

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC124963.1 + phase: 1 /pseudo
(826 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
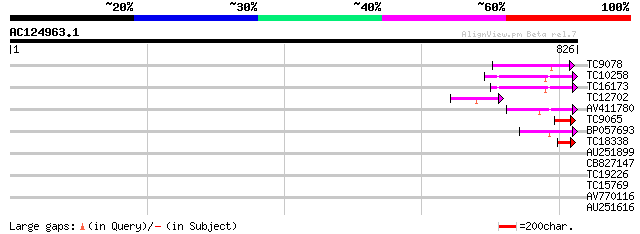
Score E
Sequences producing significant alignments: (bits) Value
TC9078 similar to AAQ89615 (AAQ89615) At3g17205, partial (12%) 74 7e-14
TC10258 similar to GB|AAP68269.1|31711826|BT008830 At1g70320 {Ar... 74 9e-14
TC16173 similar to GB|AAP68269.1|31711826|BT008830 At1g70320 {Ar... 64 1e-10
TC12702 similar to UP|AAP91821 (AAP91821) HECT ubiquitin-protein... 63 2e-10
AV411780 61 6e-10
TC9065 similar to UP|O24402 (O24402) Tap (Fragment), partial (36%) 54 1e-07
BP057693 50 2e-06
TC18338 49 3e-06
AU251899 33 0.14
CB827147 33 0.18
TC19226 similar to UP|AAP91821 (AAP91821) HECT ubiquitin-protein... 32 0.31
TC15769 similar to UP|O24402 (O24402) Tap (Fragment), partial (27%) 31 0.91
AV770116 28 4.5
AU251616 28 5.9
>TC9078 similar to AAQ89615 (AAQ89615) At3g17205, partial (12%)
Length = 666
Score = 74.3 bits (181), Expect = 7e-14
Identities = 47/122 (38%), Positives = 67/122 (54%), Gaps = 3/122 (2%)
Frame = +2
Query: 704 LCGEPNSWTFNDLLKHFKFDHGYTARSPPIMNLLEILQEFNNEERRAFVQFVTRSPRLPP 763
+ G +S +DL H + GY + I L E+L+ F+ E ++ F++FVT R P
Sbjct: 2 ISGSLDSLDVDDLRLHTNYAGGYHSEHYVIEMLWEVLKGFSMENKKKFLKFVTGCSRGPL 181
Query: 764 GGLASLDPKLTVVQKISYNHTDTDL---PSVMTCANYLKLPPYSSKERMKQKLLYAITEG 820
G L+P L +Q+ N ++ L P+ TC N LKLPPY SKE+M+ KLLYAI
Sbjct: 182 LGFRYLEP-LFCIQRAGGNASEEALDRLPTSATCMNLLKLPPYRSKEQMETKLLYAINAD 358
Query: 821 RG 822
G
Sbjct: 359 AG 364
>TC10258 similar to GB|AAP68269.1|31711826|BT008830 At1g70320 {Arabidopsis
thaliana;}, partial (68%)
Length = 907
Score = 73.9 bits (180), Expect = 9e-14
Identities = 49/139 (35%), Positives = 80/139 (57%), Gaps = 4/139 (2%)
Frame = +3
Query: 692 LRVFSEEELELILCGEPNSWTFNDLLKHFKFDHGYTARSPPIMNLLEILQEFNNEERRAF 751
+ +F+++ELEL++ G P+ +DL + + GY+A SP I E++Q F+ E++
Sbjct: 333 ISIFNDQELELLISGLPDI-DLDDLRANTDYS-GYSAASPVIPWFWEVIQGFSQEDKARL 506
Query: 752 VQFVTRSPRLPPGGLASLDPKLTVVQKI----SYNHTDTDLPSVMTCANYLKLPPYSSKE 807
+QFVT + ++P G ++L ++ QK +Y D LPS TC N L LP Y SK+
Sbjct: 507 LQFVTGTSKVPLEGFSALQG-ISGSQKFQIHKAYGSPD-HLPSAHTCFNQLDLPEYPSKQ 680
Query: 808 RMKQKLLYAITEGRGCFLF 826
++++LL AI E F F
Sbjct: 681 HLEERLLLAIHEANEGFGF 737
>TC16173 similar to GB|AAP68269.1|31711826|BT008830 At1g70320 {Arabidopsis
thaliana;}, partial (37%)
Length = 601
Score = 63.9 bits (154), Expect = 1e-10
Identities = 45/130 (34%), Positives = 72/130 (54%), Gaps = 4/130 (3%)
Frame = +3
Query: 701 ELILCGEPNSWTFNDLLKHFKFDHGYTARSPPIMNLLEILQEFNNEERRAFVQFVTRSPR 760
EL++ G P+ +DL + ++ GY+ SP I E++Q F+ E++ +QFVT + +
Sbjct: 3 ELLISGLPDI-DLDDLRANTEYS-GYSTGSPVIQWFWEVVQGFSKEDKARLLQFVTGTSK 176
Query: 761 LPPGGLASLDPKLTVVQKI----SYNHTDTDLPSVMTCANYLKLPPYSSKERMKQKLLYA 816
+P G ++L ++ QK +Y D LPS TC N L LP Y SK+ ++++LL A
Sbjct: 177 VPLEGFSALQG-ISGSQKFQIHKAYGSID-HLPSAHTCFNQLDLPEYPSKQHLEERLLLA 350
Query: 817 ITEGRGCFLF 826
I E F F
Sbjct: 351 IHEANEGFGF 380
>TC12702 similar to UP|AAP91821 (AAP91821) HECT ubiquitin-protein ligase 3,
partial (7%)
Length = 405
Score = 62.8 bits (151), Expect = 2e-10
Identities = 38/98 (38%), Positives = 47/98 (47%), Gaps = 20/98 (20%)
Frame = +1
Query: 642 FRNSKIEDLCLDFSLPGYDQTFLLPFS-IY*ITNLYSF-------------------FLV 681
FR + IEDLCLDF+LPGY L I I+NL + F
Sbjct: 112 FRGAPIEDLCLDFTLPGYPDYILKSGDEIVDISNLEEYISLVVDATVKTGIMRQIEAFRA 291
Query: 682 GFNQVFPIEHLRVFSEEELELILCGEPNSWTFNDLLKH 719
GFNQVF I L++F+ EEL+ +LCG W L H
Sbjct: 292 GFNQVFDISSLQIFTPEELDYLLCGRREMWKTETLADH 405
>AV411780
Length = 353
Score = 61.2 bits (147), Expect = 6e-10
Identities = 39/106 (36%), Positives = 58/106 (53%), Gaps = 4/106 (3%)
Frame = +1
Query: 725 GYTARSPPIMNLLEILQEFNNEERRAFVQFVTRSPRLPPGGLASLD----PKLTVVQKIS 780
GYT S + E+++ FN E+ +QFVT + ++P G +L P+ + K +
Sbjct: 10 GYTVASNVVQWFWEVVKTFNKEDMARLLQFVTGTSKVPLEGFKALQGISGPQRFQIHK-A 186
Query: 781 YNHTDTDLPSVMTCANYLKLPPYSSKERMKQKLLYAITEGRGCFLF 826
Y D LPS TC N L LP Y+SKE+++++LL AI E F F
Sbjct: 187 YGAPDR-LPSAHTCFNQLDLPEYTSKEQLQERLLLAIHEASEGFGF 321
>TC9065 similar to UP|O24402 (O24402) Tap (Fragment), partial (36%)
Length = 567
Score = 53.5 bits (127), Expect = 1e-07
Identities = 24/31 (77%), Positives = 27/31 (86%)
Frame = +3
Query: 794 CANYLKLPPYSSKERMKQKLLYAITEGRGCF 824
CANYLKLPPYS+KE M +KLLYAI EG+G F
Sbjct: 3 CANYLKLPPYSTKEIMYKKLLYAINEGQGSF 95
>BP057693
Length = 557
Score = 49.7 bits (117), Expect = 2e-06
Identities = 31/87 (35%), Positives = 48/87 (54%), Gaps = 3/87 (3%)
Frame = -1
Query: 743 FNNEERRAFVQFVTRSPRLPPGGLASLDPKLTVVQKISYNHT---DTDLPSVMTCANYLK 799
F+ E+ F+QFVT + ++P G +L ++ Q++ + LPS TC N L
Sbjct: 557 FSKEDMARFLQFVTGTSKVPLEGFKALQG-ISGPQRLQIHXAYGAPERLPSAHTCFNQLD 381
Query: 800 LPPYSSKERMKQKLLYAITEGRGCFLF 826
LP Y SKE+++++LL AI E F F
Sbjct: 380 LPEYQSKEQLQERLLLAIHEASEGFGF 300
>TC18338
Length = 439
Score = 48.9 bits (115), Expect = 3e-06
Identities = 21/26 (80%), Positives = 25/26 (95%)
Frame = +1
Query: 799 KLPPYSSKERMKQKLLYAITEGRGCF 824
KLPPYS+KE+MK+KLLYAITEG+G F
Sbjct: 1 KLPPYSTKEKMKEKLLYAITEGQGSF 78
>AU251899
Length = 174
Score = 33.5 bits (75), Expect = 0.14
Identities = 13/26 (50%), Positives = 19/26 (73%)
Frame = +1
Query: 511 WIAAQEMSSFSRPNFGVCCTNDEPEC 536
WIA++E+ S S+ + G+C TND P C
Sbjct: 82 WIASKEILSLSQ*DSGICYTNDGPTC 159
>CB827147
Length = 441
Score = 33.1 bits (74), Expect = 0.18
Identities = 14/46 (30%), Positives = 26/46 (56%)
Frame = +2
Query: 56 ELSVHVMHFLMAYLLHLQVFEFANLTKILCIIFQSTLKQNMWNQYM 101
E +VH M + L+ L V + + KI+C++++ + +WNQ M
Sbjct: 68 ETTVHRMEMQIKRLIFLMVLDLLLIGKIVCLVWRQVTVKFLWNQKM 205
>TC19226 similar to UP|AAP91821 (AAP91821) HECT ubiquitin-protein ligase 3,
partial (6%)
Length = 553
Score = 32.3 bits (72), Expect = 0.31
Identities = 17/42 (40%), Positives = 25/42 (59%)
Frame = +3
Query: 1 VLILALQIVELILQKFFSDKFIKLFIEEGVYFAISHFHHLGK 42
VL+ ALQI E++++K F K+F+ EGV A+ GK
Sbjct: 309 VLVPALQIAEILMEKL-PGTFSKIFVREGVVHAVDQLILAGK 431
>TC15769 similar to UP|O24402 (O24402) Tap (Fragment), partial (27%)
Length = 583
Score = 30.8 bits (68), Expect = 0.91
Identities = 14/22 (63%), Positives = 18/22 (81%)
Frame = +3
Query: 803 YSSKERMKQKLLYAITEGRGCF 824
YS+KE M +KLLYAI+E +G F
Sbjct: 3 YSTKEIMYKKLLYAISEXQGSF 68
>AV770116
Length = 206
Score = 28.5 bits (62), Expect = 4.5
Identities = 13/40 (32%), Positives = 21/40 (52%)
Frame = -3
Query: 787 DLPSVMTCANYLKLPPYSSKERMKQKLLYAITEGRGCFLF 826
DL ++ A+ L LPP+ ++E L + E CF+F
Sbjct: 156 DLSRAISSASTLHLPPHWTQEPQHSSLXVLLDESLTCFIF 37
>AU251616
Length = 350
Score = 28.1 bits (61), Expect = 5.9
Identities = 15/44 (34%), Positives = 24/44 (54%), Gaps = 3/44 (6%)
Frame = +3
Query: 65 LMAYLLHLQVFEFANLTKILCIIFQSTL---KQNMWNQYMFLKG 105
LM Y+++ + NL + +IF S + + +WNQ MFL G
Sbjct: 114 LMNYMINNDLALKGNLDGVELLIFPSNILPERSQLWNQMMFLWG 245
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.358 0.160 0.582
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 16,019,696
Number of Sequences: 28460
Number of extensions: 244277
Number of successful extensions: 2579
Number of sequences better than 10.0: 28
Number of HSP's better than 10.0 without gapping: 1922
Number of HSP's successfully gapped in prelim test: 54
Number of HSP's that attempted gapping in prelim test: 619
Number of HSP's gapped (non-prelim): 2029
length of query: 826
length of database: 4,897,600
effective HSP length: 98
effective length of query: 728
effective length of database: 2,108,520
effective search space: 1535002560
effective search space used: 1535002560
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 14 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 37 (21.7 bits)
S2: 59 (27.3 bits)
Medicago: description of AC124963.1


