

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC124970.4 + phase: 0 /pseudo
(204 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
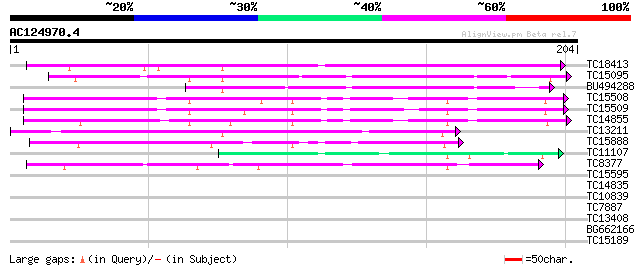
TC18413 similar to UP|P93378 (P93378) Tumor-related protein, par... 172 3e-44
TC15095 similar to UP|AAQ96377 (AAQ96377) Miraculin-like protein... 126 2e-30
BU494288 106 3e-24
TC15508 similar to UP|Q8RVX2 (Q8RVX2) Protease inhibitor precurs... 76 5e-15
TC15509 weakly similar to UP|Q8RVX2 (Q8RVX2) Protease inhibitor ... 71 1e-13
TC14855 similar to UP|Q8RVX2 (Q8RVX2) Protease inhibitor precurs... 69 7e-13
TC13211 weakly similar to UP|Q8LK19 (Q8LK19) Proteinase inhibito... 59 5e-10
TC15888 weakly similar to UP|Q8LK19 (Q8LK19) Proteinase inhibito... 53 3e-08
TC11107 52 7e-08
TC8377 similar to UP|Q9M3Z7 (Q9M3Z7) Alpha-fucosidase , partial... 47 2e-06
TC15595 homologue to UP|Q8L784 (Q8L784) Cytoplasmic aconitate hy... 28 1.1
TC14835 similar to PIR|T46895|T46895 acyl-CoA dehydrogenase [va... 27 1.9
TC10839 27 1.9
TC7887 similar to UP|Q9ZWQ8 (Q9ZWQ8) Homolog to plastid-lipid-as... 27 3.2
TC13408 25 7.2
BG662166 25 7.2
TC15189 similar to UP|TO34_PEA (Q41009) Translocase of chloropla... 25 9.4
>TC18413 similar to UP|P93378 (P93378) Tumor-related protein, partial (41%)
Length = 740
Score = 172 bits (437), Expect = 3e-44
Identities = 93/207 (44%), Positives = 125/207 (59%), Gaps = 13/207 (6%)
Frame = -3
Query: 7 LLFLLLTLFTTKPL--QGAEQPEEVRDTSGNLVRNSINYFILP-SSIQC--------GTR 55
L F +L T+P +G PE+V DT G VR + Y+I P + C G+
Sbjct: 699 LSFAILFALCTQPFLGKGDASPEQVVDTQGKNVRAGMGYYIRPVPTTPCDGRGPCVVGSG 520
Query: 56 CEMALLNTNKTCPLDVVEEE--EAMQFSFVPFNFKKGVIRVSTDLNVIHSFPTNCSTSSV 113
+ + NKTCPL+VV E +F P N KKGVIRVSTD+N+ + T C S
Sbjct: 519 FVLIARSPNKTCPLNVVVVEGFRGQAVTFTPVNPKKGVIRVSTDVNINVTLETACEES-- 346
Query: 114 TVWKVDKVDVATSQRFVTTGGVQGNPGRETVDNWFKIERFESGYKLVFCPTVCRECEVVC 173
TVWK+D D + FVTTGGV GNPG++T+ NWFKIE++E YKLVFCPTVC C+ +C
Sbjct: 345 TVWKLDDFDASAGHWFVTTGGVVGNPGKDTISNWFKIEKYEEDYKLVFCPTVCDTCKPLC 166
Query: 174 KDIGIFLDENRNTRFVLSDFPFGVKFQ 200
+++G+F+D N N R L+D + V+FQ
Sbjct: 165 RNVGVFMDSNGNRRVALTDESYKVRFQ 85
>TC15095 similar to UP|AAQ96377 (AAQ96377) Miraculin-like protein
(Fragment), partial (42%)
Length = 979
Score = 126 bits (317), Expect = 2e-30
Identities = 80/201 (39%), Positives = 115/201 (56%), Gaps = 13/201 (6%)
Frame = +3
Query: 15 FTTKPLQG---AEQPEEVRDTSGNLVRNSINYFILPSSIQCGTRCEMALLNT----NKTC 67
FTT+ L G A PE V D SG +R + Y+ILP + G + L ++ N TC
Sbjct: 228 FTTEFLIGIASAAAPEPVLDISGQKLRTGVKYYILP--VLRGKGGGLTLTSSGNINNNTC 401
Query: 68 PLDVVEEE----EAMQFSFVPFNFKKGVIRVSTDLNVIHSFPTNCSTSSVTVWKVDKVDV 123
PL VV+E+ + +F P+N K GVI STDLN+ S TN + + VWK+ K
Sbjct: 402 PLHVVQEKLEVLKGQSVTFTPYNAKGGVILTSTDLNIKSSL-TNTTCAKAPVWKLLKE-- 572
Query: 124 ATSQRFVTTGGVQGNPGRETVDNWFKIERFESGYKLVFCPTVCRECEVVCKDIGIFLDEN 183
+ F+ TGGV+GNPG T+ NWFKIE+ + Y FCP+VC+ C+ +C+++GI+ D
Sbjct: 573 LSGVWFLATGGVEGNPGMATISNWFKIEKADKDYVFSFCPSVCK-CQTLCRELGIY-DYG 746
Query: 184 RNTRFVLSDF--PFGVKFQRA 202
+ LSD PF + F++A
Sbjct: 747 GDKHLSLSDQVPPFRIMFKQA 809
>BU494288
Length = 438
Score = 106 bits (264), Expect = 3e-24
Identities = 59/137 (43%), Positives = 79/137 (57%), Gaps = 4/137 (2%)
Frame = +3
Query: 64 NKTCPLDVVEEE----EAMQFSFVPFNFKKGVIRVSTDLNVIHSFPTNCSTSSVTVWKVD 119
NK+CP +VV+ + +F P+N K GVI STDLN I SFP VWK+
Sbjct: 30 NKSCPPNVVQAKLEVWNGTPVTFTPYNAKDGVILTSTDLN-IKSFPIKSPCGPSFVWKLQ 206
Query: 120 KVDVATSQRFVTTGGVQGNPGRETVDNWFKIERFESGYKLVFCPTVCRECEVVCKDIGIF 179
K T F+ TGGV+GNPG +T+ NWFKIE+ Y L FCP+VC+ C +C+++GI+
Sbjct: 207 KE--LTGVWFLVTGGVEGNPGADTIVNWFKIEKAGKDYVLSFCPSVCK-CNTLCRELGIY 377
Query: 180 LDENRNTRFVLSDFPFG 196
+D D PFG
Sbjct: 378 IDG--------KDKPFG 404
>TC15508 similar to UP|Q8RVX2 (Q8RVX2) Protease inhibitor precursor, partial
(66%)
Length = 877
Score = 75.9 bits (185), Expect = 5e-15
Identities = 68/214 (31%), Positives = 103/214 (47%), Gaps = 18/214 (8%)
Frame = +3
Query: 6 PLLFLLLTLFTTKPLQGAEQPEEVRDTSGNLVRNSINYFILPSSIQCGTRCEMALLNT-N 64
PL FL L + T+PL E V D GN + + Y++ P S + G +AL +T N
Sbjct: 51 PLCFLSLLVLNTQPLLADEPEPAVVDMQGNPLLPGVQYYVRPVSAEKG---GLALGHTRN 221
Query: 65 KTCPLDVVEEEEAMQFSFVPFNFKK----GVIRVSTDLNV-IHSFPTNCSTSSVTVWKVD 119
KTCPLD++ + ++ S F+ G I +STDL V + + C VW++
Sbjct: 222 KTCPLDIILDPSSIIGSPTVFHTSSSSDVGYIPISTDLTVEMPILGSPCKEPK--VWRLA 395
Query: 120 KVDVATSQRFVTTGGVQGNPGRETVDNWFKIERFESG---------YKLVFCPTVCRECE 170
K V + FV+T GV GN + + FKI RFE Y FCP+V
Sbjct: 396 K--VGSDFWFVSTQGVPGN-----LVSKFKIVRFEERDNHANEGEIYSFSFCPSV---YG 545
Query: 171 VVCKDIGIFLDENRNTRFVLS---DFPFGVKFQR 201
V+C +GI++D++ + P+ V+FQ+
Sbjct: 546 VICAPVGIYVDDDGTQVMAVGAGVGKPYHVRFQK 647
>TC15509 weakly similar to UP|Q8RVX2 (Q8RVX2) Protease inhibitor precursor,
partial (77%)
Length = 797
Score = 70.9 bits (172), Expect = 1e-13
Identities = 63/213 (29%), Positives = 97/213 (44%), Gaps = 17/213 (7%)
Frame = +1
Query: 6 PLLFLLLTLFTTKPLQGAEQPEEVRDTSGNLVRNSINYFILPSSIQCGTRCEMALLNT-N 64
PL FL L + T+PL E D GN + + Y++ P S + G +AL +T N
Sbjct: 28 PLCFLSLLVLNTQPLLADEPEPAAVDMQGNPILPGVKYYVRPVSAENG---GLALGHTRN 198
Query: 65 KTCPLDVVEEEEAMQFSFV---PFNFKKGVIRVSTDLNV-IHSFPTNCSTSSVTVWKVDK 120
KTCPLD++ + ++ V + G I +STDL V + + C VW++ K
Sbjct: 199 KTCPLDIILDPFSLGTPTVFQTSSSSDLGYIPISTDLTVEMPVLKSPCKEPK--VWRIAK 372
Query: 121 VDVATSQRFVTTGGVQGNPGRETVDNWFKIERFESG---------YKLVFCPTVCRECEV 171
FV+T GV GN FKI RFE G Y +CP++
Sbjct: 373 --EGADFWFVSTQGVPGNDASR-----FKIVRFEEGDNHANEGEIYSFSYCPSI---WGY 522
Query: 172 VCKDIGIFLDENRNTRFVLS---DFPFGVKFQR 201
+C +GI++D++ + P+ V+FQ+
Sbjct: 523 ICAPVGIYVDDDGTQVMAVGAGVGKPYHVRFQK 621
>TC14855 similar to UP|Q8RVX2 (Q8RVX2) Protease inhibitor precursor, partial
(90%)
Length = 1001
Score = 68.6 bits (166), Expect = 7e-13
Identities = 66/209 (31%), Positives = 96/209 (45%), Gaps = 12/209 (5%)
Frame = +1
Query: 6 PLLFLLLTLFTTKPLQGAE-QPEEVRDTSGNLVRNSINYFILPSSIQCGTRCEMALLNT- 63
PL FL L + T+PL AE +PE V D GN + + Y+ P G + L T
Sbjct: 67 PLFFLSLLILNTQPLLAAEPEPEPVVDKQGNPLEPGVGYYAWPLWADNGG---LTLGRTR 237
Query: 64 NKTCPLDVVEEEEAM--QFSFVPFNFKKGVIRVSTDLNV-IHSFPTNCSTSSVTVWKVDK 120
NKTCPLD++ + + F + G I TDL V + + C VW++ K
Sbjct: 238 NKTCPLDIIRDPSKIGTPIEFHVADSDLGYIPTLTDLTVEMPILGSRCQEPK--VWRLLK 411
Query: 121 VDVATSQRFVTTGGVQGNPGRETVDNWFKIERFESG-----YKLVFCPTVCRECEVVCKD 175
V + F++TGGV GN + + FKIER Y FCP+V V+C
Sbjct: 412 --VGSGFWFLSTGGVAGN-----LVSKFKIERLAGEHAYEIYSFKFCPSV---PGVLCAP 561
Query: 176 IGIFLDENRNTRFVLSD--FPFGVKFQRA 202
+G + D + + D + V+FQ+A
Sbjct: 562 VGTYEDADGTQVMAIGDDIETYYVRFQKA 648
>TC13211 weakly similar to UP|Q8LK19 (Q8LK19) Proteinase inhibitor 20,
partial (44%)
Length = 558
Score = 59.3 bits (142), Expect = 5e-10
Identities = 51/169 (30%), Positives = 78/169 (45%), Gaps = 7/169 (4%)
Frame = +1
Query: 1 MKLHFPLLFLLLTLFTTKPLQGAEQPEEVRDTSGNLVRNSINYFILPSSIQCGTRCEMAL 60
+ L F L L LF PL ++ PE+V+D GN + Y I+P+ +
Sbjct: 37 LSLCFFLFALTPYLF---PLAFSQDPEQVKDAYGNPLVPGRRYSIVPALRGSRSGGVSPD 207
Query: 61 LNTNKTCPLDVVEEE------EAMQFSFVPFNFKKGVIRVSTDLNVIHSFPTNCSTSSVT 114
N+TCPL VV+E + + FS G I T L++ C+ +S
Sbjct: 208 KTGNQTCPLTVVQEYFECRDGKPVIFSVSGITPGTGTILTGTHLDIAFVEKPECADTSKW 387
Query: 115 VWKVDKVDVATSQRFVTTGGVQGNPGRETVDNWFKIERFE-SGYKLVFC 162
V + D + V+ GGV+ +PG+ +D F I++ + GYKLVFC
Sbjct: 388 VVVGNLTDYPGAP--VSIGGVEDHPGQHILDGTFNIQKLDCGGYKLVFC 528
>TC15888 weakly similar to UP|Q8LK19 (Q8LK19) Proteinase inhibitor 20,
partial (27%)
Length = 611
Score = 53.1 bits (126), Expect = 3e-08
Identities = 52/169 (30%), Positives = 79/169 (45%), Gaps = 13/169 (7%)
Frame = +3
Query: 8 LFLLLTLFTTKPLQGA--EQPEEVRDTSGNLVRNSINYFILPSSIQCGTRCEMALLNTNK 65
L +LL FTT + A + P+ V+DT+G +R+ Y+I P+ G N+
Sbjct: 69 LSVLLFAFTTYIIPSAFSQVPDLVKDTNGIPLRSDGRYYIRPALRGPGGGGVSPGETGNQ 248
Query: 66 TCPLDV------VEEEEAMQFSFVPFNFKKGVIRVSTDLNV----IHSFPTNCSTSSVTV 115
TCP+ V VE ++F P G+I L + I+ F C+ S T
Sbjct: 249 TCPVTVLQHKSEVENGNGVKFEHYP---NIGIIYEGVPLTISFGDIYGF---CADS--TK 404
Query: 116 WKVDKVDVATSQRFVTTGGVQGNPGRETVDNWFKIERFES-GYKLVFCP 163
W V D ++V GG + +PG+E V F I++++ YKLVF P
Sbjct: 405 WVVVSDDF--PGKWVGIGGAEDHPGKEIVSGLFDIQKYDDVAYKLVFTP 545
>TC11107
Length = 841
Score = 52.0 bits (123), Expect = 7e-08
Identities = 42/142 (29%), Positives = 56/142 (38%), Gaps = 18/142 (12%)
Frame = +3
Query: 76 EAMQFSFVPFNFKKGVIRVSTDLNVIHSFPTNCSTSSVTVWKVDKVDVATSQRFVTTGGV 135
E F PF +++ D V+ S T C S T W V + + +R V TG
Sbjct: 21 EGFPVKFTPFAKGDKYVKLGKDFTVVFSAATTCVQS--TAWSVGSTEAKSGRRLVVTG-- 188
Query: 136 QGNPGRETVDNWFKIERFESG---YKLVFCPT-VCRECEVVCKDIGIFLDENRNTRFVL- 190
G+ G ++F+I E Y + +CPT VC C C GI L EN L
Sbjct: 189 -GDVGERAYGSYFRIVAVEGRAGIYNIQWCPTDVCNICRFRCGTAGI-LRENGKILLALD 362
Query: 191 -------------SDFPFGVKF 199
+ FP GVKF
Sbjct: 363 GGMLPVVFQKV*FAAFPLGVKF 428
>TC8377 similar to UP|Q9M3Z7 (Q9M3Z7) Alpha-fucosidase , partial (83%)
Length = 1014
Score = 47.0 bits (110), Expect = 2e-06
Identities = 49/197 (24%), Positives = 88/197 (43%), Gaps = 11/197 (5%)
Frame = +1
Query: 7 LLFLLLTLFTTK--PLQGAEQPEEVRDTSGNLVRNSINYFILPSSIQCGTRCEMALLNTN 64
L FL +TK + E+V D +G+ + Y+I P+ I+ + L T
Sbjct: 52 LCFLFFAFISTKLPTAFSSNDVEQVLDVNGDSIFPGGRYYIRPA-IRGPPGGGVRLGKTG 228
Query: 65 KT-CPLDVVEEEEAMQFSFVPFNFK------KGVIRVSTDLNVIHSFPTNCSTSSVTVWK 117
+ CP+ ++E ++ + +P F+ G+I T+L++ + +C S +
Sbjct: 229 DSDCPVTALQEYSEVK-NGIPVRFRIAAEVSTGIIFTGTELDIEFTVKPHCVESPKWIVF 405
Query: 118 VDKVDVATSQRFVTTGGVQGNPGRETVDNWFKIERFESG--YKLVFCPTVCRECEVVCKD 175
VD + V GG + +PG++T F I++ +G YKLVF C + C D
Sbjct: 406 VDN---EIGKACVGIGGAKDHPGKQTFSGTFNIQKNSAGFGYKLVF----CIKGSPTCLD 564
Query: 176 IGIFLDENRNTRFVLSD 192
IG + + R L++
Sbjct: 565 IGRYDNGEGGRRLNLTE 615
>TC15595 homologue to UP|Q8L784 (Q8L784) Cytoplasmic aconitate hydratase
(At2g05710), partial (26%)
Length = 1077
Score = 28.1 bits (61), Expect = 1.1
Identities = 15/47 (31%), Positives = 26/47 (54%)
Frame = -1
Query: 88 KKGVIRVSTDLNVIHSFPTNCSTSSVTVWKVDKVDVATSQRFVTTGG 134
+ G+ +ST L + S STSS+ WK+ ++ +A S+R + G
Sbjct: 612 RAGLRGISTCLEEVFSQSIIFSTSSLVTWKLSQLRMADSRRILMEYG 472
>TC14835 similar to PIR|T46895|T46895 acyl-CoA dehydrogenase [validated] -
Arabidopsis thaliana {Arabidopsis
thaliana;} , partial (31%)
Length = 741
Score = 27.3 bits (59), Expect = 1.9
Identities = 11/29 (37%), Positives = 18/29 (61%)
Frame = +2
Query: 35 NLVRNSINYFILPSSIQCGTRCEMALLNT 63
N + N+ +YF++ S+I G C M L N+
Sbjct: 599 NSISNASSYFLVLSTIHVGPYCHMCLSNS 685
>TC10839
Length = 769
Score = 27.3 bits (59), Expect = 1.9
Identities = 15/44 (34%), Positives = 25/44 (56%), Gaps = 3/44 (6%)
Frame = -1
Query: 78 MQFSFVPFNFKKGVIRVSTDLNVIHSF---PTNCSTSSVTVWKV 118
+QFS V ++++KG I+ + D H F P+ S SS ++ V
Sbjct: 643 LQFSCVIYSYRKGEIKKAVDWKFTHGFYK*PSLLSCSSASLLAV 512
>TC7887 similar to UP|Q9ZWQ8 (Q9ZWQ8) Homolog to plastid-lipid-associated
protein, partial (65%)
Length = 1004
Score = 26.6 bits (57), Expect = 3.2
Identities = 10/32 (31%), Positives = 18/32 (56%)
Frame = -2
Query: 155 SGYKLVFCPTVCRECEVVCKDIGIFLDENRNT 186
+ Y ++ CP +VC+ FLD+N++T
Sbjct: 697 TNYYIIICPKTKTTICLVCQKRAAFLDKNKHT 602
>TC13408
Length = 583
Score = 25.4 bits (54), Expect = 7.2
Identities = 10/20 (50%), Positives = 14/20 (70%)
Frame = -1
Query: 184 RNTRFVLSDFPFGVKFQRAC 203
R+T+FVL + FG+KF C
Sbjct: 577 RSTKFVLINCQFGLKFNLGC 518
>BG662166
Length = 331
Score = 25.4 bits (54), Expect = 7.2
Identities = 10/32 (31%), Positives = 16/32 (49%)
Frame = -1
Query: 156 GYKLVFCPTVCRECEVVCKDIGIFLDENRNTR 187
GY++ C + CR C + I + + N N R
Sbjct: 331 GYEITHCSSCCRRCIIQTLKICVLICCNSNKR 236
>TC15189 similar to UP|TO34_PEA (Q41009) Translocase of chloroplast 34 (34
kDa chloroplast outer envelope protein) (GTP-binding
protein OEP34) (GTP-binding protein IAP34), partial
(36%)
Length = 738
Score = 25.0 bits (53), Expect = 9.4
Identities = 11/26 (42%), Positives = 15/26 (57%)
Frame = +3
Query: 2 KLHFPLLFLLLTLFTTKPLQGAEQPE 27
KL PLL+ L F KP++G Q +
Sbjct: 222 KLWIPLLYALQYFFVIKPIEGLIQQD 299
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.324 0.139 0.432
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 4,036,848
Number of Sequences: 28460
Number of extensions: 56238
Number of successful extensions: 403
Number of sequences better than 10.0: 34
Number of HSP's better than 10.0 without gapping: 390
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 391
length of query: 204
length of database: 4,897,600
effective HSP length: 86
effective length of query: 118
effective length of database: 2,450,040
effective search space: 289104720
effective search space used: 289104720
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.0 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.6 bits)
S2: 53 (25.0 bits)
Medicago: description of AC124970.4


