

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
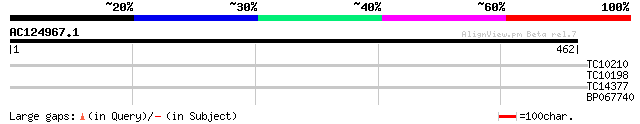
Query= AC124967.1 - phase: 0 /pseudo
(462 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
TC10210 29 1.9
TC10198 27 5.5
TC14377 similar to UP|PSQ2_ARATH (Q41932) Oxygen-evolving enhanc... 27 7.1
BP067740 27 9.3
>TC10210
Length = 765
Score = 28.9 bits (63), Expect = 1.9
Identities = 9/14 (64%), Positives = 9/14 (64%)
Frame = -1
Query: 307 SWWKSFCFDWYTDC 320
SWWK CF W T C
Sbjct: 753 SWWKPICFMWCTRC 712
>TC10198
Length = 644
Score = 27.3 bits (59), Expect = 5.5
Identities = 13/40 (32%), Positives = 19/40 (47%)
Frame = +2
Query: 326 HQRYLFL**YSFNCYYRHWCYSLFHCFRMCL*TGFDCV*Y 365
+Q +F + FN H LF+CF+ C F C+ Y
Sbjct: 338 YQTLIFRNIFLFNFMKIHCSMFLFYCFKTCKFPSFSCLLY 457
>TC14377 similar to UP|PSQ2_ARATH (Q41932) Oxygen-evolving enhancer protein
3-2, chloroplast precursor (OEE3) (16 kDa subunit of
oxygen evolving system of photosystem II) (OEC 16 kDa
subunit), partial (96%)
Length = 936
Score = 26.9 bits (58), Expect = 7.1
Identities = 11/21 (52%), Positives = 11/21 (52%)
Frame = -1
Query: 132 CC*VCCKICGACKVLSPLCPG 152
CC CC CG VLS L G
Sbjct: 216 CCCCCCCYCGCSSVLSHLVGG 154
>BP067740
Length = 529
Score = 26.6 bits (57), Expect = 9.3
Identities = 7/16 (43%), Positives = 11/16 (68%)
Frame = -3
Query: 337 FNCYYRHWCYSLFHCF 352
F+C+ + WC F+CF
Sbjct: 83 FHCFLKKWCSLFFNCF 36
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.376 0.169 0.742
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 9,375,755
Number of Sequences: 28460
Number of extensions: 154728
Number of successful extensions: 3043
Number of sequences better than 10.0: 8
Number of HSP's better than 10.0 without gapping: 1198
Number of HSP's successfully gapped in prelim test: 94
Number of HSP's that attempted gapping in prelim test: 1702
Number of HSP's gapped (non-prelim): 1455
length of query: 462
length of database: 4,897,600
effective HSP length: 93
effective length of query: 369
effective length of database: 2,250,820
effective search space: 830552580
effective search space used: 830552580
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 13 ( 7.0 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 35 (21.5 bits)
S2: 57 (26.6 bits)
Medicago: description of AC124967.1


