

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC124217.4 + phase: 0
(583 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
Score E
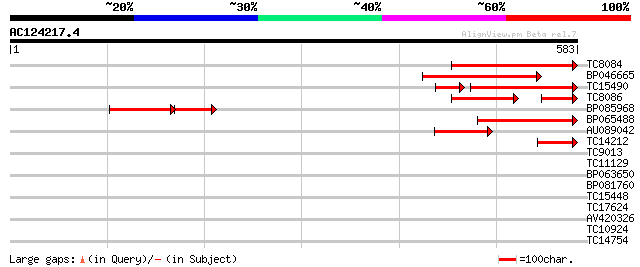
Sequences producing significant alignments: (bits) Value
TC8084 similar to UP|PIF1_YEAST (P07271) DNA repair and recombin... 169 9e-43
BP046665 152 1e-37
TC15490 weakly similar to UP|Q9LTU4 (Q9LTU4) Helicase-like prote... 128 2e-36
TC8086 similar to UP|DMC1_HUMAN (Q14565) Meiotic recombination p... 93 2e-27
BP085968 80 1e-24
BP065488 90 1e-18
AU089042 65 2e-11
TC14212 similar to UP|Q9FN61 (Q9FN61) Gb|AAF07369.1, partial (10%) 47 1e-05
TC9013 35 0.034
TC11129 weakly similar to UP|PDA6_MEDSA (P38661) Probable protei... 35 0.044
BP063650 32 0.22
BP081760 28 3.2
TC15448 weakly similar to UP|Q9FIH0 (Q9FIH0) Brain and reproduct... 28 4.1
TC17624 28 4.1
AV420326 28 5.4
TC10924 homologue to UP|Q8H9D4 (Q8H9D4) 26S proteasome AAA-ATPas... 27 7.0
TC14754 similar to UP|Q9ZTX0 (Q9ZTX0) Similarity to SCAMP37, par... 27 9.2
>TC8084 similar to UP|PIF1_YEAST (P07271) DNA repair and recombination
protein PIF1, mitochondrial precursor, partial (4%)
Length = 560
Score = 169 bits (429), Expect = 9e-43
Identities = 80/129 (62%), Positives = 102/129 (79%)
Frame = +1
Query: 455 LCNGTRLIITRMGRYVLEGKVISGSNVGDRVFIPRLSLSPSDVRIPFKFQRKQFPLAVSF 514
LCNGTRLI+ +G YV++ VI+G+N+GD +FIPRL + PSD PFKF+R+ FP+++ F
Sbjct: 61 LCNGTRLIVVDLGTYVIKATVITGTNIGDDIFIPRLDMVPSDSGYPFKFERR*FPISLCF 240
Query: 515 AMTINKSQGQSLQNVGVYLPAPVFSHGQLYVAVSRVTSRGGLKILITDEDGDDTNLTSNV 574
AMTINKSQGQSL +V +YL PVF+HGQLYVA+SRV SR GLK+L+ DE+ TN T NV
Sbjct: 241 AMTINKSQGQSLSHVSLYLSRPVFTHGQLYVALSRVRSRKGLKLLVLDEEEKVTNTTKNV 420
Query: 575 VYEEVFRNV 583
VY EVF N+
Sbjct: 421 VYREVFENI 447
>BP046665
Length = 524
Score = 152 bits (384), Expect = 1e-37
Identities = 71/123 (57%), Positives = 94/123 (75%)
Frame = -1
Query: 425 IVASGLPNHKLRLKEGVPVMLLRNLDTKNGLCNGTRLIITRMGRYVLEGKVISGSNVGDR 484
I G+PNHK+ LKEG P+MLLRN+ G CNGTRLI+ +G V++ VI+ +N+GD
Sbjct: 521 ITCFGIPNHKITLKEGAPIMLLRNIYQAVGFCNGTRLIVADLGTNVIKATVIT*TNIGDD 342
Query: 485 VFIPRLSLSPSDVRIPFKFQRKQFPLAVSFAMTINKSQGQSLQNVGVYLPAPVFSHGQLY 544
+FIPR+ + PSD PFKF+R+QFP+++ AMTINKSQGQSL +VG+YL VF+HGQLY
Sbjct: 341 IFIPRMDMVPSDSGYPFKFERRQFPISLCSAMTINKSQGQSLSHVGLYLSRHVFTHGQLY 162
Query: 545 VAV 547
VA+
Sbjct: 161 VAL 153
Score = 37.7 bits (86), Expect = 0.005
Identities = 19/45 (42%), Positives = 27/45 (59%)
Frame = -2
Query: 532 YLPAPVFSHGQLYVAVSRVTSRGGLKILITDEDGDDTNLTSNVVY 576
Y+ ++SH +S + SR GLK+L+ DE+ TN T NVVY
Sbjct: 202 YIFLAMYSHMVSCTLLSMLRSRKGLKLLVLDEEEKVTNTTKNVVY 68
>TC15490 weakly similar to UP|Q9LTU4 (Q9LTU4) Helicase-like protein, partial
(8%)
Length = 634
Score = 128 bits (321), Expect(2) = 2e-36
Identities = 61/109 (55%), Positives = 83/109 (75%)
Frame = +2
Query: 475 VISGSNVGDRVFIPRLSLSPSDVRIPFKFQRKQFPLAVSFAMTINKSQGQSLQNVGVYLP 534
VI+G+++GD + IPR+ + PSD PFKF+R+Q P+++ FAMTINKSQG+SL +VG+YL
Sbjct: 149 VITGTHIGDDISIPRMDMVPSDSSYPFKFERRQSPISLCFAMTINKSQGRSLSHVGLYLS 328
Query: 535 APVFSHGQLYVAVSRVTSRGGLKILITDEDGDDTNLTSNVVYEEVFRNV 583
PV +HG LYVA+ RV SR LK+L+ DE+ TN T NVVY E+F N+
Sbjct: 329 RPVSTHG*LYVALPRVRSRK*LKLLVLDEEEKMTNTTKNVVYREIFENI 475
Score = 41.6 bits (96), Expect(2) = 2e-36
Identities = 18/30 (60%), Positives = 23/30 (76%)
Frame = +1
Query: 438 KEGVPVMLLRNLDTKNGLCNGTRLIITRMG 467
KEG +MLLRN+ +GLCNGTRLI+ +G
Sbjct: 43 KEGALIMLLRNIVQASGLCNGTRLIVVDLG 132
>TC8086 similar to UP|DMC1_HUMAN (Q14565) Meiotic recombination protein
DMC1/LIM15 homolog, partial (12%)
Length = 628
Score = 93.2 bits (230), Expect(2) = 2e-27
Identities = 41/69 (59%), Positives = 56/69 (80%)
Frame = +1
Query: 455 LCNGTRLIITRMGRYVLEGKVISGSNVGDRVFIPRLSLSPSDVRIPFKFQRKQFPLAVSF 514
LC+GTRLI+ +G YV++ VI+G+N+GD +FIPRL + PSD PFKF+R+ FP+++ F
Sbjct: 169 LCHGTRLIVVDLGTYVIKATVITGTNIGDDIFIPRLDMVPSDSGYPFKFERR*FPISLCF 348
Query: 515 AMTINKSQG 523
AMTINKSQG
Sbjct: 349 AMTINKSQG 375
Score = 46.6 bits (109), Expect(2) = 2e-27
Identities = 22/36 (61%), Positives = 26/36 (72%)
Frame = +3
Query: 548 SRVTSRGGLKILITDEDGDDTNLTSNVVYEEVFRNV 583
SRV SR GLK+L+ DE+ TN T NVVY EVF N+
Sbjct: 369 SRVRSRKGLKLLVLDEEEKVTNTTKNVVYREVFENI 476
>BP085968
Length = 341
Score = 79.7 bits (195), Expect(2) = 1e-24
Identities = 37/68 (54%), Positives = 50/68 (73%)
Frame = +2
Query: 103 TDEQHKVYDTIMARVDEKKPGIFFLYGYGGTGKTFIWKAMSAAIRSRGEIVLTVASSGIA 162
T +Q KVY IM+ V G +FLYG+GG+GKTF+W S+ +RS+G +VL VASSGIA
Sbjct: 11 TAKQSKVYKQIMSSVWSDDGGFYFLYGFGGSGKTFVWNTWSSGLRSQGLMVLNVASSGIA 190
Query: 163 ALLIPGGR 170
+ L+PGG+
Sbjct: 191 SWLLPGGK 214
Score = 50.8 bits (120), Expect(2) = 1e-24
Identities = 22/43 (51%), Positives = 31/43 (71%)
Frame = +3
Query: 170 RTAHSRFGIPFTVDECSTCGVKPNTPLAQLVVQARLIIWDEAP 212
RTAHSRF I ++++ STC +K + A+L+ +A IIWDEAP
Sbjct: 213 RTAHSRFSIHISINDISTCNIKQGSQKAELLQKASSIIWDEAP 341
>BP065488
Length = 439
Score = 89.7 bits (221), Expect = 1e-18
Identities = 50/104 (48%), Positives = 68/104 (65%), Gaps = 2/104 (1%)
Frame = -1
Query: 482 GDRVFIPRLSLSPSDVRIPFKFQRKQFPLAVSFAMTINKSQGQ-SLQNVGVYLPAPVFSH 540
GD +FIPR+++ PS P KF+R QFP+++ FAMTINKSQ V + +SH
Sbjct: 379 GDDIFIPRMNMVPSVSGDPLKFERCQFPISLCFAMTINKSQXSVHYPTVALISFLACYSH 200
Query: 541 -GQLYVAVSRVTSRGGLKILITDEDGDDTNLTSNVVYEEVFRNV 583
+LYVA+S V SR GLK+L+ E+ TN T N VY+EVF+N+
Sbjct: 199 MDKLYVALSGVRSRKGLKLLVLGEEEKVTNATKNEVYQEVFKNI 68
>AU089042
Length = 191
Score = 65.5 bits (158), Expect = 2e-11
Identities = 30/60 (50%), Positives = 42/60 (70%)
Frame = +3
Query: 437 LKEGVPVMLLRNLDTKNGLCNGTRLIITRMGRYVLEGKVISGSNVGDRVFIPRLSLSPSD 496
LK GVPVML+ NL GLCNGTRLI+ +G V+ ++SG+++G V+I ++L PSD
Sbjct: 9 LKXGVPVMLMXNLXISTGLCNGTRLIVDYLGPNVIGATILSGTHIGXVVYISMMNLXPSD 188
>TC14212 similar to UP|Q9FN61 (Q9FN61) Gb|AAF07369.1, partial (10%)
Length = 613
Score = 46.6 bits (109), Expect = 1e-05
Identities = 22/41 (53%), Positives = 29/41 (70%)
Frame = +2
Query: 543 LYVAVSRVTSRGGLKILITDEDGDDTNLTSNVVYEEVFRNV 583
LYVAVSRV S+ GLKILI+ + T N+VY+EVF+ +
Sbjct: 2 LYVAVSRVKSKDGLKILISSDGTSTPGATKNIVYKEVFQKI 124
>TC9013
Length = 766
Score = 35.0 bits (79), Expect = 0.034
Identities = 17/26 (65%), Positives = 19/26 (72%)
Frame = +3
Query: 558 ILITDEDGDDTNLTSNVVYEEVFRNV 583
ILI D D ++N TSNVVY EVFR V
Sbjct: 627 ILICDGDDSNSNSTSNVVYTEVFRTV 704
>TC11129 weakly similar to UP|PDA6_MEDSA (P38661) Probable protein disulfide
isomerase A6 precursor (P5) , partial (7%)
Length = 600
Score = 34.7 bits (78), Expect = 0.044
Identities = 14/24 (58%), Positives = 19/24 (78%)
Frame = +2
Query: 379 VEKINEYMLDMVPGEEKVYLSYDS 402
VEK+NE+MLD++PG YLS D+
Sbjct: 473 VEKVNEFMLDLLPGNTTEYLSSDT 544
>BP063650
Length = 497
Score = 32.3 bits (72), Expect = 0.22
Identities = 13/31 (41%), Positives = 22/31 (70%)
Frame = +1
Query: 255 QILPVIPRGIRPEIVHATINSSHLWRHCEVL 285
+ L V+ +G +I+HA +N+S+L HC+VL
Sbjct: 58 KFLTVVYKGRMQDIIHAIVNASYL*DHCQVL 150
>BP081760
Length = 355
Score = 28.5 bits (62), Expect = 3.2
Identities = 12/30 (40%), Positives = 20/30 (66%)
Frame = -2
Query: 539 SHGQLYVAVSRVTSRGGLKILITDEDGDDT 568
SHG ++ + S + L++L +DEDG+DT
Sbjct: 282 SHGAVWTSSSPPSGPLSLRMLFSDEDGEDT 193
>TC15448 weakly similar to UP|Q9FIH0 (Q9FIH0) Brain and reproductive
organ-expressed protein-like, partial (32%)
Length = 631
Score = 28.1 bits (61), Expect = 4.1
Identities = 17/47 (36%), Positives = 27/47 (57%), Gaps = 5/47 (10%)
Frame = +2
Query: 58 LSDYPPMPTADPSLMPDLQNHFIHEE-INYNRR----ELQEEHNQLM 99
+SDYP P +PSLM + F+ +E +N+ R+ +L + H Q M
Sbjct: 227 ISDYPWSPRWEPSLMAERIGEFLADESLNFKRQCSEGQL*Q*HGQSM 367
>TC17624
Length = 615
Score = 28.1 bits (61), Expect = 4.1
Identities = 12/13 (92%), Positives = 13/13 (99%)
Frame = +1
Query: 571 TSNVVYEEVFRNV 583
TSNVVY+EVFRNV
Sbjct: 535 TSNVVYKEVFRNV 573
>AV420326
Length = 416
Score = 27.7 bits (60), Expect = 5.4
Identities = 12/31 (38%), Positives = 18/31 (57%)
Frame = +2
Query: 423 NTIVASGLPNHKLRLKEGVPVMLLRNLDTKN 453
N+ AS NH+LRL+ V V + ++ T N
Sbjct: 32 NSFCASSADNHRLRLRRAVTVSVSASVSTSN 124
>TC10924 homologue to UP|Q8H9D4 (Q8H9D4) 26S proteasome AAA-ATPase subunit
RPT4a, partial (49%)
Length = 648
Score = 27.3 bits (59), Expect = 7.0
Identities = 14/34 (41%), Positives = 20/34 (58%)
Frame = +2
Query: 113 IMARVDEKKPGIFFLYGYGGTGKTFIWKAMSAAI 146
+ RV K P LYG GTGKT + +A+++ I
Sbjct: 533 LFLRVGIKPPKGVLLYGPPGTGKTLLARAIASNI 634
>TC14754 similar to UP|Q9ZTX0 (Q9ZTX0) Similarity to SCAMP37, partial (38%)
Length = 651
Score = 26.9 bits (58), Expect = 9.2
Identities = 10/21 (47%), Positives = 15/21 (70%)
Frame = +3
Query: 8 VSFLHQCYTLPLTYVNKCYVF 28
V++ HQ L + +VNKCY+F
Sbjct: 372 VNYSHQDILLRVNFVNKCYLF 434
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.322 0.139 0.413
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 9,630,298
Number of Sequences: 28460
Number of extensions: 127079
Number of successful extensions: 599
Number of sequences better than 10.0: 35
Number of HSP's better than 10.0 without gapping: 595
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 598
length of query: 583
length of database: 4,897,600
effective HSP length: 95
effective length of query: 488
effective length of database: 2,193,900
effective search space: 1070623200
effective search space used: 1070623200
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.9 bits)
S2: 58 (26.9 bits)
Medicago: description of AC124217.4


