

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC124953.2 + phase: 0
(53 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
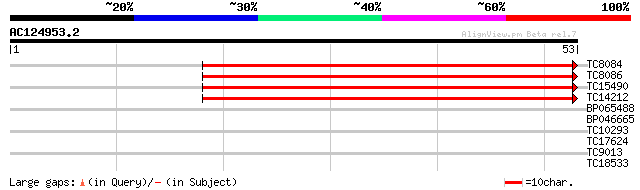
TC8084 similar to UP|PIF1_YEAST (P07271) DNA repair and recombin... 45 3e-06
TC8086 similar to UP|DMC1_HUMAN (Q14565) Meiotic recombination p... 45 3e-06
TC15490 weakly similar to UP|Q9LTU4 (Q9LTU4) Helicase-like prote... 41 6e-05
TC14212 similar to UP|Q9FN61 (Q9FN61) Gb|AAF07369.1, partial (10%) 38 5e-04
BP065488 37 0.001
BP046665 31 0.044
TC10293 similar to UP|Q84TG3 (Q84TG3) At2g35930, partial (19%) 28 0.49
TC17624 27 0.64
TC9013 27 0.84
TC18533 weakly similar to UP|Q9LUL3 (Q9LUL3) Genomic DNA, chromo... 23 9.3
>TC8084 similar to UP|PIF1_YEAST (P07271) DNA repair and recombination
protein PIF1, mitochondrial precursor, partial (4%)
Length = 560
Score = 45.1 bits (105), Expect = 3e-06
Identities = 20/35 (57%), Positives = 26/35 (74%)
Frame = +1
Query: 19 RVTSRKGLKILITDENGDCIDNTTNVVYKEVFQNV 53
RV SRKGLK+L+ DE + T NVVY+EVF+N+
Sbjct: 343 RVRSRKGLKLLVLDEEEKVTNTTKNVVYREVFENI 447
>TC8086 similar to UP|DMC1_HUMAN (Q14565) Meiotic recombination protein
DMC1/LIM15 homolog, partial (12%)
Length = 628
Score = 45.1 bits (105), Expect = 3e-06
Identities = 20/35 (57%), Positives = 26/35 (74%)
Frame = +3
Query: 19 RVTSRKGLKILITDENGDCIDNTTNVVYKEVFQNV 53
RV SRKGLK+L+ DE + T NVVY+EVF+N+
Sbjct: 372 RVRSRKGLKLLVLDEEEKVTNTTKNVVYREVFENI 476
>TC15490 weakly similar to UP|Q9LTU4 (Q9LTU4) Helicase-like protein, partial
(8%)
Length = 634
Score = 40.8 bits (94), Expect = 6e-05
Identities = 18/35 (51%), Positives = 25/35 (71%)
Frame = +2
Query: 19 RVTSRKGLKILITDENGDCIDNTTNVVYKEVFQNV 53
RV SRK LK+L+ DE + T NVVY+E+F+N+
Sbjct: 371 RVRSRK*LKLLVLDEEEKMTNTTKNVVYREIFENI 475
>TC14212 similar to UP|Q9FN61 (Q9FN61) Gb|AAF07369.1, partial (10%)
Length = 613
Score = 37.7 bits (86), Expect = 5e-04
Identities = 18/35 (51%), Positives = 23/35 (65%)
Frame = +2
Query: 19 RVTSRKGLKILITDENGDCIDNTTNVVYKEVFQNV 53
RV S+ GLKILI+ + T N+VYKEVFQ +
Sbjct: 20 RVKSKDGLKILISSDGTSTPGATKNIVYKEVFQKI 124
>BP065488
Length = 439
Score = 36.6 bits (83), Expect = 0.001
Identities = 17/34 (50%), Positives = 23/34 (67%)
Frame = -1
Query: 20 VTSRKGLKILITDENGDCIDNTTNVVYKEVFQNV 53
V SRKGLK+L+ E + T N VY+EVF+N+
Sbjct: 169 VRSRKGLKLLVLGEEEKVTNATKNEVYQEVFKNI 68
>BP046665
Length = 524
Score = 31.2 bits (69), Expect = 0.044
Identities = 14/25 (56%), Positives = 17/25 (68%)
Frame = -2
Query: 22 SRKGLKILITDENGDCIDNTTNVVY 46
SRKGLK+L+ DE + T NVVY
Sbjct: 142 SRKGLKLLVLDEEEKVTNTTKNVVY 68
>TC10293 similar to UP|Q84TG3 (Q84TG3) At2g35930, partial (19%)
Length = 757
Score = 27.7 bits (60), Expect = 0.49
Identities = 13/31 (41%), Positives = 20/31 (63%)
Frame = +3
Query: 22 SRKGLKILITDENGDCIDNTTNVVYKEVFQN 52
S GLK L++ +NGD I + T V+ K F++
Sbjct: 654 SEPGLKTLLSFKNGDFIASLTRVMQKGFFES 746
>TC17624
Length = 615
Score = 27.3 bits (59), Expect = 0.64
Identities = 11/14 (78%), Positives = 14/14 (99%)
Frame = +1
Query: 40 NTTNVVYKEVFQNV 53
+T+NVVYKEVF+NV
Sbjct: 532 STSNVVYKEVFRNV 573
>TC9013
Length = 766
Score = 26.9 bits (58), Expect = 0.84
Identities = 13/26 (50%), Positives = 18/26 (69%)
Frame = +3
Query: 28 ILITDENGDCIDNTTNVVYKEVFQNV 53
ILI D + ++T+NVVY EVF+ V
Sbjct: 627 ILICDGDDSNSNSTSNVVYTEVFRTV 704
>TC18533 weakly similar to UP|Q9LUL3 (Q9LUL3) Genomic DNA, chromosome 3, P1
clone: MLN21, partial (4%)
Length = 669
Score = 23.5 bits (49), Expect = 9.3
Identities = 9/14 (64%), Positives = 9/14 (64%)
Frame = -3
Query: 13 FFWADYRVTSRKGL 26
FFW Y VT KGL
Sbjct: 133 FFWFQYHVTVSKGL 92
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.327 0.140 0.436
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 926,039
Number of Sequences: 28460
Number of extensions: 7650
Number of successful extensions: 46
Number of sequences better than 10.0: 20
Number of HSP's better than 10.0 without gapping: 46
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 46
length of query: 53
length of database: 4,897,600
effective HSP length: 29
effective length of query: 24
effective length of database: 4,072,260
effective search space: 97734240
effective search space used: 97734240
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.1 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.7 bits)
S2: 49 (23.5 bits)
Medicago: description of AC124953.2


