

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
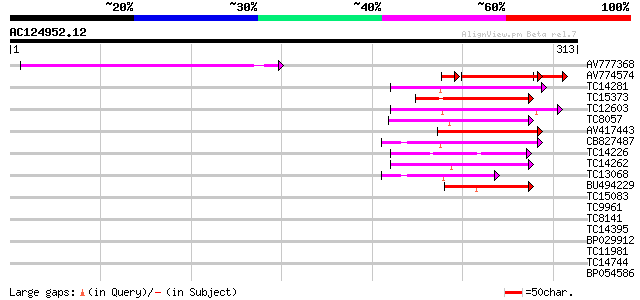
Query= AC124952.12 + phase: 2 /pseudo
(313 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
AV777368 119 9e-28
AV774574 64 4e-14
TC14281 similar to UP|Q9XHC6 (Q9XHC6) Cytochrome P450 monooxygen... 59 1e-09
TC15373 weakly similar to UP|Q9STH9 (Q9STH9) Cytochrome P450 hom... 54 3e-08
TC12603 weakly similar to UP|AAP49697 (AAP49697) Cytochrome P-45... 54 3e-08
TC8057 weakly similar to UP|Q8W228 (Q8W228) Cytochrome P450, par... 52 1e-07
AV417443 51 2e-07
CB827487 51 3e-07
TC14226 UP|Q9MBE4 (Q9MBE4) Cytochrome P450, complete 44 4e-05
TC14262 UP|Q9MBE5 (Q9MBE5) Cytochrome P450, complete 43 6e-05
TC13068 weakly similar to UP|O48924 (O48924) CYP83D1p (Fragment)... 40 4e-04
BU494229 39 9e-04
TC15083 similar to UP|GAR1_YEAST (P28007) snoRNP protein GAR1, p... 35 0.016
TC9961 homologue to UP|C982_SOYBN (O48922) Cytochrome P450 98A2 ... 34 0.028
TC8141 weakly similar to UP|C79B_ARATH (O81346) Cytochrome P450 ... 33 0.062
TC14395 weakly similar to UP|C823_SOYBN (O49858) Cytochrome P450... 33 0.081
BP029912 31 0.31
TC11981 27 5.8
TC14744 UP|C7DB_LOTJA (O22307) Cytochrome P450 71D11 (Fragment)... 27 5.8
BP054586 27 5.8
>AV777368
Length = 604
Score = 119 bits (297), Expect = 9e-28
Identities = 69/145 (47%), Positives = 81/145 (55%)
Frame = +2
Query: 7 YFHPHLDCQ*LATTFN*EHFLIALFNPSHKSMVL**CCILVSFLCWWFPQYIWPRKSCKH 66
+ H H +C LA + N*E FLIALF S KS VL CCI V F CWWFP WP K CK
Sbjct: 176 FLHLHQNCHSLAISIN*EQFLIALFKLSRKSTVLCCCCIWVIFPCWWFPLQTWPEK*CKP 355
Query: 67 MA*FLRTDQARH*QKPYSMVVRTLRFHLMAIHGDRKRNCV*MNC*VRKGCNQYSLLEKSN 126
M F + D H QKPYS T + GDRK N V +N * KGC+ +SL EK
Sbjct: 356 MTQFFQADPTSHQQKPYSTGAMT*DLRPTVMRGDRKGNFVCLNY*ALKGCSLFSLSEKRK 535
Query: 127 LLHWLIRYVKL*V*VVVVAV*TLVR 151
WL +Y V+V+A+*TL R
Sbjct: 536 *QLWLRKYA-----VMVLAL*TLGR 595
>AV774574
Length = 510
Score = 64.3 bits (155), Expect(3) = 4e-14
Identities = 30/45 (66%), Positives = 39/45 (86%)
Frame = +1
Query: 250 LQDDGLTEFELTQNDLKALLMNMFLGGSDTTSTTLEWAMAACKKS 294
LQ+ G++EFELTQ+DLKALL++M L G+DT+STTLEWAMA K+
Sbjct: 64 LQEGGMSEFELTQDDLKALLLDMCLAGTDTSSTTLEWAMAELVKN 198
Score = 27.7 bits (60), Expect(3) = 4e-14
Identities = 12/19 (63%), Positives = 15/19 (78%)
Frame = +3
Query: 290 ACKKSSHNEESTGGGKENC 308
A +KSSHNEES GK++C
Sbjct: 186 AGEKSSHNEESPRRGKKSC 242
Score = 21.2 bits (43), Expect(3) = 4e-14
Identities = 8/10 (80%), Positives = 10/10 (100%)
Frame = +2
Query: 239 KKKDFLDILL 248
++KDFLDILL
Sbjct: 32 RRKDFLDILL 61
>TC14281 similar to UP|Q9XHC6 (Q9XHC6) Cytochrome P450 monooxygenaseCYP93D1,
partial (97%)
Length = 1768
Score = 58.9 bits (141), Expect = 1e-09
Identities = 30/91 (32%), Positives = 54/91 (58%), Gaps = 5/91 (5%)
Frame = +3
Query: 211 KKIKDSSEEMDDFLDRVIVEHKMSRR-----DPKKKDFLDILLQLQDDGLTEFELTQNDL 265
KK KD ++D +++V+ EH+ +R +KKD DILL L + + +LT++
Sbjct: 762 KKNKDMHRKLDGMMEKVLKEHEEARATEGADSDRKKDLFDILLNLIEADGADNKLTRDSA 941
Query: 266 KALLMNMFLGGSDTTSTTLEWAMAACKKSSH 296
KA ++MFL G++ ++ LEW++A ++ H
Sbjct: 942 KAFALDMFLAGTNGPASVLEWSLAELIRNPH 1034
>TC15373 weakly similar to UP|Q9STH9 (Q9STH9) Cytochrome P450 homolog,
partial (61%)
Length = 1059
Score = 54.3 bits (129), Expect = 3e-08
Identities = 30/65 (46%), Positives = 45/65 (69%)
Frame = +1
Query: 225 DRVIVEHKMSRRDPKKKDFLDILLQLQDDGLTEFELTQNDLKALLMNMFLGGSDTTSTTL 284
+RV +E + R + KDFL LL L+++G ++ L+ +KALLM+M GGSDT+S T+
Sbjct: 196 ERVKMESEGKRSE--SKDFLQFLLNLKEEGDSKTPLSITHVKALLMDMLAGGSDTSSNTV 369
Query: 285 EWAMA 289
E+AMA
Sbjct: 370 EFAMA 384
>TC12603 weakly similar to UP|AAP49697 (AAP49697) Cytochrome P-450-like
protein (Fragment), partial (24%)
Length = 364
Score = 53.9 bits (128), Expect = 3e-08
Identities = 32/109 (29%), Positives = 56/109 (51%), Gaps = 14/109 (12%)
Frame = +1
Query: 211 KKIKDSSEEMDDFLDRVIVEHKMSRRD-------PKKKDFLDILLQLQDDGLTEFELTQN 263
KK+K +S+E+DD + +HK R++ + DF+D+ L + DD
Sbjct: 22 KKMKKTSKELDDLATVWLEQHKQKRKNNGSGEWKKGEYDFMDVFLSIVDDEGFHGHDGDT 201
Query: 264 DLKALLMNMFLGGSDTTSTTLEWAMA-------ACKKSSHNEESTGGGK 305
+KA + + L G+DTT+ TL WA++ A K++H ++ GG+
Sbjct: 202 IIKATCLQLILAGTDTTTATLTWAISLLLNNREALNKATHEIDTQMGGR 348
>TC8057 weakly similar to UP|Q8W228 (Q8W228) Cytochrome P450, partial (29%)
Length = 1148
Score = 52.4 bits (124), Expect = 1e-07
Identities = 27/85 (31%), Positives = 47/85 (54%), Gaps = 5/85 (5%)
Frame = +2
Query: 210 LKKIKDSSEEMDDFLDRVIVEHKMSRRDPKKK-----DFLDILLQLQDDGLTEFELTQND 264
++K K + EE D L+++I +H+ R +K DF D LL + + + +
Sbjct: 56 VRKFKKAHEEFDQMLEQIIKDHEAPSRLSDQKNGQSIDFTDTLLSHMKQSKDKHIINKTN 235
Query: 265 LKALLMNMFLGGSDTTSTTLEWAMA 289
LKA+L+++ +G DTT T +WAM+
Sbjct: 236 LKAILLDIIIGSLDTTIMTTDWAMS 310
>AV417443
Length = 421
Score = 51.2 bits (121), Expect = 2e-07
Identities = 22/58 (37%), Positives = 39/58 (66%)
Frame = +2
Query: 237 DPKKKDFLDILLQLQDDGLTEFELTQNDLKALLMNMFLGGSDTTSTTLEWAMAACKKS 294
D + D +DI L++ ++ F+LT + +KA+LMN+F+ G+DT+S + WAM + K+
Sbjct: 29 DKEVADIIDIFLEIMNNHSLSFDLTIDHIKAVLMNIFIAGTDTSSALVVWAMTSLLKN 202
>CB827487
Length = 569
Score = 50.8 bits (120), Expect = 3e-07
Identities = 30/92 (32%), Positives = 48/92 (51%), Gaps = 3/92 (3%)
Frame = +3
Query: 206 SLAKLKKIKDSSEEMDDFLDRVIVEHKMSRR---DPKKKDFLDILLQLQDDGLTEFELTQ 262
SLA+L K +S D F +V+ EH R ++ D +D LLQL++ G +LT
Sbjct: 21 SLARLDKTINS---FDAFFQQVLDEHLDPNRIKDQTQEDDIVDTLLQLRNQGSLSIDLTD 191
Query: 263 NDLKALLMNMFLGGSDTTSTTLEWAMAACKKS 294
+KA +M++ +G +DT+ W M K+
Sbjct: 192 EHIKAFMMDLLIGSTDTSVAASVWLMTGLMKN 287
>TC14226 UP|Q9MBE4 (Q9MBE4) Cytochrome P450, complete
Length = 1878
Score = 43.9 bits (102), Expect = 4e-05
Identities = 25/78 (32%), Positives = 43/78 (55%)
Frame = +3
Query: 211 KKIKDSSEEMDDFLDRVIVEHKMSRRDPKKKDFLDILLQLQDDGLTEFELTQNDLKALLM 270
K++K S + D FL ++ EH+ ++ +D LL LQ+ + T +K L +
Sbjct: 765 KRVKGISSKTDRFLRGLLQEHR-DKKQRTANTMIDHLLTLQESQPEYY--TDQIIKGLAL 935
Query: 271 NMFLGGSDTTSTTLEWAM 288
M L G+D+++ TLEW+M
Sbjct: 936 AMLLAGTDSSAVTLEWSM 989
>TC14262 UP|Q9MBE5 (Q9MBE5) Cytochrome P450, complete
Length = 1960
Score = 43.1 bits (100), Expect = 6e-05
Identities = 23/90 (25%), Positives = 48/90 (52%), Gaps = 11/90 (12%)
Frame = +1
Query: 211 KKIKDSSEEMDDFLDRVIVEHKMSRRDPKKKD-----------FLDILLQLQDDGLTEFE 259
K+I + + D +++VI + + + K+++ FLD LL+ D E +
Sbjct: 769 KRIDEIFNKFDPVIEKVIKKRQEIIKRRKERNGELEEGEQSVVFLDTLLEYAADENMEIK 948
Query: 260 LTQNDLKALLMNMFLGGSDTTSTTLEWAMA 289
+T+ +K L+++ F G+D+T+ +WA+A
Sbjct: 949 ITKEQIKGLVVDFFSAGTDSTAVATDWALA 1038
>TC13068 weakly similar to UP|O48924 (O48924) CYP83D1p (Fragment), partial
(5%)
Length = 544
Score = 40.4 bits (93), Expect = 4e-04
Identities = 26/68 (38%), Positives = 39/68 (57%), Gaps = 3/68 (4%)
Frame = +1
Query: 206 SLAKLKKIKDSSEEMDDFLDRVIVEHKMSRRDP---KKKDFLDILLQLQDDGLTEFELTQ 262
SLA L K +S D F RV+ EH R+ +++D +DILLQL++ G +LT
Sbjct: 199 SLAHLDKTINS---FDAFFQRVLDEHLDPNRNKDQTQEEDIVDILLQLRNQGSLSIDLTN 369
Query: 263 NDLKALLM 270
+ +KA +M
Sbjct: 370 DHIKAFMM 393
>BU494229
Length = 412
Score = 39.3 bits (90), Expect = 9e-04
Identities = 18/51 (35%), Positives = 33/51 (64%), Gaps = 2/51 (3%)
Frame = +1
Query: 241 KDFLDILLQLQDDGLT--EFELTQNDLKALLMNMFLGGSDTTSTTLEWAMA 289
+D +D+LL+LQ + ++ LT ++LKA++ +MF G +T S + W M+
Sbjct: 82 EDLVDVLLRLQKEQQEPDQYALTDDNLKAVIQDMFAAGGETISEVVLWVMS 234
>TC15083 similar to UP|GAR1_YEAST (P28007) snoRNP protein GAR1, partial (7%)
Length = 1367
Score = 35.0 bits (79), Expect = 0.016
Identities = 22/79 (27%), Positives = 42/79 (52%), Gaps = 3/79 (3%)
Frame = +1
Query: 218 EEMDDFLDRVIVEHKMSRRDPKKKD---FLDILLQLQDDGLTEFELTQNDLKALLMNMFL 274
++ +D L +I K+++ D ++D LL+L+ + +L ++++ L +
Sbjct: 766 KDQEDVLVPLIRARKLAKEGKPVNDVVSYVDTLLELELPE-EKRKLNESEMVTLCSEILN 942
Query: 275 GGSDTTSTTLEWAMAACKK 293
G+DTTST LEW +A K
Sbjct: 943 AGTDTTSTALEWVLANLDK 999
>TC9961 homologue to UP|C982_SOYBN (O48922) Cytochrome P450 98A2 , partial
(45%)
Length = 846
Score = 34.3 bits (77), Expect = 0.028
Identities = 14/38 (36%), Positives = 26/38 (67%)
Frame = +1
Query: 257 EFELTQNDLKALLMNMFLGGSDTTSTTLEWAMAACKKS 294
+++L+++ + LL +M G DTT+ ++EWAMA K+
Sbjct: 19 KYDLSEDTIIGLLWDMITAGMDTTAISVEWAMAELIKN 132
>TC8141 weakly similar to UP|C79B_ARATH (O81346) Cytochrome P450 79B2 ,
partial (60%)
Length = 1919
Score = 33.1 bits (74), Expect = 0.062
Identities = 24/81 (29%), Positives = 39/81 (47%), Gaps = 3/81 (3%)
Frame = +2
Query: 212 KIKDSSEEMDDFLDRVIVEHKMSRRDPKK---KDFLDILLQLQDDGLTEFELTQNDLKAL 268
KI + M + D +I + D K +D LD+L++L+D LT +LKA
Sbjct: 818 KIMKAMRIMRKYHDPIIDDRIKQWNDGLKTVEEDLLDVLIKLKDANNKPL-LTLKELKAQ 994
Query: 269 LMNMFLGGSDTTSTTLEWAMA 289
++ + + D S EWA+A
Sbjct: 995 IIELAIEMVDNPSNAFEWALA 1057
>TC14395 weakly similar to UP|C823_SOYBN (O49858) Cytochrome P450 82A3
(P450 CP6) , partial (38%)
Length = 1079
Score = 32.7 bits (73), Expect = 0.081
Identities = 15/46 (32%), Positives = 28/46 (60%), Gaps = 7/46 (15%)
Frame = +3
Query: 267 ALLMNMFLGGSDTTSTTLEWAMA-------ACKKSSHNEESTGGGK 305
A + + + G+DTT++TL WA++ A KK++H ++ GG+
Sbjct: 3 ATCLQLIVAGTDTTTSTLTWALSLLLNNREALKKATHELDTQMGGR 140
>BP029912
Length = 559
Score = 30.8 bits (68), Expect = 0.31
Identities = 18/53 (33%), Positives = 31/53 (57%), Gaps = 3/53 (5%)
Frame = -3
Query: 221 DDFLDRVI---VEHKMSRRDPKKKDFLDILLQLQDDGLTEFELTQNDLKALLM 270
D D+VI V+ + ++ + DFL LL L+++G ++ L+ +KALLM
Sbjct: 278 DRIFDKVIGERVKMEAKGKENESXDFLQFLLDLKEEGDSKVPLSITHVKALLM 120
>TC11981
Length = 510
Score = 26.6 bits (57), Expect = 5.8
Identities = 8/23 (34%), Positives = 16/23 (68%)
Frame = -3
Query: 46 LVSFLCWWFPQYIWPRKSCKHMA 68
++ +C+ FPQY+ P ++ H+A
Sbjct: 133 MIILICYPFPQYVTPTQTYSHIA 65
>TC14744 UP|C7DB_LOTJA (O22307) Cytochrome P450 71D11 (Fragment) , complete
Length = 1753
Score = 26.6 bits (57), Expect = 5.8
Identities = 20/48 (41%), Positives = 22/48 (45%)
Frame = +1
Query: 54 FPQYIWPRKSCKHMA*FLRTDQARH*QKPYSMVVRTLRFHLMAIHGDR 101
FPQ RKS K M L Q MV +T FHLM I GD+
Sbjct: 271 FPQQSMLRKS*KPMMSHLHPGLVLFSQI*CFMVPQT*AFHLMVITGDK 414
>BP054586
Length = 340
Score = 26.6 bits (57), Expect = 5.8
Identities = 17/52 (32%), Positives = 26/52 (49%)
Frame = +1
Query: 223 FLDRVIVEHKMSRRDPKKKDFLDILLQLQDDGLTEFELTQNDLKALLMNMFL 274
FL + ++HK SRR PK + + L+ L +L N L A++ FL
Sbjct: 79 FLSSLFLDHKHSRRLPKLS--ITQIFALERKALVHMQLESNLLLAVMNLRFL 228
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.353 0.156 0.562
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 5,905,518
Number of Sequences: 28460
Number of extensions: 81956
Number of successful extensions: 1166
Number of sequences better than 10.0: 46
Number of HSP's better than 10.0 without gapping: 1153
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 1164
length of query: 313
length of database: 4,897,600
effective HSP length: 90
effective length of query: 223
effective length of database: 2,336,200
effective search space: 520972600
effective search space used: 520972600
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 14 ( 7.1 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 37 (21.5 bits)
S2: 55 (25.8 bits)
Medicago: description of AC124952.12


