

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC124953.18 + phase: 0
(391 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
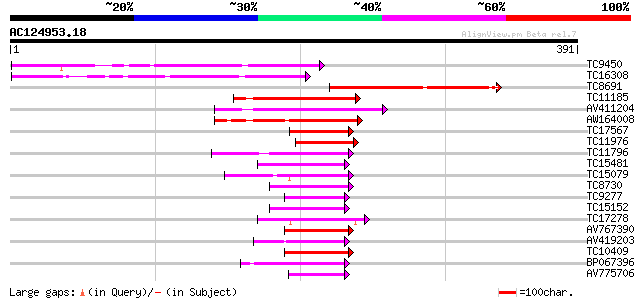
Score E
Sequences producing significant alignments: (bits) Value
TC9450 similar to GB|AAM16172.1|20334832|AY094016 At2g39720/T5I7... 169 1e-42
TC16308 similar to GB|AAM16172.1|20334832|AY094016 At2g39720/T5I... 158 1e-39
TC8691 similar to UP|Q940T5 (Q940T5) AT5g59550/f2o15_210, partia... 108 2e-24
TC11185 similar to UP|Q9SXW6 (Q9SXW6) Ring finger protein (Fragm... 90 6e-19
AV411204 87 6e-18
AW164008 86 1e-17
TC17567 similar to UP|Q8RXD3 (Q8RXD3) ABI3-interacting protein 2... 59 2e-09
TC11976 similar to GB|AAP12856.1|30017245|BT006207 At3g02340 {Ar... 57 4e-09
TC11796 similar to GB|AAP12870.1|30017273|BT006221 At4g35840 {Ar... 56 1e-08
TC15481 similar to UP|Q9LUR1 (Q9LUR1) RING zinc finger protein-l... 54 3e-08
TC15079 similar to UP|Q84KA9 (Q84KA9) RING/C3HC4/PHD zinc finger... 54 6e-08
TC8730 similar to PIR|H96764|H96764 protein RING zinc finger pro... 53 8e-08
TC9277 similar to GB|AAP13413.1|30023760|BT006305 At3g55530 {Ara... 53 1e-07
TC15152 similar to UP|Q8RZY3 (Q8RZY3) RING zinc finger protein-l... 52 1e-07
TC17278 similar to UP|Q8GT74 (Q8GT74) NEP1-interacting protein 2... 52 2e-07
AV767390 52 2e-07
AV419203 51 3e-07
TC10409 similar to UP|Q8GUU2 (Q8GUU2) RES protein, partial (17%) 51 3e-07
BP067396 51 4e-07
AV775706 51 4e-07
>TC9450 similar to GB|AAM16172.1|20334832|AY094016 At2g39720/T5I7.2
{Arabidopsis thaliana;}, partial (22%)
Length = 646
Score = 169 bits (427), Expect = 1e-42
Identities = 91/218 (41%), Positives = 132/218 (59%), Gaps = 2/218 (0%)
Frame = +2
Query: 2 SSHWCYRCNKFVRVWRLGMPICPDCDSGFLEDV--EQSTHSANTVGGRRMRFPMAAAMYM 59
+S+WCY C +FV + G CP C++GF+E++ +QS + A+ G + ++ +
Sbjct: 53 ASYWCYSCTRFVHLSAAGNIACPHCETGFVEEIHPDQSPNHAHAHGLSPLPEDPVSSRRL 232
Query: 60 IGHRNNNYNQNTFRRHRRNNVNGGDISPFNPIIMIRGGGGSSEGTSREREENNEFELFYE 119
FRR RR+ G SPFNP+I++RG + +G +E FEL+Y+
Sbjct: 233 -----------GFRRRRRDT---GSRSPFNPVIVLRGT--ADDGADQEGGAAAAFELYYD 364
Query: 120 DGAGSGLRALPPRMSELILGSGFERVMEQLSHVEANRSGNEGHNQQHLPALKSAVELLPT 179
DG G+GLR LPP +SE +LGSGF+R++EQ S +E N G ++ PA K+A+E +PT
Sbjct: 365 DGDGTGLRPLPPTVSEFLLGSGFDRMLEQFSQIEMNGFGRP----ENPPASKAAIESMPT 532
Query: 180 IEINESHMNVESHCAVCKEPFELGISAREMPCKHIYHN 217
+EI E E+ CAVCKE F LG ARE+PCKHIYH+
Sbjct: 533 VEIGEMEACNEAQCAVCKEEF*LGAEARELPCKHIYHS 646
>TC16308 similar to GB|AAM16172.1|20334832|AY094016 At2g39720/T5I7.2
{Arabidopsis thaliana;}, partial (13%)
Length = 640
Score = 158 bits (400), Expect = 1e-39
Identities = 85/206 (41%), Positives = 122/206 (58%)
Frame = +3
Query: 2 SSHWCYRCNKFVRVWRLGMPICPDCDSGFLEDVEQSTHSANTVGGRRMRFPMAAAMYMIG 61
SS+WCY CN+FV + +CP C SGF+E++ HS A +++ G
Sbjct: 96 SSYWCYSCNRFVHLLDHNDVVCPHCQSGFVEEIHAG-HSP------------AVSLFADG 236
Query: 62 HRNNNYNQNTFRRHRRNNVNGGDISPFNPIIMIRGGGGSSEGTSREREENNEFELFYEDG 121
+ + FRR RRN G SPFNP+I++RG G + G + + FELFY+DG
Sbjct: 237 IHASR--RQGFRRRRRN---AGSRSPFNPVIVLRGPGDDAAGADND---GSSFELFYDDG 392
Query: 122 AGSGLRALPPRMSELILGSGFERVMEQLSHVEANRSGNEGHNQQHLPALKSAVELLPTIE 181
G+GLR LPP MSEL+LGSGF+R+++Q S +E N G ++ A K+ +E +PT+E
Sbjct: 393 DGTGLRPLPPTMSELLLGSGFDRLLDQFSQIEINGFG----RPENPXASKAXIESMPTVE 560
Query: 182 INESHMNVESHCAVCKEPFELGISAR 207
I E+ + E+HCAVCK+ FE+ AR
Sbjct: 561 ITEAEVQAETHCAVCKDAFEMXXEAR 638
>TC8691 similar to UP|Q940T5 (Q940T5) AT5g59550/f2o15_210, partial (16%)
Length = 942
Score = 108 bits (269), Expect = 2e-24
Identities = 59/119 (49%), Positives = 76/119 (63%)
Frame = +2
Query: 221 LPWLAIQNSCPVCRHELPCESPQINNEISNSNEDENVGLTIWRLPGGGFAVGRFSGDGGG 280
+PWL+++NSCPVCRHELP + ++E +GLTIWRLPGGGFAVGRFSG
Sbjct: 2 VPWLSMRNSCPVCRHELPSDLNANPLRAPGQIDEEAIGLTIWRLPGGGFAVGRFSGGRSA 181
Query: 281 GENRMEHPIVYTEVDGAFNNVGEPRRISWSLTSSRGGIGRSRGGAFRRMLSNLFGCLRG 339
GE+ + P VYTE+DG N+ PRR+S +++ R R G FR LS FG +RG
Sbjct: 182 GESHL--PEVYTEMDGGVNSNRAPRRVSPTVSGHRVRERRGVGRVFRGFLS-FFG-IRG 346
>TC11185 similar to UP|Q9SXW6 (Q9SXW6) Ring finger protein (Fragment),
partial (79%)
Length = 807
Score = 90.1 bits (222), Expect = 6e-19
Identities = 42/89 (47%), Positives = 55/89 (61%), Gaps = 1/89 (1%)
Frame = +2
Query: 155 NRSGNEGHNQQHLPALKSAVELLPTIEINESHMN-VESHCAVCKEPFELGISAREMPCKH 213
N SG +G PA KS VE LP +E+ E + + CA+CK+ L R +PC H
Sbjct: 242 NESGLKGSP----PAAKSFVESLPLVELTEEELRGKDMACAICKDEIMLEEKVRRLPCSH 409
Query: 214 IYHNECILPWLAIQNSCPVCRHELPCESP 242
YH +CILPWL+I+N+CPVCR ELP + P
Sbjct: 410 CYHGDCILPWLSIRNTCPVCRFELPTDDP 496
>AV411204
Length = 427
Score = 86.7 bits (213), Expect = 6e-18
Identities = 44/120 (36%), Positives = 65/120 (53%), Gaps = 1/120 (0%)
Frame = +2
Query: 142 FERVMEQLSHVEANRSGNEGHNQQHLPALKSAVELLPTIEINESH-MNVESHCAVCKEPF 200
FE ++E L+ +++R G PA S V LP I I + H + E CA+CK+
Sbjct: 89 FEDLLEHLAENDSSRRGAP-------PAAVSFVNNLPRIVIGKEHEKHGELVCAICKDVL 247
Query: 201 ELGISAREMPCKHIYHNECILPWLAIQNSCPVCRHELPCESPQINNEISNSNEDENVGLT 260
G ++PC H+YH CILPWL+ +NSCP+CR+ELP + NS+ + +T
Sbjct: 248 APGTEVNQLPCSHLYHPSCILPWLSARNSCPLCRYELPTDDKDYEEGKQNSDAGNVIHVT 427
>AW164008
Length = 351
Score = 85.5 bits (210), Expect = 1e-17
Identities = 43/102 (42%), Positives = 65/102 (63%)
Frame = +3
Query: 142 FERVMEQLSHVEANRSGNEGHNQQHLPALKSAVELLPTIEINESHMNVESHCAVCKEPFE 201
+E +M+ L+ E++ G +G PA K+AVE LPT++I V CA+CK+
Sbjct: 69 YEALMQILA--ESDGGGRKGAP----PASKAAVEALPTVKIISESDTVA--CAICKDLMG 224
Query: 202 LGISAREMPCKHIYHNECILPWLAIQNSCPVCRHELPCESPQ 243
+G SA+ +PC H YH +CI+PWL+ +NSCPVCR +LP + +
Sbjct: 225 VGQSAKRLPCGHQYHGDCIVPWLSSRNSCPVCRFQLPTDDKE 350
>TC17567 similar to UP|Q8RXD3 (Q8RXD3) ABI3-interacting protein 2, partial
(25%)
Length = 511
Score = 58.5 bits (140), Expect = 2e-09
Identities = 21/44 (47%), Positives = 30/44 (67%)
Frame = +1
Query: 194 AVCKEPFELGISAREMPCKHIYHNECILPWLAIQNSCPVCRHEL 237
A+C+E + +E+PCKH +H C+ PWL NSCP+CR+EL
Sbjct: 1 AICRENLVVNDKMQELPCKHTFHPPCLKPWLDEHNSCPICRYEL 132
>TC11976 similar to GB|AAP12856.1|30017245|BT006207 At3g02340 {Arabidopsis
thaliana;}, partial (11%)
Length = 530
Score = 57.4 bits (137), Expect = 4e-09
Identities = 20/43 (46%), Positives = 30/43 (69%)
Frame = +3
Query: 198 EPFELGISAREMPCKHIYHNECILPWLAIQNSCPVCRHELPCE 240
+ F +G + +PC H Y +CI+PWL I+N+CPVCR+E P +
Sbjct: 3 DDFVVGEEVKPLPCSHRYPGDCIVPWLGIRNTCPVCRYEFPTD 131
>TC11796 similar to GB|AAP12870.1|30017273|BT006221 At4g35840 {Arabidopsis
thaliana;}, partial (86%)
Length = 1114
Score = 55.8 bits (133), Expect = 1e-08
Identities = 30/99 (30%), Positives = 46/99 (46%), Gaps = 1/99 (1%)
Frame = +1
Query: 140 SGFERVMEQLSHVEANRSGNEGHNQQHLPALKSAVELLPTIEINESHMNVESHCAVCKEP 199
+GF+ V G G + + +P +K T + N C+VC +
Sbjct: 469 TGFDEVQNIFDTGCGGAKGLSGDSVEKIPKIKI------TTDNNADASGERVSCSVCLQD 630
Query: 200 FELGISAREMP-CKHIYHNECILPWLAIQNSCPVCRHEL 237
F+LG + R +P C H++H CI WL SCP+CR +L
Sbjct: 631 FQLGETVRSLPHCHHMFHLPCIDKWLFRHGSCPLCRRDL 747
>TC15481 similar to UP|Q9LUR1 (Q9LUR1) RING zinc finger protein-like,
partial (32%)
Length = 706
Score = 54.3 bits (129), Expect = 3e-08
Identities = 25/65 (38%), Positives = 36/65 (54%), Gaps = 2/65 (3%)
Frame = +1
Query: 172 SAVELLPTIEIN-ESHMNVESHCAVCKEPFELGISAREMP-CKHIYHNECILPWLAIQNS 229
S + LLP + ++H CAVC FE G + R +P CKH +H +CI W ++
Sbjct: 358 SVITLLPVFSYDPKAHPEGPPECAVCLSEFESGETGRVLPKCKHAFHTDCIDMWFHSHST 537
Query: 230 CPVCR 234
CP+CR
Sbjct: 538 CPLCR 552
>TC15079 similar to UP|Q84KA9 (Q84KA9) RING/C3HC4/PHD zinc finger-like
protein, partial (20%)
Length = 921
Score = 53.5 bits (127), Expect = 6e-08
Identities = 33/96 (34%), Positives = 46/96 (47%), Gaps = 7/96 (7%)
Frame = +3
Query: 149 LSHVEANRSGNEGHNQQHLPALKSAVELLPTIEINESHMNVES------HCAVCKEPFEL 202
L+H A SG+ N+ + ++ PT SH+ CAVC FE
Sbjct: 480 LAHAAAGGSGHRQLNELSNGLNQEVIDTFPTFLY--SHVKCLKIGKGTLACAVCLNEFED 653
Query: 203 GISAREMP-CKHIYHNECILPWLAIQNSCPVCRHEL 237
+ R +P C H+YH+ CI WLA ++CPVCR L
Sbjct: 654 DETLRLIPICNHVYHHSCIDLWLASHSTCPVCRASL 761
>TC8730 similar to PIR|H96764|H96764 protein RING zinc finger protein
F25P22.18 [imported] - Arabidopsis
thaliana {Arabidopsis thaliana;}, partial (23%)
Length = 583
Score = 53.1 bits (126), Expect = 8e-08
Identities = 19/58 (32%), Positives = 34/58 (57%)
Frame = +3
Query: 180 IEINESHMNVESHCAVCKEPFELGISAREMPCKHIYHNECILPWLAIQNSCPVCRHEL 237
I + S ++ C +C+E +E ++ C+H +H +CI WL ++N CPVC+ E+
Sbjct: 168 ISNDTSKHQLDRKCTICQEEYESDDELGKLKCEHCFHFQCIKQWLVLKNFCPVCKQEV 341
>TC9277 similar to GB|AAP13413.1|30023760|BT006305 At3g55530 {Arabidopsis
thaliana;}, partial (79%)
Length = 1297
Score = 52.8 bits (125), Expect = 1e-07
Identities = 21/45 (46%), Positives = 27/45 (59%)
Frame = +3
Query: 190 ESHCAVCKEPFELGISAREMPCKHIYHNECILPWLAIQNSCPVCR 234
E C+VC E +G R +PC H +H CI PWL Q +CPVC+
Sbjct: 873 ELTCSVCLEQVNVGDILRSLPCLHQFHANCIDPWLRQQGTCPVCK 1007
>TC15152 similar to UP|Q8RZY3 (Q8RZY3) RING zinc finger protein-like,
partial (11%)
Length = 779
Score = 52.4 bits (124), Expect = 1e-07
Identities = 18/55 (32%), Positives = 33/55 (59%)
Frame = +3
Query: 180 IEINESHMNVESHCAVCKEPFELGISAREMPCKHIYHNECILPWLAIQNSCPVCR 234
++I++ V+ +C +C+E +E+ + C H YH +CI W+ +N CPVC+
Sbjct: 297 LQISDDASQVDKNCTICQEEYEVEEEIGRLNCGHSYHYQCIQQWVEQKNFCPVCK 461
>TC17278 similar to UP|Q8GT74 (Q8GT74) NEP1-interacting protein 2, partial
(62%)
Length = 610
Score = 52.0 bits (123), Expect = 2e-07
Identities = 27/84 (32%), Positives = 47/84 (55%), Gaps = 7/84 (8%)
Frame = +3
Query: 172 SAVELLPTIEINESHMNVESH---CAVCKEPFELGISAREMP-CKHIYHNECILPWLAIQ 227
++V +P + I ++ + C+VC + F+LG + R +P C H++H CI WL+
Sbjct: 246 ASVAKIPQVTITGNNGDASGQRDSCSVCLQDFQLGETVRSLPYCHHMFHLPCIDEWLSKH 425
Query: 228 NSCPVCRHEL---PCESPQINNEI 248
SCP+CR ++ C S +I + I
Sbjct: 426 VSCPLCRRDM*FCICTSLKIGSTI 497
>AV767390
Length = 447
Score = 51.6 bits (122), Expect = 2e-07
Identities = 16/48 (33%), Positives = 30/48 (62%)
Frame = -3
Query: 190 ESHCAVCKEPFELGISAREMPCKHIYHNECILPWLAIQNSCPVCRHEL 237
++ C +C ++ G+ RE+PC H +H C+ WL I +CP+C++ +
Sbjct: 439 DAECCICLSSYDDGVELRELPCGHHFHCGCVDKWLHINATCPLCKYNI 296
>AV419203
Length = 393
Score = 51.2 bits (121), Expect = 3e-07
Identities = 24/66 (36%), Positives = 34/66 (51%)
Frame = +1
Query: 169 ALKSAVELLPTIEINESHMNVESHCAVCKEPFELGISAREMPCKHIYHNECILPWLAIQN 228
A+++ ++ LP + + S C VC E F G R +PC H +H ECI WL +
Sbjct: 187 AVEALIQELPKFRLKAVPTDC-SECPVCLEEFHEGNEVRGLPCAHNFHVECIDEWLRLNV 363
Query: 229 SCPVCR 234
CP CR
Sbjct: 364 KCPRCR 381
>TC10409 similar to UP|Q8GUU2 (Q8GUU2) RES protein, partial (17%)
Length = 715
Score = 51.2 bits (121), Expect = 3e-07
Identities = 15/48 (31%), Positives = 29/48 (60%)
Frame = +3
Query: 190 ESHCAVCKEPFELGISAREMPCKHIYHNECILPWLAIQNSCPVCRHEL 237
++ C +C +E G +PC H +H+ CI+ WL + +CP+C++ +
Sbjct: 36 DAECCICISSYEDGAELHSLPCNHHFHSTCIVKWLKMNATCPLCKYNI 179
>BP067396
Length = 506
Score = 50.8 bits (120), Expect = 4e-07
Identities = 24/76 (31%), Positives = 42/76 (54%), Gaps = 1/76 (1%)
Frame = -1
Query: 160 EGHNQQHLPALKSAVELLPTIEINESHMNVESHCAVCKEPFELGISAREMP-CKHIYHNE 218
+ HN + + +S +LL + S +V S C++C ++ + R +P C H+YH
Sbjct: 479 DDHNMHN--SFESYPKLLYSEVEKRSDSSVSSCCSICLGDYKEADTLRMLPDCGHVYHLA 306
Query: 219 CILPWLAIQNSCPVCR 234
C+ PWL ++CP+CR
Sbjct: 305 CVDPWLKFHSTCPICR 258
>AV775706
Length = 462
Score = 50.8 bits (120), Expect = 4e-07
Identities = 19/42 (45%), Positives = 23/42 (54%)
Frame = -2
Query: 193 CAVCKEPFELGISAREMPCKHIYHNECILPWLAIQNSCPVCR 234
CAVC E F+ + C H YH+ C+LPWL CP CR
Sbjct: 293 CAVCLEDFQPKQQVMNLSCSHTYHSACLLPWLEAHPPCPCCR 168
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.318 0.136 0.431
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 8,054,415
Number of Sequences: 28460
Number of extensions: 125718
Number of successful extensions: 1039
Number of sequences better than 10.0: 127
Number of HSP's better than 10.0 without gapping: 993
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 1024
length of query: 391
length of database: 4,897,600
effective HSP length: 92
effective length of query: 299
effective length of database: 2,279,280
effective search space: 681504720
effective search space used: 681504720
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 56 (26.2 bits)
Medicago: description of AC124953.18


