

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
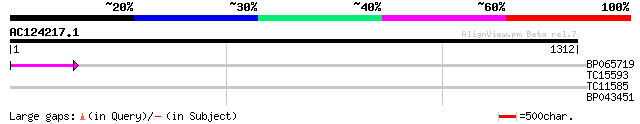
Query= AC124217.1 - phase: 0 /pseudo
(1312 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
BP065719 49 5e-06
TC15593 weakly similar to UP|AAR15453 (AAR15453) Nuclear divisio... 29 4.2
TC11585 similar to UP|Q9ZWQ5 (Q9ZWQ5) UDP-glycose:flavonoid glyc... 29 5.5
BP043451 28 9.4
>BP065719
Length = 567
Score = 48.9 bits (115), Expect = 5e-06
Identities = 51/159 (32%), Positives = 74/159 (46%), Gaps = 1/159 (0%)
Frame = +2
Query: 1 WFQNGSIQ-A*ICIGLI*ICVNGRIA*GVENP*INQIWWRNY*IHCRTHCLLFDRGR*YR 59
WFQ G + C+ L R + +E P I+ + +H RTH + +R R
Sbjct: 89 WFQRGFRE*TPFCVSLYRGGSRIRTSSRLEGPQIH*VLRGFRRVHGRTHSQVPNRSRRLS 268
Query: 60 KQREFKNEVFFKFFN*KCVHLVYNITPMIYLYLESIRKGIS*TVLHGPI*D*S*RVGQCS 119
Q +FKNEV +FFN KC++LV++ + Y+ + K *TVL G + G C
Sbjct: 269 HQ*KFKNEVLPEFFNQKCLYLVHHPRS*VSPYMGPVGKNFP*TVL*GRVQSKXEGPG*CQ 448
Query: 120 T*NV*IY**LFE*I*TP*G*MLYHAT*T*IGRNGCGWSR 158
+Y**L + +* * ML+ *T WSR
Sbjct: 449 KKASRVY**LPKQV*DA*ISMLHPCV*TRASCLXRWWSR 565
>TC15593 weakly similar to UP|AAR15453 (AAR15453) Nuclear division RFT1-like
protein, partial (17%)
Length = 753
Score = 29.3 bits (64), Expect = 4.2
Identities = 14/46 (30%), Positives = 23/46 (49%)
Frame = -2
Query: 505 TPSSKSPTEKWVFSSGKKSSYTSPPAKWVKRFTGTDNQNKAPVSKK 550
T SPTEKW+ + G+K S++ ++ T N+ P K+
Sbjct: 185 TQKQASPTEKWIINDGQKLSWSRNTFSEMRVITPESNRIIHPEGKQ 48
>TC11585 similar to UP|Q9ZWQ5 (Q9ZWQ5) UDP-glycose:flavonoid
glycosyltransferase, partial (40%)
Length = 609
Score = 28.9 bits (63), Expect = 5.5
Identities = 19/49 (38%), Positives = 24/49 (48%), Gaps = 4/49 (8%)
Frame = +2
Query: 493 TRHQNRGQVRTFTPSSKSPTEKWVFSSGKKS----SYTSPPAKWVKRFT 537
T H + V T SS SP KWV G ++ TSPPA++ R T
Sbjct: 176 TEHADTTSVFT---SSNSPAPKWVSLVGSRTCSPHPTTSPPARYTWRLT 313
>BP043451
Length = 544
Score = 28.1 bits (61), Expect = 9.4
Identities = 13/31 (41%), Positives = 21/31 (66%), Gaps = 3/31 (9%)
Frame = +1
Query: 597 FKMIRKPATERIFPPLSA---VKENLHVEDE 624
F M+ +++I+ PLSA +KENLH+E +
Sbjct: 343 FPMVFSSCSQQIYSPLSA*ALLKENLHLESD 435
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.358 0.160 0.568
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 21,207,288
Number of Sequences: 28460
Number of extensions: 284687
Number of successful extensions: 3082
Number of sequences better than 10.0: 9
Number of HSP's better than 10.0 without gapping: 2274
Number of HSP's successfully gapped in prelim test: 76
Number of HSP's that attempted gapping in prelim test: 763
Number of HSP's gapped (non-prelim): 2425
length of query: 1312
length of database: 4,897,600
effective HSP length: 101
effective length of query: 1211
effective length of database: 2,023,140
effective search space: 2450022540
effective search space used: 2450022540
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 14 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 37 (21.7 bits)
S2: 61 (28.1 bits)
Medicago: description of AC124217.1


