

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
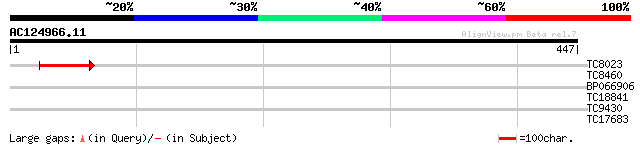
Query= AC124966.11 - phase: 0
(447 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
TC8023 51 3e-07
TC8460 similar to GB|AAN73294.1|25141199|BT002297 At5g12010/F14F... 30 0.62
BP066906 28 2.3
TC18841 28 4.0
TC9430 similar to UP|Q96569 (Q96569) L-lactate dehydrogenase (L... 27 6.8
TC17683 weakly similar to UP|P93816 (P93816) F19P19.10, partial ... 27 8.9
>TC8023
Length = 1089
Score = 51.2 bits (121), Expect = 3e-07
Identities = 26/45 (57%), Positives = 32/45 (70%), Gaps = 1/45 (2%)
Frame = +3
Query: 24 PFDIEAYKQKRDIEDTYIVNRFIQRRKELEEGSGSR-SRKYLNRD 67
PFD+EAYK++ +IED YI N F +RR + EG R RKYLNRD
Sbjct: 54 PFDVEAYKRQSEIEDRYIFNLFRKRRNLILEGYTPRVKRKYLNRD 188
>TC8460 similar to GB|AAN73294.1|25141199|BT002297 At5g12010/F14F18_180
{Arabidopsis thaliana;}, partial (36%)
Length = 880
Score = 30.4 bits (67), Expect = 0.62
Identities = 30/132 (22%), Positives = 56/132 (41%), Gaps = 1/132 (0%)
Frame = +3
Query: 82 NEPTYGDAMFRRRYRMQKHVFLRIVGDLSITDNYFTQRVDAANKEGISPLAKCTTAMRML 141
+ P + + FRR +RM K F I +L D+ T++ + +E I + + L
Sbjct: 357 SHPDFPEEEFRRSFRMSKATFEMICREL---DSAVTKK-NTMLREAIPVRQRVAVCIWRL 524
Query: 142 AYGVAADAVDEYIKIGGTTALECLRRFCKGIIRLYEEVYLRAPNQDDLQRILHVSE-MRG 200
A G V + +G +T + + C I + +LR P++ + E + G
Sbjct: 525 ATGDPLRLVAKRFGLGISTCHKLVLEVCSAIKTVLMPKFLRWPDEAAMTAAKSEFEALSG 704
Query: 201 FPGMIGSIDCMH 212
P + G++ H
Sbjct: 705 IPNIGGAMYTTH 740
>BP066906
Length = 386
Score = 28.5 bits (62), Expect = 2.3
Identities = 10/32 (31%), Positives = 16/32 (49%)
Frame = +3
Query: 228 RGDKGTTTVILEAVASHDLWIWHAFFGCPGTL 259
R ++ T I+ + S W+WH FG P +
Sbjct: 225 RHEENYTRYIIWILPSFGFWLWHVIFGFPNNI 320
>TC18841
Length = 568
Score = 27.7 bits (60), Expect = 4.0
Identities = 15/37 (40%), Positives = 19/37 (50%), Gaps = 1/37 (2%)
Frame = +2
Query: 210 CMHWEWKN-CPKAWEGQFTRGDKGTTTVILEAVASHD 245
C HWEW+N C K +G +G +L VA HD
Sbjct: 326 CSHWEWENSCQKYLKG----FSQGYMKPLLAGVA*HD 424
>TC9430 similar to UP|Q96569 (Q96569) L-lactate dehydrogenase (LDH) ,
partial (49%)
Length = 866
Score = 26.9 bits (58), Expect = 6.8
Identities = 15/39 (38%), Positives = 19/39 (48%)
Frame = +2
Query: 400 AQPYSTEVLPTFANHARARSELRDPNVHHELQADLVKHI 438
AQP + E LP A+H R+ S L HH Q H+
Sbjct: 602 AQPPAEEPLPLQADHTRSGSLLPGLCPHHRFQPRRCPHL 718
>TC17683 weakly similar to UP|P93816 (P93816) F19P19.10, partial (5%)
Length = 561
Score = 26.6 bits (57), Expect = 8.9
Identities = 10/20 (50%), Positives = 14/20 (70%)
Frame = +2
Query: 6 ITNLIPIYFQYHQRLSMDPF 25
+T LIP+ +H+ SMDPF
Sbjct: 287 LTPLIPLQMFFHEYFSMDPF 346
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.323 0.138 0.428
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 7,737,889
Number of Sequences: 28460
Number of extensions: 101531
Number of successful extensions: 500
Number of sequences better than 10.0: 12
Number of HSP's better than 10.0 without gapping: 498
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 500
length of query: 447
length of database: 4,897,600
effective HSP length: 93
effective length of query: 354
effective length of database: 2,250,820
effective search space: 796790280
effective search space used: 796790280
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.5 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (22.0 bits)
S2: 57 (26.6 bits)
Medicago: description of AC124966.11


