Sbjct: 338 NNHLFPYLHKFATEQGGLSVDVIKKAFSATEEEFLHLV 451
>TC17500 UP|Q9ZPL9 (Q9ZPL9) Nodule-enhanced protein phosphatase type 2C,
complete
Length = 1325
Score = 60.5 bits (145), Expect = 5e-10
Identities = 68/257 (26%), Positives = 108/257 (41%), Gaps = 7/257 (2%)
Frame = +3
Query: 41 DRKENDY---HCSIATSQSNTVMEDFYQVEFG--KNSLFVGVYDGHKGLDAARFIRVCLF 95
+RK +D + S++ S MED VE G + V+DGH G A R L+
Sbjct: 366 ERKRDDGVLPYGSVSVVGSRKEMEDAVSVETGCVTKCDYFAVFDGHGGAQVAEACRERLY 545
Query: 96 PELSRLVTENKVVSEDIMEQAVDFIE--KGFKEYVTNNIDDDGRVGSVGSCCLFGIIWGR 153
RLV E + +E+ VD+ E +G + + + + +VGS + ++
Sbjct: 546 ----RLVAEEVERCGNGVEE-VDWEEVMEGCFRNMDGEVAGNAALRTVGSTAVVAVVAAA 710
Query: 154 TLFVANVGDSRAILGSSKGFFKRPHVVQLTVDHHVSHAAAREEIRNHITNDPFVLCKNRG 213
+ +AN GD RA+LG + V L+ DH E +R + N
Sbjct: 711 EVVIANCGDCRAVLG------RGGEAVDLSSDHKPDR--PDELMRIEEAGGKVI---NWN 857
Query: 214 SLRVKSLIEITRSIGDAYLKWSDPHPSFETFSRYEANVISEKPFTDRRDIDESDKFLIFA 273
RV ++ +RSIGD YL+ P+ + KP D+FLI A
Sbjct: 858 GQRVLGVLATSRSIGDQYLR---PY-------------VISKPEVTVTKRSSKDEFLILA 989
Query: 274 SHGFWKLMTNSEAADIV 290
S G W ++++ A +V
Sbjct: 990 SDGLWDVISSEMACQVV 1040
>AV415023
Length = 385
Score = 56.6 bits (135), Expect = 7e-09
Identities = 32/74 (43%), Positives = 45/74 (60%), Gaps = 3/74 (4%)
Frame = +2
Query: 34 DNLVWCEDRKEN---DYHCSIATSQSNTVMEDFYQVEFGKNSLFVGVYDGHKGLDAARFI 90
D L+W D K + D+ SIA +Q+N+ +ED QV ++ +VGVYDGH G A+RF+
Sbjct: 161 DGLLWHTDLKPHASGDF--SIAVAQANSSLEDQSQVFTSPSATYVGVYDGHSGPQASRFV 334
Query: 91 RVCLFPELSRLVTE 104
LFP L + TE
Sbjct: 335 NNHLFPYLHKFATE 376
>TC15266 similar to UP|Q9M3V0 (Q9M3V0) Protein phosphatase 2C (PP2C) ,
partial (25%)
Length = 725
Score = 54.3 bits (129), Expect = 3e-08
Identities = 26/71 (36%), Positives = 47/71 (65%), Gaps = 1/71 (1%)
Frame = +1
Query: 278 WKLMTNSEAADIVYNNSQDGISKRLVRAALEKAINDI-ITYCNLQNLKAGNGLLGRRHYY 336
W+ ++N +A DIV N+ G ++RL++ AL++A + Y +L+ + G RRH++
Sbjct: 1 WEHLSNQDAVDIVQNHPHHGSARRLIKTALQEAAKKREMRYSDLKKIDRGV----RRHFH 168
Query: 337 DDVTVIVIFLN 347
DD+TV+V+FL+
Sbjct: 169 DDITVVVVFLD 201
>AV424435
Length = 280
Score = 52.4 bits (124), Expect = 1e-07
Identities = 37/105 (35%), Positives = 51/105 (48%), Gaps = 8/105 (7%)
Frame = +1
Query: 74 FVGVYDGHKGLDAARFIRVCLFPELSRLVTEN--------KVVSEDIMEQAVDFIEKGFK 125
F GV+DGH G AA+F+R L R++ E+ KVV++ +E +F +
Sbjct: 7 FYGVFDGHGGKSAAQFVR----DHLPRVIVEDADFPLELEKVVTKSFLETDAEFAKTCSS 174
Query: 126 EYVTNNIDDDGRVGSVGSCCLFGIIWGRTLFVANVGDSRAILGSS 170
E S G+ L II GR+L VAN GD RA+L S
Sbjct: 175 E-------------SSGTTALTAIILGRSLLVANAGDCRAVLSRS 270
>TC10673 homologue to UP|Q8S8Z0 (Q8S8Z0) Protein phosphatase 2C, partial
(48%)
Length = 669
Score = 50.8 bits (120), Expect = 4e-07
Identities = 37/112 (33%), Positives = 55/112 (49%), Gaps = 4/112 (3%)
Frame = +3
Query: 60 MEDFYQVEF----GKNSLFVGVYDGHKGLDAARFIRVCLFPELSRLVTENKVVSEDIMEQ 115
MEDFY+ G+ GV+DGH G AA +++ LF S L++ K +S D
Sbjct: 348 MEDFYETRIDGVEGEVVGLFGVFDGHGGARAAEYVKQNLF---SNLISHPKFIS-DTKSA 515
Query: 116 AVDFIEKGFKEYVTNNIDDDGRVGSVGSCCLFGIIWGRTLFVANVGDSRAIL 167
D E++ + + + GS S I+ G L VANVGDS+A++
Sbjct: 516 IADAYNHTDSEFLKSENNQNRDAGSTASTA---ILVGDRLLVANVGDSKAVI 662
>BP048498
Length = 470
Score = 35.4 bits (80), Expect(2) = 5e-07
Identities = 15/28 (53%), Positives = 21/28 (74%)
Frame = -1
Query: 266 SDKFLIFASHGFWKLMTNSEAADIVYNN 293
+D FLIFAS G W+ ++N +A DIV +N
Sbjct: 467 NDLFLIFASDGLWEHLSNEKAVDIVNSN 384
Score = 34.7 bits (78), Expect(2) = 5e-07
Identities = 21/52 (40%), Positives = 33/52 (63%), Gaps = 1/52 (1%)
Frame = -2
Query: 297 GISKRLVRAALEKAINDI-ITYCNLQNLKAGNGLLGRRHYYDDVTVIVIFLN 347
G +K LV+AAL +A + Y +L+ + RRH++DD++VIV+FLN
Sbjct: 301 GSAKILVKAALHEAARKREMRYSDLRKIDQKV----RRHFHDDISVIVLFLN 158
>TC11813 similar to UP|Q8LJ68 (Q8LJ68) Protein phosphatase 2C-like protein,
partial (36%)
Length = 917
Score = 50.4 bits (119), Expect = 5e-07
Identities = 44/136 (32%), Positives = 61/136 (44%), Gaps = 1/136 (0%)
Frame = +1
Query: 155 LFVANVGDSRAILGSSKGFFKRPHVVQLTVDHHVSHAAAREEIRNHITNDPFVLCKNRGS 214
+ V+N GDSRA+L K + L+VDH + RE+ I + + G
Sbjct: 34 IIVSNCGDSRAVLCRGK------EPLALSVDHKPN----REDEYARIEAAGGKVIQWNGH 183
Query: 215 LRVKSLIEITRSIGDAYLK-WSDPHPSFETFSRYEANVISEKPFTDRRDIDESDKFLIFA 273
RV ++ ++RSIGD YLK W P P R + D+ LI A
Sbjct: 184 -RVFGVLAMSRSIGDRYLKPWIIPEPEVRFLPRAK-----------------EDECLILA 309
Query: 274 SHGFWKLMTNSEAADI 289
S G W +MTN EA D+
Sbjct: 310 SDGLWDVMTNEEACDL 357
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.320 0.137 0.404
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 6,638,538
Number of Sequences: 28460
Number of extensions: 81519
Number of successful extensions: 696
Number of sequences better than 10.0: 77
Number of HSP's better than 10.0 without gapping: 602
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 632
length of query: 397
length of database: 4,897,600
effective HSP length: 92
effective length of query: 305
effective length of database: 2,279,280
effective search space: 695180400
effective search space used: 695180400
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 56 (26.2 bits)
Medicago: description of AC124971.1


