

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC124967.9 - phase: 0
(122 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
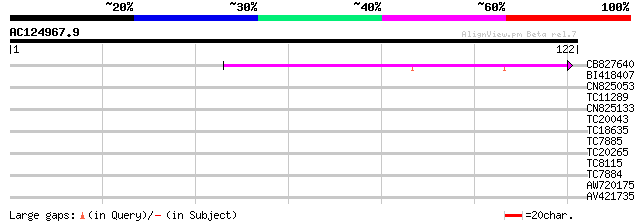
Score E
Sequences producing significant alignments: (bits) Value
CB827640 38 4e-04
BI418407 30 0.088
CN825053 29 0.20
TC11289 weakly similar to UP|Q9LUE5 (Q9LUE5) Emb|CAB86483.1, par... 29 0.20
CN825133 27 0.75
TC20043 similar to UP|Q940G7 (Q940G7) G protein-coupled receptor... 26 2.2
TC18635 similar to UP|Q9MA20 (Q9MA20) T5E21.13, partial (6%) 25 4.8
TC7885 similar to UP|RL4B_ARATH (Q9SF40) 60S ribosomal protein L... 24 6.3
TC20265 similar to UP|COL6_ARATH (Q9LQZ7) Zinc finger protein co... 24 8.3
TC8115 similar to PIR|A24727|A24727 phenylalanine ammonia-lyase ... 24 8.3
TC7884 similar to UP|RL4_PRUAR (Q9XF97) 60S ribosomal protein L4... 24 8.3
AW720175 24 8.3
AV421735 24 8.3
>CB827640
Length = 550
Score = 38.1 bits (87), Expect = 4e-04
Identities = 25/92 (27%), Positives = 44/92 (47%), Gaps = 17/92 (18%)
Frame = +1
Query: 47 TVFVGKSRRLYRVTSDVVDNPVFRELVERSRETEQQNDT---------------VNVVAC 91
TV VG+ +R++ V ++ FR L++ + Q D V V
Sbjct: 202 TVAVGREKRVFLVDPFILQENQFRILMDHTSMKMDQGDKEDHHHHLGFSTKQHGVVFVDV 381
Query: 92 EVVLFEHLLWMLEN--ADPQPESLDELVDFYA 121
+ +LF+H+LW++ N + +L E++DFYA
Sbjct: 382 DTILFQHMLWLVHNDYSSLCNLNLKEIIDFYA 477
>BI418407
Length = 422
Score = 30.4 bits (67), Expect = 0.088
Identities = 23/85 (27%), Positives = 41/85 (48%), Gaps = 14/85 (16%)
Frame = +2
Query: 35 DEDSSSSRSDLHTVFVGKSRRLYRVTSDVVDNPVFRELVERSRE----TEQQND------ 84
+E+ S SR V VGK +R++ V ++ FR L++ S E++ +
Sbjct: 164 EEEYSDSRV---MVVVGKEKRVFLVDPFILQENPFRLLMDISMRKVPAAEKKKNHFHHFT 334
Query: 85 ----TVNVVACEVVLFEHLLWMLEN 105
+ V + +LFEH+LW++ N
Sbjct: 335 TRKRKIIFVDVDDILFEHMLWLMNN 409
>CN825053
Length = 608
Score = 29.3 bits (64), Expect = 0.20
Identities = 12/29 (41%), Positives = 20/29 (68%)
Frame = +1
Query: 17 WNSFGNGKQSRHSISAVADEDSSSSRSDL 45
WN F + ++++ + VADE SS+S S+L
Sbjct: 109 WNGFTSSRKNQRVVLRVADERSSNSNSNL 195
>TC11289 weakly similar to UP|Q9LUE5 (Q9LUE5) Emb|CAB86483.1, partial (32%)
Length = 765
Score = 29.3 bits (64), Expect = 0.20
Identities = 24/98 (24%), Positives = 45/98 (45%), Gaps = 1/98 (1%)
Frame = +2
Query: 15 KKWNSFGNGKQSRHSISAVADEDSSSSRSDLHTVFVGKSRRLYRVTSDVVDNPVFRELVE 74
K W S G S + VA E S V+VG+ + + + ++ V++P+F+ L+E
Sbjct: 263 KSWPSRGK------STTVVAPEGCFS-------VYVGQQMQRFVIKTEYVNHPLFKMLLE 403
Query: 75 RSR-ETEQQNDTVNVVACEVVLFEHLLWMLENADPQPE 111
+ E + V+ C V +F +L ++ P+
Sbjct: 404 EAESEYGYSSQGPIVLPCNVDVFYKVLMEMDEETSTPD 517
>CN825133
Length = 711
Score = 27.3 bits (59), Expect = 0.75
Identities = 26/89 (29%), Positives = 40/89 (44%)
Frame = +3
Query: 19 SFGNGKQSRHSISAVADEDSSSSRSDLHTVFVGKSRRLYRVTSDVVDNPVFRELVERSRE 78
SFG+GK+ +H+I ++ E +DL K RRL N R+ V+RS
Sbjct: 336 SFGDGKEDQHAIESLTYE-----ITDLIKRSEKKLRRLSAAGPSEDSN--VRKNVQRSLA 494
Query: 79 TEQQNDTVNVVACEVVLFEHLLWMLENAD 107
T+ QN +V + + + L E D
Sbjct: 495 TDLQNLSVELRKKQSTYLKRLRQQKEGQD 581
>TC20043 similar to UP|Q940G7 (Q940G7) G protein-coupled receptor-like
protein, partial (15%)
Length = 574
Score = 25.8 bits (55), Expect = 2.2
Identities = 12/25 (48%), Positives = 15/25 (60%)
Frame = +1
Query: 21 GNGKQSRHSISAVADEDSSSSRSDL 45
G + RH S++ D DSS S SDL
Sbjct: 199 GGSDEGRHESSSLLDLDSSLSSSDL 273
>TC18635 similar to UP|Q9MA20 (Q9MA20) T5E21.13, partial (6%)
Length = 610
Score = 24.6 bits (52), Expect = 4.8
Identities = 10/20 (50%), Positives = 15/20 (75%)
Frame = -1
Query: 49 FVGKSRRLYRVTSDVVDNPV 68
F+G S +L +SDVVD+P+
Sbjct: 568 FLGYS*KLMEASSDVVDHPI 509
>TC7885 similar to UP|RL4B_ARATH (Q9SF40) 60S ribosomal protein L4-2 (L1),
complete
Length = 1554
Score = 24.3 bits (51), Expect = 6.3
Identities = 11/35 (31%), Positives = 20/35 (56%), Gaps = 1/35 (2%)
Frame = +3
Query: 1 MAKGGKLMKLKSALKKWNSFGNGKQSRHS-ISAVA 34
M +GG++ ++W+ N Q RH+ +SA+A
Sbjct: 372 MCRGGRMFAPTKIWRRWHRKINVNQKRHALVSAIA 476
>TC20265 similar to UP|COL6_ARATH (Q9LQZ7) Zinc finger protein constans-like
6, partial (32%)
Length = 623
Score = 23.9 bits (50), Expect = 8.3
Identities = 12/42 (28%), Positives = 23/42 (54%)
Frame = -2
Query: 10 LKSALKKWNSFGNGKQSRHSISAVADEDSSSSRSDLHTVFVG 51
L+++L W + + R + SAV + ++SS H +F+G
Sbjct: 235 LEASLLAWWTRWSQPSQRAASSAVQNTEASSLLQTSHWIFIG 110
>TC8115 similar to PIR|A24727|A24727 phenylalanine ammonia-lyase - kidney
bean (fragment) {Phaseolus vulgaris;} ,
partial (34%)
Length = 760
Score = 23.9 bits (50), Expect = 8.3
Identities = 8/15 (53%), Positives = 11/15 (73%)
Frame = +3
Query: 9 KLKSALKKWNSFGNG 23
K+KS + WN+ GNG
Sbjct: 459 KVKSLILSWNALGNG 503
>TC7884 similar to UP|RL4_PRUAR (Q9XF97) 60S ribosomal protein L4 (L1),
partial (97%)
Length = 1476
Score = 23.9 bits (50), Expect = 8.3
Identities = 11/35 (31%), Positives = 20/35 (56%), Gaps = 1/35 (2%)
Frame = +1
Query: 1 MAKGGKLMKLKSALKKWNSFGNGKQSRHS-ISAVA 34
M +GG++ ++W+ N Q RH+ +SA+A
Sbjct: 331 MCRGGRMFAPTRIWRRWHRKINVNQKRHALVSAIA 435
>AW720175
Length = 451
Score = 23.9 bits (50), Expect = 8.3
Identities = 17/57 (29%), Positives = 26/57 (44%), Gaps = 3/57 (5%)
Frame = +3
Query: 66 NPVFRELVERSRETEQQNDTVNVVACEVVLFEHLLWMLENADPQPESLDEL---VDF 119
NP + S E ++ N + + E+V+ W+L + QP DEL VDF
Sbjct: 84 NPAVEHQITVSAEVKE-NQGGDAIVSEMVIECESNWILSESQIQPMKEDELMLSVDF 251
>AV421735
Length = 286
Score = 23.9 bits (50), Expect = 8.3
Identities = 12/42 (28%), Positives = 23/42 (54%)
Frame = -2
Query: 10 LKSALKKWNSFGNGKQSRHSISAVADEDSSSSRSDLHTVFVG 51
L+++L W + + R + SAV + ++SS H +F+G
Sbjct: 252 LEASLLAWWTRWSQPSQRAASSAVQNTEASSLLQTSHWIFIG 127
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.315 0.130 0.371
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 1,965,126
Number of Sequences: 28460
Number of extensions: 20422
Number of successful extensions: 121
Number of sequences better than 10.0: 26
Number of HSP's better than 10.0 without gapping: 121
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 121
length of query: 122
length of database: 4,897,600
effective HSP length: 79
effective length of query: 43
effective length of database: 2,649,260
effective search space: 113918180
effective search space used: 113918180
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.6 bits)
S2: 49 (23.5 bits)
Medicago: description of AC124967.9


