

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC124958.16 + phase: 0
(248 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
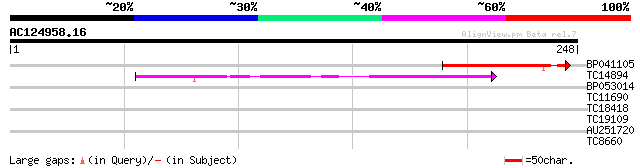
BP041105 46 5e-06
TC14894 weakly similar to UP|Q9LD82 (Q9LD82) ESTs AU029648, part... 40 3e-04
BP053014 27 3.3
TC11690 similar to UP|6PGL_DROME (Q9VZ64) Potential 6-phosphoglu... 27 3.3
TC18418 27 4.3
TC19109 similar to UP|Q84LE0 (Q84LE0) Phytocyanin protein, PUP2,... 26 5.6
AU251720 26 7.3
TC8660 similar to UP|AAR28754 (AAR28754) Bax inhibitor, partial ... 25 9.5
>BP041105
Length = 537
Score = 46.2 bits (108), Expect = 5e-06
Identities = 27/59 (45%), Positives = 37/59 (61%), Gaps = 3/59 (5%)
Frame = -3
Query: 190 VVKFSGVASDEVVKEKVEKLRLSLERDGFKVIGDFLLGRYNPP---WTLPMFRTNEVMI 245
V+KFSG +++VV+EK + LR ++ +DG K LL RYN P WT M NEV+I
Sbjct: 508 VIKFSGKPTEDVVREKEKTLRSNIMKDGLKPELGCLLARYNDPGRTWTFTM--RNEVLI 338
>TC14894 weakly similar to UP|Q9LD82 (Q9LD82) ESTs AU029648, partial (84%)
Length = 982
Score = 40.4 bits (93), Expect = 3e-04
Identities = 42/160 (26%), Positives = 66/160 (41%), Gaps = 2/160 (1%)
Frame = +3
Query: 56 VETPKYEVTKTTQDYEIRIYAPSV--AAEVTYDPSQFKGNKDGGFMVLANYIGALGNPQN 113
+E P Y+V + YEIR Y +V + D S + + GF+ L NYI +N
Sbjct: 147 IECPSYDVIQVGNGYEIRRYNSTVWISNSPIQDISLVEATRT-GFLRLFNYI----QGKN 311
Query: 114 TKPEKIAMTAPVITKGSAEKIAMTAPVVTKSSEEGERNKMVTMQFILPSSYEKAEEAPKP 173
+KI MTAPV+++ S P S F++ K +A P
Sbjct: 312 DYSQKIEMTAPVLSEVSPS----DGPFCESS-------------FVVSFFVPKVNQANPP 440
Query: 174 TDERVVIREEGERKYGVVKFSGVASDEVVKEKVEKLRLSL 213
+ + ++ V +F G SD V E+ L+ S+
Sbjct: 441 PAKGLHVQRWKPVNVAVRQFGGFVSDASVGEEAAALKASI 560
>BP053014
Length = 441
Score = 26.9 bits (58), Expect = 3.3
Identities = 16/45 (35%), Positives = 23/45 (50%)
Frame = -3
Query: 165 EKAEEAPKPTDERVVIREEGERKYGVVKFSGVASDEVVKEKVEKL 209
E+A P P ++ EEGE YG + + +E+VKE KL
Sbjct: 232 EEAVLLPNPG----MLTEEGEEGYGKPRVMELGEEELVKEGCRKL 110
>TC11690 similar to UP|6PGL_DROME (Q9VZ64) Potential
6-phosphogluconolactonase (6PGL) , partial (16%)
Length = 542
Score = 26.9 bits (58), Expect = 3.3
Identities = 20/60 (33%), Positives = 27/60 (44%), Gaps = 6/60 (10%)
Frame = +3
Query: 108 LGNPQNTKPEKIAMTAPVITKGSAEKIAMT------APVVTKSSEEGERNKMVTMQFILP 161
L N PE+I T PVI SA IAM A V + E+G + + +Q + P
Sbjct: 9 LKNAPKPPPERITFTLPVI--NSASNIAMVVTGAGKADAVYSALEKGPTDYKLPVQLVSP 182
>TC18418
Length = 437
Score = 26.6 bits (57), Expect = 4.3
Identities = 29/130 (22%), Positives = 55/130 (42%), Gaps = 3/130 (2%)
Frame = +3
Query: 20 ATSTTHKNLLLPTSFNSHFTLSHSRSIMGMVFGKIGVETPKYEVTKTTQDYEIRIYAPSV 79
++S +HK L F SHF+LS S ++ + + T + + PS
Sbjct: 93 SSSHSHKTLSFFIEFVSHFSLSLSLTV-------------RQKQRTRTLSLSLSLAFPSF 233
Query: 80 AAEVTYDPSQFKGNKDGGFMVLANYIGALGNPQNTKPEKIAMTAPVITKGSAEK---IAM 136
A ++ S + GG +N+ G+ +++ P ++ AP + S ++ + M
Sbjct: 234 AIMISSRCSFLSNSGLGG---SSNFSGS--QRRHSAPLRVLCMAPSSSTKSGDRNGSVVM 398
Query: 137 TAPVVTKSSE 146
P +T S E
Sbjct: 399 ETPTLTPSKE 428
>TC19109 similar to UP|Q84LE0 (Q84LE0) Phytocyanin protein, PUP2, partial
(5%)
Length = 479
Score = 26.2 bits (56), Expect = 5.6
Identities = 9/24 (37%), Positives = 18/24 (74%)
Frame = +2
Query: 112 QNTKPEKIAMTAPVITKGSAEKIA 135
++ KP+K TAPV++ G+ E+++
Sbjct: 227 KDPKPKKTDATAPVVSSGARERVS 298
>AU251720
Length = 374
Score = 25.8 bits (55), Expect = 7.3
Identities = 11/31 (35%), Positives = 17/31 (54%)
Frame = -1
Query: 7 IQISYPLLYKGLKATSTTHKNLLLPTSFNSH 37
+++SY L AT +HK L+P +F H
Sbjct: 296 LEVSYSPCRHPLAATEKSHKQALVP*NFQKH 204
>TC8660 similar to UP|AAR28754 (AAR28754) Bax inhibitor, partial (97%)
Length = 1053
Score = 25.4 bits (54), Expect = 9.5
Identities = 19/69 (27%), Positives = 31/69 (44%)
Frame = -3
Query: 108 LGNPQNTKPEKIAMTAPVITKGSAEKIAMTAPVVTKSSEEGERNKMVTMQFILPSSYEKA 167
+ NP++T+P + + + TK + EK A V T + KM + S +
Sbjct: 529 MDNPEDTRPPRYKYSLRLATKAAPEKHAKAREVPTNAL------KMRLGSI*MAKSIKGP 368
Query: 168 EEAPKPTDE 176
EAP +DE
Sbjct: 367 IEAP*NSDE 341
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.315 0.133 0.376
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 3,442,876
Number of Sequences: 28460
Number of extensions: 39506
Number of successful extensions: 168
Number of sequences better than 10.0: 16
Number of HSP's better than 10.0 without gapping: 167
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 167
length of query: 248
length of database: 4,897,600
effective HSP length: 88
effective length of query: 160
effective length of database: 2,393,120
effective search space: 382899200
effective search space used: 382899200
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (22.0 bits)
S2: 54 (25.4 bits)
Medicago: description of AC124958.16


