

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC124966.3 + phase: 0
(470 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
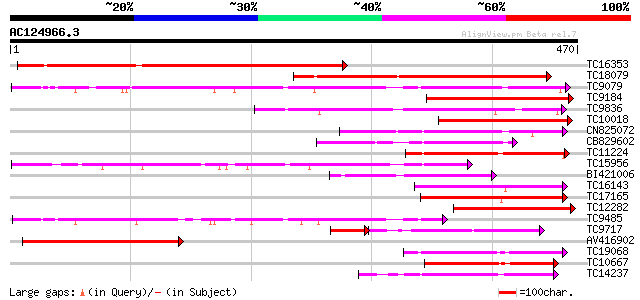
Score E
Sequences producing significant alignments: (bits) Value
TC16353 weakly similar to UP|HQGT_ARATH (Q9M156) Probable hydroq... 254 3e-68
TC18079 weakly similar to UP|UFO5_MANES (Q40287) Flavonol 3-O-gl... 206 7e-54
TC9079 weakly similar to UP|Q8W4G1 (Q8W4G1) UDP-glucose glucosyl... 171 3e-43
TC9184 weakly similar to UP|HQGT_RAUSE (Q9AR73) Hydroquinone glu... 167 3e-42
TC9836 weakly similar to UP|Q9LXV0 (Q9LXV0) Glucosyltransferase-... 166 8e-42
TC10018 weakly similar to UP|HQGT_RAUSE (Q9AR73) Hydroquinone gl... 155 1e-38
CN825072 130 4e-31
CB829602 124 3e-29
TC11224 weakly similar to UP|Q8S996 (Q8S996) Glucosyltransferase... 123 6e-29
TC15956 weakly similar to UP|LGT_CITUN (Q9MB73) Limonoid UDP-glu... 120 6e-28
BI421006 114 3e-26
TC16143 similar to UP|Q7XZD0 (Q7XZD0) Isoflavonoid glucosyltrans... 113 6e-26
TC17165 similar to UP|Q9SMG6 (Q9SMG6) Betanidin-5-O-glucosyltran... 112 1e-25
TC12282 similar to UP|Q8S996 (Q8S996) Glucosyltransferase-13 (Fr... 112 2e-25
TC9485 similar to UP|Q8S999 (Q8S999) Glucosyltransferase-10, par... 109 1e-24
TC9717 weakly similar to UP|Q9LNI1 (Q9LNI1) F6F3.22 protein, par... 89 2e-24
AV416902 107 5e-24
TC19068 weakly similar to UP|AAR06913 (AAR06913) UDP-glycosyltra... 105 2e-23
TC10667 similar to UP|Q9M051 (Q9M051) Glucuronosyl transferase-l... 103 8e-23
TC14237 weakly similar to UP|AAR06913 (AAR06913) UDP-glycosyltra... 102 1e-22
>TC16353 weakly similar to UP|HQGT_ARATH (Q9M156) Probable hydroquinone
glucosyltransferase (Arbutin synthase) , partial (52%)
Length = 885
Score = 254 bits (648), Expect = 3e-68
Identities = 125/274 (45%), Positives = 176/274 (63%)
Frame = +1
Query: 7 IAVVPGVGYSHLVPILQFSKRLVQLHPNFHVTCFIPTLGSPSNATKSILQTLPSNINHTF 66
+A++P G HL+P+++F+KRL+ LH N VT IPT PS A ++LQ+LP++I+H F
Sbjct: 73 VAMMPSPGMGHLIPMIEFAKRLI-LHHNLQVTFIIPTEAPPSKAQITVLQSLPNSISHIF 249
Query: 67 LPPVNPNDLPQGTTMESQILLTLTNSLPYLHQGLKSLAKEIPLVALVVDAFSVEVLNIGK 126
LPPV+ DLPQ +E +I LT+ SLP L + L+ AL+VD F + ++ +
Sbjct: 250 LPPVDLTDLPQNAMIEIRISLTVLRSLPSLRHAFRPLSA----TALLVDLFGTDAFDVAR 417
Query: 127 ELNMLSYIYFPSAATTLAWCFYLPKLDEETSCEYRDILEPIKIPGCVPLHGRDFLSIAQD 186
E YI+FPS AT L+ YLP+LD E CE+R++ EP+KIPG VP+HG D L QD
Sbjct: 418 EFGASPYIFFPSTATALSLFLYLPRLDREVHCEFRELAEPVKIPGSVPIHGSDLLDPVQD 597
Query: 187 RSSQAYKHFLPFVKLLSSADGVLVNSFLEIEMGPLSAMKEEGGDNPPVYPVGPIIETETK 246
R ++AYK L S ADG+L NSF+E+E G + +++E P VYPVGP+++ +
Sbjct: 598 RKNEAYKWVLHHATRYSQADGILENSFMELEPGAIKELQKEEPGKPRVYPVGPLVKVDGV 777
Query: 247 SGDDANGLECLAWLDKQQPCSVLYVSFGSGGTLS 280
G ECL+WLD+Q SVL+V FGSGGTL+
Sbjct: 778 QAGSGPGSECLSWLDEQPHGSVLFVCFGSGGTLT 879
>TC18079 weakly similar to UP|UFO5_MANES (Q40287) Flavonol
3-O-glucosyltransferase 5 (UDP-glucose flavonoid
3-O-glucosyltransferase 5) , partial (33%)
Length = 889
Score = 206 bits (524), Expect = 7e-54
Identities = 100/214 (46%), Positives = 149/214 (68%)
Frame = +3
Query: 236 PVGPIIETETKSGDDANGLECLAWLDKQQPCSVLYVSFGSGGTLSQEQIVELALGLELSN 295
P+ +E++ + G+D + E L WLDKQ SV+YVSFGSGGT+ Q Q+ E+ALGLELS
Sbjct: 6 PLVRTVESKPRHGEDQD--EILRWLDKQPAGSVIYVSFGSGGTMPQGQMTEIALGLELSQ 179
Query: 296 TKFLWVLRAPSSSSSSAGYLSAENDIDTLQFLPSGFLERTKEKGFVITSWAPQIQILSPN 355
+F+WV+R P+ + +SA + N++ L +LP GF+ RT+E G V+ WAPQ +IL
Sbjct: 180 HRFVWVVRPPNEADASATFFGFGNEM-ALNYLPEGFMNRTREFGVVVPMWAPQAEILGHP 356
Query: 356 SVGGFLTHCGWNSTLESVVHGVPLITWPLFAEQKMNAVLLSEGLKVGLRASVNENGIVER 415
+ GGF+T CGWNS LESV++GVP++ WPL++EQKMNA +LSE L V +R V E G+V R
Sbjct: 357 ATGGFVTPCGWNSVLESVLNGVPMVAWPLYSEQKMNAYMLSEELGVAVRVKVAEGGVVCR 536
Query: 416 VEVAKVIKYLMEGDEGEKLRNNMKELKEAASNAV 449
+V +++ +M +EG +++ ++ELK + A+
Sbjct: 537 DQVVDMVRRVMVSEEGMGMKDRVRELKLSGDEAL 638
>TC9079 weakly similar to UP|Q8W4G1 (Q8W4G1) UDP-glucose
glucosyltransferase, partial (65%)
Length = 1585
Score = 171 bits (432), Expect = 3e-43
Identities = 144/505 (28%), Positives = 223/505 (43%), Gaps = 41/505 (8%)
Frame = +1
Query: 2 EKTVHIAVVPGVGYSHLVPILQFSKRLVQLHPNFHVTCFIPTLGSPSNATKS----ILQT 57
E+ H + P H+ P+ Q +K L+ L FH+T F+ T + KS L
Sbjct: 7 ERKPHAVLTPSPLQGHINPLFQLAK-LLHLR-GFHIT-FVHTEYNHKRLIKSRGPNALDG 177
Query: 58 LPSNINHTFLPPVNPNDLPQGTTMESQILLTLTNS------LPY------LHQGLKSLAK 105
LP + T P+ L G SQ L +L +S LP+ LH+ S
Sbjct: 178 LPDFVFETI-----PDGLEDGDGEVSQDLYSLCDSIRNKCHLPFQDLLAKLHRSA-SAGL 339
Query: 106 EIPLVALVVDAFSVEVLNIGKELNMLSYIYFPSAATTLAWCFYLPKLDEETSCEYRDILE 165
P+ LV D + ++L + + P++A+T + L ++ +D
Sbjct: 340 VPPVTCLVSDLLMTFTIQAAQQLQLPILLLCPASASTFMSFLHFQTLLDKGVIPLKDESY 519
Query: 166 PIK---------IPGCVPLHGRDFLSIAQ--DRSSQAYKHFLPFVKLLSSADGVLVNSFL 214
IPG +D + D + + + A ++ N+F
Sbjct: 520 LTNGYLDTKVDWIPGMQNFRLKDLPDFIRTTDPNDIMLEFMVEVADRAKRASAIVFNTFN 699
Query: 215 EIEMGPLSAMKEEGGDNPPVYPVGPIIETETKSGDDA----------NGLECLAWLDKQQ 264
E+E LSA+ P +YP+GP ++ + +CL WL+ ++
Sbjct: 700 ELERDVLSALSTM---LPSLYPIGPFPSFLNQAPQNQLASLGSNLWKEDTKCLQWLESKE 870
Query: 265 PCSVLYVSFGSGGTLSQEQIVELALGLELSNTKFLWVLRAPSSSSSSAGYLSAENDIDTL 324
P SV+YV+FGS +S EQ++E A GL FLW++R + SA
Sbjct: 871 PGSVVYVNFGSITVMSPEQLLEFAWGLANGKKPFLWIIRPDLVADGSA------------ 1014
Query: 325 QFLPSGFLERTKEKGFVITSWAPQIQILSPNSVGGFLTHCGWNSTLESVVHGVPLITWPL 384
L F++ T +G +I SW PQ ++L+ SVGGFLTHCGWNST+ES+ GVP++ WP
Sbjct: 1015-ILSPKFVDETSNRG-LIASWCPQEKVLNHPSVGGFLTHCGWNSTIESICAGVPMLCWPF 1188
Query: 385 FAEQKMNAVLLSEGLKVGLRASVNENGIVERVEVAKVIKYLMEGDEGEKLRNNMKELKEA 444
A+Q N + +G+ N V+R EV +I LM GD+G+K+R ELK+
Sbjct: 1189VADQPTNCRYICNEWDIGIEVDTN----VKREEVENLIHELMVGDKGKKMRQRTMELKKK 1356
Query: 445 ASNAVKEDGSS----TKTISQIALK 465
A + G S K I ++ LK
Sbjct: 1357AEEDTRPGGCSYMNLDKLIKEVLLK 1431
>TC9184 weakly similar to UP|HQGT_RAUSE (Q9AR73) Hydroquinone
glucosyltransferase (Arbutin synthase) , partial (26%)
Length = 643
Score = 167 bits (424), Expect = 3e-42
Identities = 77/122 (63%), Positives = 102/122 (83%)
Frame = +3
Query: 346 APQIQILSPNSVGGFLTHCGWNSTLESVVHGVPLITWPLFAEQKMNAVLLSEGLKVGLRA 405
APQ ++LS S+GGFL+HCGWNS LESV +GVPL+ WPL+AEQ+MNAVL++E +KV LR
Sbjct: 3 APQAKVLSHASIGGFLSHCGWNSVLESVANGVPLVAWPLYAEQRMNAVLVTEDVKVALRP 182
Query: 406 SVNENGIVERVEVAKVIKYLMEGDEGEKLRNNMKELKEAASNAVKEDGSSTKTISQIALK 465
V ENG+VE VE+A+V++ LM+G+EG+KLR MK+LKEAA+ +KE+GSST I Q+ALK
Sbjct: 183 KVGENGLVELVEIARVVRTLMQGEEGKKLRCKMKDLKEAAATTLKENGSSTNQIFQLALK 362
Query: 466 WR 467
W+
Sbjct: 363 WK 368
>TC9836 weakly similar to UP|Q9LXV0 (Q9LXV0) Glucosyltransferase-like
protein, partial (55%)
Length = 1286
Score = 166 bits (420), Expect = 8e-42
Identities = 97/272 (35%), Positives = 150/272 (54%), Gaps = 14/272 (5%)
Frame = +1
Query: 204 SADGVLVNSFLEIEMGPLSAMKEEGGDNPPVYPVGPII-ETETKSGDDANGLE---CLAW 259
++DG L N+ + + L + + N P + +GP++ T + S G+ C W
Sbjct: 217 NSDGFLFNTVADFDSMGLRYVARKL--NRPAWAIGPVLLSTGSGSRGKGGGISPELCRKW 390
Query: 260 LDKQQPCSVLYVSFGSGGTLSQEQIVELALGLELSNTKFLWVLRAPSSSSSSAGYLSAEN 319
LD + SVL+VSFGS T+S Q+++LA L+ S F+WV+R P ++ + + E
Sbjct: 391 LDTKPSNSVLFVSFGSMNTISASQMMQLATALDRSGRNFIWVVRPPIGFDINSEFRAEE- 567
Query: 320 DIDTLQFLPSGFLERTKEKGFVITSWAPQIQILSPNSVGGFLTHCGWNSTLESVVHGVPL 379
+LP GFL R +KG V+ WAPQ++ILS +V FL+HCGWNS LES+ HGVP+
Sbjct: 568 ------WLPEGFLGRVAKKGLVVHDWAPQVEILSHGAVSAFLSHCGWNSVLESLSHGVPI 729
Query: 380 ITWPLFAEQKMNAVLLSEGLKV------GLRASVNENGIVERVEVAKVIKYLMEGDEGEK 433
+ WP+ AEQ N +L E L V G R V +VE++E+ + E + G K
Sbjct: 730 LGWPMAAEQFFNCKMLEEELGVCVEVARGKRCEVRHEDLVEKIELV-----MNEAESGVK 894
Query: 434 LRNNMKELKEAASNAVKED----GSSTKTISQ 461
+R N ++E +AV+++ GSS + I +
Sbjct: 895 IRKNAGNIREMIRDAVRDEDGYKGSSVRAIDE 990
>TC10018 weakly similar to UP|HQGT_RAUSE (Q9AR73) Hydroquinone
glucosyltransferase (Arbutin synthase) , partial (24%)
Length = 586
Score = 155 bits (393), Expect = 1e-38
Identities = 73/111 (65%), Positives = 92/111 (82%)
Frame = +1
Query: 356 SVGGFLTHCGWNSTLESVVHGVPLITWPLFAEQKMNAVLLSEGLKVGLRASVNENGIVER 415
S GGFLTHCGWNS LESVV+GVPL+ WPLFAEQKMNAVLL++ +KV LR V E+G+VE+
Sbjct: 7 STGGFLTHCGWNSILESVVYGVPLVAWPLFAEQKMNAVLLTQEIKVALRPRVGEDGLVEK 186
Query: 416 VEVAKVIKYLMEGDEGEKLRNNMKELKEAASNAVKEDGSSTKTISQIALKW 466
E+A V+K LMEG+EG+KLR MK +KEA + ++ E+GSS+ ISQ+ALKW
Sbjct: 187 HEIASVVKCLMEGEEGKKLRYQMKNMKEAGARSLGENGSSSNHISQLALKW 339
>CN825072
Length = 712
Score = 130 bits (328), Expect = 4e-31
Identities = 76/200 (38%), Positives = 110/200 (55%), Gaps = 11/200 (5%)
Frame = -3
Query: 274 GSGGTLSQEQIVELALGLELSNTKFLWVLRAPSSSSSSAGYLSAENDIDTLQFLPSGFLE 333
GS G+ + QI E+A +E S +F+W LR P S G D D LP GFL+
Sbjct: 710 GSRGSFDRAQITEIARAVENSGVRFVWSLRKPXLKGSMVGPSDYSVD-DLASVLPEGFLD 534
Query: 334 RTKEKGFVITSWAPQIQILSPNSVGGFLTHCGWNSTLESVVHGVPLITWPLFAEQKMNAV 393
RT G VI WAPQ ++L+ + GGF++HCGWNSTLES+ GVP+ TWPL+AEQ+ NA
Sbjct: 533 RTTGIGRVI-GWAPQTRVLTHPATGGFVSHCGWNSTLESIYFGVPIATWPLYAEQQTNAF 357
Query: 394 LLSEGLKVGLRASVNENGIVERVEVAKVIKYLMEGDEGE-----------KLRNNMKELK 442
+L LK+ + S++ RVE+ YL+ D+ E ++R +KE+
Sbjct: 356 VLVRELKIAVEISLD-----YRVEINGGPNYLLTADKIEGGIRSVLDKDGEVRKRVKEMS 192
Query: 443 EAASNAVKEDGSSTKTISQI 462
E + + E G S + ++
Sbjct: 191 EKSRKTLLEGGCSYSYLDRL 132
>CB829602
Length = 542
Score = 124 bits (312), Expect = 3e-29
Identities = 69/167 (41%), Positives = 100/167 (59%)
Frame = +1
Query: 255 ECLAWLDKQQPCSVLYVSFGSGGTLSQEQIVELALGLELSNTKFLWVLRAPSSSSSSAGY 314
ECL WLD Q+P SVLYV+FGS ++ +Q+VELA G+ S KF+WV+R P A
Sbjct: 91 ECLKWLDSQEPSSVLYVNFGSVINMTPQQLVELAWGIANSKKKFIWVIR-PDLVEGEASI 267
Query: 315 LSAENDIDTLQFLPSGFLERTKEKGFVITSWAPQIQILSPNSVGGFLTHCGWNSTLESVV 374
+ E + TK++G ++ SW PQ Q+L +++GGFLTHCGWNST+ES+
Sbjct: 268 VLPE------------IVAETKDRGIML-SWCPQEQVLKHSALGGFLTHCGWNSTIESIS 408
Query: 375 HGVPLITWPLFAEQKMNAVLLSEGLKVGLRASVNENGIVERVEVAKV 421
GVPLI P F +Q +N+ + G+ + ++ V+R EV K+
Sbjct: 409 SGVPLICSPFFNDQFVNSRYICSEWDFGMEMNSDD---VKRDEVEKL 540
>TC11224 weakly similar to UP|Q8S996 (Q8S996) Glucosyltransferase-13
(Fragment), partial (19%)
Length = 596
Score = 123 bits (309), Expect = 6e-29
Identities = 69/139 (49%), Positives = 94/139 (66%), Gaps = 3/139 (2%)
Frame = +3
Query: 329 SGFLERTKEKGFVITSWAPQIQILSPNSVGGFLTHCGWNSTLESVVHGVPLITWPLFAEQ 388
+GFLERTK +G V+ SWAPQ++IL SVG FLTH GWNS LE +V GVP+I P F +Q
Sbjct: 3 NGFLERTKSQGKVV-SWAPQMEILKHASVGVFLTHGGWNSVLECIVGGVPMIGRPFFGDQ 179
Query: 389 KMNAVLLSEGLKVGLRASVNENGIVERVEVAKVIKYLMEGDEGEKLRNNMKELKEAASNA 448
++N +L++ L VG+ +NG++ + + K +K M G+EG+ +R M ELKE A A
Sbjct: 180 RINIWMLAKLLGVGVGL---QNGVLAKETILKTLKSTMTGEEGKVMRQKMAELKEMAWKA 350
Query: 449 VKEDGSSTK---TISQIAL 464
V+ DGSSTK T+ QI L
Sbjct: 351 VEPDGSSTKNLCTLMQIIL 407
>TC15956 weakly similar to UP|LGT_CITUN (Q9MB73) Limonoid
UDP-glucosyltransferase (Limonoid glucosyltransferase)
(Limonoid GTase) (LGTase) , partial (65%)
Length = 1169
Score = 120 bits (300), Expect = 6e-28
Identities = 116/411 (28%), Positives = 189/411 (45%), Gaps = 29/411 (7%)
Frame = +2
Query: 2 EKTVHIAVVPGVGYSHLVPILQFSKRLVQLHPNFHVTCFIPTLGSPSNATKSILQTLPSN 61
E +HI +V H+ P+L+ +K L + TL + A KS+ +N
Sbjct: 26 ESPLHILLVSFPAQGHINPLLRLAKCLAAKGSSV-------TLATTQTAGKSMRTA--TN 178
Query: 62 INHTFLPPVNPNDL----------PQGTTMESQILLTLTNSLPYLHQGLKSLAKEIP--- 108
I P++ L P ++ + L L Q + + KE
Sbjct: 179 ITDKAPTPISDGSLKFEFFDDGLGPDDDSIRGNLSLYLQQLELVGRQQILKMIKEHADSN 358
Query: 109 --LVALVVDAFSVEVLNIGKELNMLS-YIYFPSAATTLAWCFYLPKLDEETSCEYRDILE 165
+ ++ + F V+++ +E + S ++ S+A A+ Y L S E D L
Sbjct: 359 RQISCIINNPFLPWVVDVAEEQGIPSALLWIQSSAVFTAYYSYFHNLVTFPSKE--DPLA 532
Query: 166 PIKIPGC--VPLHGR--DFLSIAQDRSSQAYKHF----LPFVKLLSSADGVLVNSFLEIE 217
+++P V H DFL S Y L K L+ +LV+++ E+E
Sbjct: 533 DVQLPSTSIVLKHCEIPDFL-----HPSCTYPFLGTLILEQFKNLNKTFCILVDTYEELE 697
Query: 218 MGPLSAMKEEGGDNPPVYPVGPIIET-ETKS----GDDANGLECLAWLDKQQPCSVLYVS 272
+ + + + PVGP+ ++ + KS GD +C+ WL+ + SV+YVS
Sbjct: 698 HDFIDYLSKTF----LIRPVGPLFKSSQAKSAAIRGDFMKSDDCIEWLNSKPQSSVVYVS 865
Query: 273 FGSGGTLSQEQIVELALGLELSNTKFLWVLRAPSSSSSSAGYLSAENDIDTLQFLPSGFL 332
FGS L QEQ+ E+A GL+ S FLWV++ P + ++ LP GFL
Sbjct: 866 FGSIVYLPQEQVDEIAFGLKKSQVSFLWVMKPPPEVTGLQPHV-----------LPYGFL 1012
Query: 333 ERTKEKGFVITSWAPQIQILSPNSVGGFLTHCGWNSTLESVVHGVPLITWP 383
E T E+G V+ W+PQ ++L+ SV FLTHCGWNS++E++ GVP++T+P
Sbjct: 1013EETGERGRVV-QWSPQEEVLAHPSVSCFLTHCGWNSSMEALTSGVPVLTFP 1162
>BI421006
Length = 519
Score = 114 bits (285), Expect = 3e-26
Identities = 61/138 (44%), Positives = 81/138 (58%)
Frame = +2
Query: 266 CSVLYVSFGSGGTLSQEQIVELALGLELSNTKFLWVLRAPSSSSSSAGYLSAENDIDTLQ 325
CS+ F S T +EQI ++A GLE S KF+WVLR G E
Sbjct: 86 CSICIFWFFS--TFIEEQIEQIANGLEQSKQKFIWVLRDADKGDIFDGDKVKERG----- 244
Query: 326 FLPSGFLERTKEKGFVITSWAPQIQILSPNSVGGFLTHCGWNSTLESVVHGVPLITWPLF 385
LP GF +R + G V+ WAPQ++ILS S GGF++HCGWNS +ES+ GVP+ WP+
Sbjct: 245 -LPKGFEKRVEGMGLVVRDWAPQLEILSHPSTGGFMSHCGWNSCIESMSMGVPIAAWPMH 421
Query: 386 AEQKMNAVLLSEGLKVGL 403
++Q N VL++E LKV L
Sbjct: 422 SDQPRNTVLITEVLKVAL 475
Score = 30.0 bits (66), Expect = 0.85
Identities = 12/18 (66%), Positives = 15/18 (82%)
Frame = +3
Query: 257 LAWLDKQQPCSVLYVSFG 274
+ WLD+Q+ SVLYVSFG
Sbjct: 54 MEWLDRQEQKSVLYVSFG 107
>TC16143 similar to UP|Q7XZD0 (Q7XZD0) Isoflavonoid glucosyltransferase,
partial (30%)
Length = 604
Score = 113 bits (283), Expect = 6e-26
Identities = 57/137 (41%), Positives = 82/137 (59%), Gaps = 10/137 (7%)
Frame = +1
Query: 336 KEKGFVITSWAPQIQILSPNSVGGFLTHCGWNSTLESVVHGVPLITWPLFAEQKMNAVLL 395
+EKG +I W PQ+ IL +VG F+THCGWNST+E+V GVP+ITWP+ EQ N L+
Sbjct: 1 REKGMIIRGWVPQVVILGHRAVGAFVTHCGWNSTVEAVSAGVPMITWPVVGEQFYNEKLI 180
Query: 396 SEGLKVGLRASVNE---------NGIVERVEVAKVIKYLME-GDEGEKLRNNMKELKEAA 445
+E +G+ E +V R + K ++ LM+ GDEGE++R +E E A
Sbjct: 181 TEVRGIGVEVGAEECCLMGFDKREKLVSRESIEKAVRRLMDGGDEGEQIRRRAQEYGEKA 360
Query: 446 SNAVKEDGSSTKTISQI 462
AV+E GSS K ++ +
Sbjct: 361 RQAVEEGGSSHKNLTAL 411
>TC17165 similar to UP|Q9SMG6 (Q9SMG6) Betanidin-5-O-glucosyltransferase,
partial (24%)
Length = 546
Score = 112 bits (280), Expect = 1e-25
Identities = 57/129 (44%), Positives = 81/129 (62%), Gaps = 7/129 (5%)
Frame = +1
Query: 341 VITSWAPQIQILSPNSVGGFLTHCGWNSTLESVVHGVPLITWPLFAEQKMNAVLLSEGLK 400
+I WAPQ+ IL ++G F+THCGWNSTLE V GVP++TWP+ AEQ N L++E LK
Sbjct: 1 IIRGWAPQVLILEHEAIGAFVTHCGWNSTLEGVAAGVPMVTWPIAAEQFYNEKLVTEVLK 180
Query: 401 VGLRAS-------VNENGIVERVEVAKVIKYLMEGDEGEKLRNNMKELKEAASNAVKEDG 453
+G+ V E V+ V K +K +MEG+E E++RN +K L + A AV+E G
Sbjct: 181 IGVPVGAKKWGRMVLEGDGVKSDAVEKGVKRIMEGEEAEEMRNRVKVLSQQAQWAVEEGG 360
Query: 454 SSTKTISQI 462
SS ++ +
Sbjct: 361 SSYSDLNAL 387
>TC12282 similar to UP|Q8S996 (Q8S996) Glucosyltransferase-13 (Fragment),
partial (7%)
Length = 511
Score = 112 bits (279), Expect = 2e-25
Identities = 54/102 (52%), Positives = 77/102 (74%), Gaps = 1/102 (0%)
Frame = +3
Query: 369 TLESVVHGVPLITWPLFAEQKMNAVLLSEGLKVGLRASVN-ENGIVERVEVAKVIKYLME 427
TLESVV+GVP+I WPLFAEQ+MNA ++++ LK+G+R V+ E G+V+R E+A+ +K +ME
Sbjct: 3 TLESVVNGVPMIVWPLFAEQRMNAKIITDALKIGVRPKVDGETGMVKREEIARGVKRIME 182
Query: 428 GDEGEKLRNNMKELKEAASNAVKEDGSSTKTISQIALKWRNL 469
GDE ++ +KEL + A+ A+ E GSS IS +ALKW NL
Sbjct: 183 GDESLEICRRVKELSDGAAAALSEHGSSRNAISSLALKWNNL 308
>TC9485 similar to UP|Q8S999 (Q8S999) Glucosyltransferase-10, partial (75%)
Length = 1195
Score = 109 bits (272), Expect = 1e-24
Identities = 110/403 (27%), Positives = 179/403 (44%), Gaps = 42/403 (10%)
Frame = +1
Query: 3 KTVHIAVVPGVGYSHLVPILQFSKRLVQLHPNFHVTCFIPTLGSPSNATKS----ILQTL 58
K H+ +P H+ P+L+ +K L+ FHVT F+ T + KS L L
Sbjct: 82 KKPHVVCIPYPAQGHINPMLKLAK-LLHFKGGFHVT-FVNTEYNHKRLLKSRGPDSLNGL 255
Query: 59 PSNINHTFLPPVNPNDLPQGTTMESQILLTLTNSLPYLHQGLKSL---AKEIPLVALVV- 114
PS T + D+ + S + T LP+ + L L + ++P V +V
Sbjct: 256 PSFRFETIPDGLPETDVDVTQDIPSLCISTRKTCLPHFKKLLSKLNDVSSDVPPVTCIVS 435
Query: 115 DAFSVEVLNIGKELNMLSYIYFPSAATTLAWCFYLPKLDEETSCEYRDILE----PIK-- 168
D L+ ELN+ +++ ++A C ++ + +YR+++E P+K
Sbjct: 436 DGCMSFTLDAAIELNIPEVLFWTTSA-----CGFMGYV------QYRELIEKGIIPLKDS 582
Query: 169 --------------IPGCVPLHGRDFLSIAQDRSSQAYKHFLPFV----KLLSSADGVLV 210
+PG + +D S R++ L F+ + A +++
Sbjct: 583 SDITNGYLETTIEWLPGMKNIRLKDLPSFL--RTTDPNDKMLDFLTGECQRALKASAIIL 756
Query: 211 NSFLEIEMGPLSAMKEEGGDNPPVYPVGPI---IETETKSGDDANGL-------ECLAWL 260
N+F +E L A PPVY +GP+ I+ T ++ G ECL WL
Sbjct: 757 NTFDALEHDVLEAFSSI---LPPVYSIGPLHLLIKDVTDKNLNSLGSNLWKEDSECLKWL 927
Query: 261 DKQQPCSVLYVSFGSGGTLSQEQIVELALGLELSNTKFLWVLRAPSSSSSSAGYLSAEND 320
D ++P SV+YV+FGS ++ EQ+VE A GL SN FLWV+R + A
Sbjct: 928 DTKEPNSVVYVNFGSITVMTSEQMVEFAWGLANSNKTFLWVIRPDLVAGKHA-------- 1083
Query: 321 IDTLQFLPSGFLERTKEKGFVITSWAPQIQILSPNSVGGFLTH 363
LP F+ T ++G ++SW PQ +L+ ++GGFLTH
Sbjct: 1084-----VLPEEFVAATNDRG-RLSSWTPQEDVLTHPAIGGFLTH 1194
>TC9717 weakly similar to UP|Q9LNI1 (Q9LNI1) F6F3.22 protein, partial (18%)
Length = 902
Score = 88.6 bits (218), Expect(2) = 2e-24
Identities = 52/147 (35%), Positives = 83/147 (56%), Gaps = 1/147 (0%)
Frame = +2
Query: 298 FLWVLRAPSSSSSSAGYLSAENDIDTLQFLPSGFLERTKEKGFVITSWAPQIQILSPNSV 357
F WVL+ +S S+ + D ++ ++ +KG V +WAPQ++IL+ S+
Sbjct: 98 FFWVLKKQNSDSADSH--------DWIE-------NQSNKKGMVWRTWAPQMRILAHKSI 232
Query: 358 GGFLTHCGWNSTLESVVHGVPLITWPLFAEQKMNAVLLSEGLKVGLRASVNE-NGIVERV 416
GGFLTHCGW+S +E + G PLI P F ++M EG K+G++ S N+ +G R
Sbjct: 233 GGFLTHCGWSSVIEGLQVGCPLIMLP-FPNEQMLVARFMEGKKLGVKVSKNQHDGKFTRD 409
Query: 417 EVAKVIKYLMEGDEGEKLRNNMKELKE 443
VAK ++ +M +EG+ R +EL +
Sbjct: 410 SVAKALRSVMLEEEGKSYRCQAEELSK 490
Score = 40.8 bits (94), Expect(2) = 2e-24
Identities = 21/32 (65%), Positives = 23/32 (71%)
Frame = +1
Query: 267 SVLYVSFGSGGTLSQEQIVELALGLELSNTKF 298
SV+YV+FGS TLS E ELALGLELS F
Sbjct: 4 SVIYVAFGSEVTLSNEDFTELALGLELSGFPF 99
>AV416902
Length = 404
Score = 107 bits (266), Expect = 5e-24
Identities = 55/134 (41%), Positives = 81/134 (60%)
Frame = +3
Query: 11 PGVGYSHLVPILQFSKRLVQLHPNFHVTCFIPTLGSPSNATKSILQTLPSNINHTFLPPV 70
P SHL+P+++ ++RLV H + HVT IPTL + + K++L TLP N N T LP V
Sbjct: 3 PSPELSHLIPLVELARRLVLHHHDLHVTLLIPTLVPLTASMKALLNTLPPNTNFTVLPQV 182
Query: 71 NPNDLPQGTTMESQILLTLTNSLPYLHQGLKSLAKEIPLVALVVDAFSVEVLNIGKELNM 130
DLP ++ L + +S P++H+ L SL LVALV D FS + ++ K+LN+
Sbjct: 183 KIEDLPHNINPVTKWKLIVQHSTPFIHEALTSLISSTRLVALVYDMFSFDAHHVAKKLNL 362
Query: 131 LSYIYFPSAATTLA 144
LSY++F S A L+
Sbjct: 363 LSYLFFISGAVVLS 404
>TC19068 weakly similar to UP|AAR06913 (AAR06913) UDP-glycosyltransferase
85A8, partial (26%)
Length = 565
Score = 105 bits (261), Expect = 2e-23
Identities = 54/136 (39%), Positives = 81/136 (58%)
Frame = +1
Query: 327 LPSGFLERTKEKGFVITSWAPQIQILSPNSVGGFLTHCGWNSTLESVVHGVPLITWPLFA 386
L S F+ +G +I SW Q +L+ S+GGFLTHCGWNST+ES+ GVP++ WP FA
Sbjct: 16 LSSEFINEISSRG-LIASWCSQEHVLNHPSIGGFLTHCGWNSTIESISAGVPMLCWPFFA 192
Query: 387 EQKMNAVLLSEGLKVGLRASVNENGIVERVEVAKVIKYLMEGDEGEKLRNNMKELKEAAS 446
+Q N + +G++ ++ NG +R EV K+I LM G++G+K+R ELK+ A
Sbjct: 193 DQLTNCRSICNEWDIGMQ--IDTNG--KREEVEKLINELMVGEKGKKMRQKTMELKKRAE 360
Query: 447 NAVKEDGSSTKTISQI 462
+ G S + Q+
Sbjct: 361 EDTRPGGCSYMNLDQV 408
>TC10667 similar to UP|Q9M051 (Q9M051) Glucuronosyl transferase-like
protein, partial (20%)
Length = 824
Score = 103 bits (256), Expect = 8e-23
Identities = 55/111 (49%), Positives = 73/111 (65%)
Frame = +1
Query: 345 WAPQIQILSPNSVGGFLTHCGWNSTLESVVHGVPLITWPLFAEQKMNAVLLSEGLKVGLR 404
WAPQ Q+L ++G F THCGWNSTLESV GVP+I P F +QK+NA +S+ KVG++
Sbjct: 22 WAPQEQVLKHPAIGAFWTHCGWNSTLESVCEGVPMICSPCFGDQKVNAKYVSDVWKVGVQ 201
Query: 405 ASVNENGIVERVEVAKVIKYLMEGDEGEKLRNNMKELKEAASNAVKEDGSS 455
N+ G R E+ K I LM GDE ++R N+ +LKE A+ + E GSS
Sbjct: 202 LQ-NKPG---RGEIEKTISLLMLGDEAYEIRGNILKLKEKANVCLSEGGSS 342
>TC14237 weakly similar to UP|AAR06913 (AAR06913) UDP-glycosyltransferase
85A8, partial (27%)
Length = 999
Score = 102 bits (255), Expect = 1e-22
Identities = 59/166 (35%), Positives = 91/166 (54%)
Frame = +2
Query: 290 GLELSNTKFLWVLRAPSSSSSSAGYLSAENDIDTLQFLPSGFLERTKEKGFVITSWAPQI 349
G+ S FLW+LR + E+ I+ +P FL+ K +G+ I SW Q
Sbjct: 194 GIANSKVPFLWILRPD--------VVMGEDSIN----VPQEFLDEIKGRGY-IASWCFQE 334
Query: 350 QILSPNSVGGFLTHCGWNSTLESVVHGVPLITWPLFAEQKMNAVLLSEGLKVGLRASVNE 409
++LS S+G FLTHCGWNST+E + G+PLI WP F+EQ N+ ++G+
Sbjct: 335 EVLSHPSIGAFLTHCGWNSTIEGISAGLPLICWPFFSEQHTNSRYACTTWEIGMEV---- 502
Query: 410 NGIVERVEVAKVIKYLMEGDEGEKLRNNMKELKEAASNAVKEDGSS 455
N V+R E+ ++ +M+G++G+++R E K+ A A GSS
Sbjct: 503 NHDVKRDEITTLVNEMMKGEKGKEMRKKGLEWKKKAIEATGLGGSS 640
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.316 0.135 0.395
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 8,499,680
Number of Sequences: 28460
Number of extensions: 126019
Number of successful extensions: 821
Number of sequences better than 10.0: 99
Number of HSP's better than 10.0 without gapping: 785
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 789
length of query: 470
length of database: 4,897,600
effective HSP length: 94
effective length of query: 376
effective length of database: 2,222,360
effective search space: 835607360
effective search space used: 835607360
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.6 bits)
S2: 57 (26.6 bits)
Medicago: description of AC124966.3


