

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC124951.10 - phase: 0
(446 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
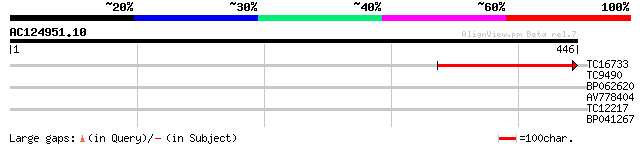
TC16733 weakly similar to UP|Q8W106 (Q8W106) AT3g27930/K24A2_2, ... 199 6e-52
TC9490 homologue to UP|Q8W1R5 (Q8W1R5) 26S proteasome regulatory... 29 1.4
BP062620 28 4.0
AV778404 27 6.8
TC12217 27 6.8
BP041267 27 8.9
>TC16733 weakly similar to UP|Q8W106 (Q8W106) AT3g27930/K24A2_2, partial
(26%)
Length = 629
Score = 199 bits (507), Expect = 6e-52
Identities = 93/110 (84%), Positives = 103/110 (93%)
Frame = +3
Query: 337 LVKAKAGPKSSTMALAFKSWWKPSFTFSISATRDRADGQVQYGFGLQSESLREASYQRAD 396
L+K K GP+ S+MALAFKSWWKPSFTFSISATRDRADGQ+QYGFG+ SESLREASYQRAD
Sbjct: 3 LLKXKVGPRISSMALAFKSWWKPSFTFSISATRDRADGQIQYGFGIXSESLREASYQRAD 182
Query: 397 PNFVMLTPSKEHLAEGIVWETGKRPMLQSDIDAGHFDGLPRELRPLDKIL 446
PN+VMLTPSKEHLAEGI W TG+RPMLQSD+DAG F+GLPR+LRPLDKIL
Sbjct: 183 PNYVMLTPSKEHLAEGIAWTTGQRPMLQSDLDAGQFEGLPRDLRPLDKIL 332
>TC9490 homologue to UP|Q8W1R5 (Q8W1R5) 26S proteasome regulatory subunit
IV, partial (20%)
Length = 552
Score = 29.3 bits (64), Expect = 1.4
Identities = 17/54 (31%), Positives = 25/54 (45%)
Frame = +1
Query: 133 SCAYYPRYGFGAFGVFPLLLKNRTSSQIATFTADKVRQLYRETSQDSGVMGLRY 186
S YYPR F FG++ L+++ S + + + LY SQ G L Y
Sbjct: 280 SILYYPRLSFFFFGIYVLVMQTNAHSFVLS-KLSFIHLLYSG*SQSWGQFNLHY 438
>BP062620
Length = 555
Score = 27.7 bits (60), Expect = 4.0
Identities = 12/30 (40%), Positives = 15/30 (50%), Gaps = 8/30 (26%)
Frame = +2
Query: 2 LIWKWIVSWYRNEVE--------SPPPVVL 23
LIW W V W +N+ + SPPP L
Sbjct: 236 LIWNWFVEWMKNKPDLKGGNPDRSPPPADL 325
>AV778404
Length = 401
Score = 26.9 bits (58), Expect = 6.8
Identities = 11/20 (55%), Positives = 12/20 (60%)
Frame = +1
Query: 13 NEVESPPPVVLVPPLFDFPP 32
NE SPPP+ L PPL P
Sbjct: 340 NEPTSPPPLTLSPPLSQLTP 399
>TC12217
Length = 495
Score = 26.9 bits (58), Expect = 6.8
Identities = 10/25 (40%), Positives = 16/25 (64%)
Frame = -3
Query: 90 PLDQKPDENINGNALFRWQSDVNDP 114
PL+ P EN+ +A++ WQ V+ P
Sbjct: 289 PLNLNPQENLTTDAIYCWQFCVSSP 215
>BP041267
Length = 520
Score = 26.6 bits (57), Expect = 8.9
Identities = 12/35 (34%), Positives = 18/35 (51%)
Frame = +2
Query: 288 LEENTVVGITNYIDFGFELQTSVDDAIAANNISDS 322
LEEN V + ++ G T VD + N +SD+
Sbjct: 242 LEENNVAVVEKNVEAGSSTTTEVDKSGCVNTVSDA 346
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.320 0.135 0.413
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 8,037,919
Number of Sequences: 28460
Number of extensions: 108544
Number of successful extensions: 580
Number of sequences better than 10.0: 12
Number of HSP's better than 10.0 without gapping: 559
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 580
length of query: 446
length of database: 4,897,600
effective HSP length: 93
effective length of query: 353
effective length of database: 2,250,820
effective search space: 794539460
effective search space used: 794539460
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 57 (26.6 bits)
Medicago: description of AC124951.10


