

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC124216.5 - phase: 0
(340 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
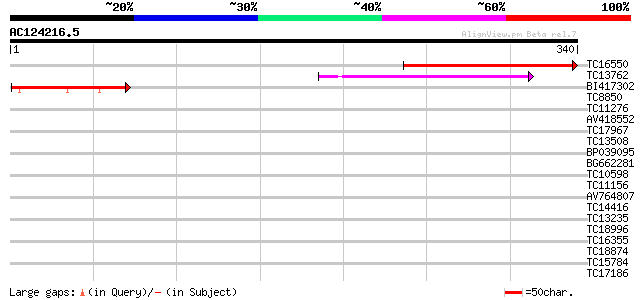
Score E
Sequences producing significant alignments: (bits) Value
TC16550 similar to UP|AAR25824 (AAR25824) Basic pentacysteine 4,... 187 2e-48
TC13762 similar to UP|Q9LDE2 (Q9LDE2) F10B6.5 (T5E21.17) (At1g14... 115 8e-27
BI417302 65 2e-11
TC8850 similar to UP|RK34_SPIOL (P82244) 50S ribosomal protein L... 29 1.3
TC11276 weakly similar to UP|HDA1_CHICK (P56517) Histone deacety... 29 1.3
AV418552 28 1.7
TC17967 similar to UP|Q7XA66 (Q7XA66) At1g49590, partial (17%) 28 1.7
TC13508 28 2.2
BP039095 28 2.2
BG662281 28 2.2
TC10598 weakly similar to GB|AAM78036.1|21928015|AY125526 At2g01... 28 2.2
TC11156 28 2.9
AV764807 27 3.8
TC14416 similar to UP|Q8SMH8 (Q8SMH8) RNA-binding protein, parti... 27 4.9
TC13235 similar to UP|Q9LQB6 (Q9LQB6) F4N2.1, partial (38%) 27 4.9
TC18996 similar to UP|O24323 (O24323) Cysteine proteinase precur... 27 6.4
TC16355 similar to UP|Q9M4U6 (Q9M4U6) Histone H1 variant, partia... 27 6.4
TC18874 similar to UP|Q9FJJ0 (Q9FJJ0) Genomic DNA, chromosome 5,... 27 6.4
TC15784 similar to UP|PE31_ARATH (Q9LHA7) Peroxidase 31 precurso... 27 6.4
TC17186 homologue to UP|AAD03372 (AAD03372) Expressed protein, p... 27 6.4
>TC16550 similar to UP|AAR25824 (AAR25824) Basic pentacysteine 4, partial
(35%)
Length = 641
Score = 187 bits (475), Expect = 2e-48
Identities = 78/104 (75%), Positives = 95/104 (91%)
Frame = +2
Query: 237 LNQVAYDDSTMPAPACSCTGVLRQCYKWGNGGWQSACCTTTLSVYPLPAVPNKRHARVGG 296
LN +A+D++ MP P C+CTG+ RQCYKWGNGGWQS+CCTTTLSVYPLP +PNKRHAR+GG
Sbjct: 5 LNLIAFDETIMPVPVCTCTGIARQCYKWGNGGWQSSCCTTTLSVYPLPQLPNKRHARIGG 184
Query: 297 RKMSGSAFNKLLSRLAAEGHDLSHPVDLKDHWAKHGTNRYITIK 340
RKMSGS F +LLS+LA+EGHD+S P+DLK++WA+HGTNRYITIK
Sbjct: 185 RKMSGSVFTRLLSKLASEGHDISIPLDLKNYWARHGTNRYITIK 316
>TC13762 similar to UP|Q9LDE2 (Q9LDE2) F10B6.5 (T5E21.17)
(At1g14680/F10B6_22), partial (42%)
Length = 568
Score = 115 bits (289), Expect = 8e-27
Identities = 59/129 (45%), Positives = 77/129 (58%)
Frame = +3
Query: 186 VKKEGDDLNKTMFANNKELEWKSSEEIINGDDDLNKQLAISKADWKPQDLALNQVAYDDS 245
VKKEG K + L +++ D+ N + K KP +L +N + D S
Sbjct: 186 VKKEGGQSKKRQ--SKGALTTPKAKKPRKPKDNSNVAVQRVKPPKKPMELVINGIDMDIS 359
Query: 246 TMPAPACSCTGVLRQCYKWGNGGWQSACCTTTLSVYPLPAVPNKRHARVGGRKMSGSAFN 305
+P P CSCTG +QCY+WG GGWQSACCTT +S+YPLP +R AR+ GRKMS AF
Sbjct: 360 GLPVPVCSCTGSPQQCYRWGCGGWQSACCTTNVSIYPLPMSIKRRGARIAGRKMSQGAFK 539
Query: 306 KLLSRLAAE 314
K+L +LAAE
Sbjct: 540 KVLEKLAAE 566
>BI417302
Length = 499
Score = 65.1 bits (157), Expect = 2e-11
Identities = 41/80 (51%), Positives = 51/80 (63%), Gaps = 9/80 (11%)
Frame = +2
Query: 2 DDR--ENGRHKADQYKSAQGQWLM-QQHQHQHPSM----KQIMSIMAERDAAIQERNL-- 52
DDR ENGRHK + Y+ A+ W M QHQ + P+ K++ SIMAER AA+ E L
Sbjct: 257 DDRQHENGRHKMEYYRGARSPWDMDHQHQVKEPNALVMNKKLRSIMAERQAALLELELEA 436
Query: 53 ALSEKKAALAERDMAFLQRD 72
A+SEK ALA RD+A QRD
Sbjct: 437 AISEKNEALAARDLALRQRD 496
>TC8850 similar to UP|RK34_SPIOL (P82244) 50S ribosomal protein L34,
chloroplast precursor, partial (49%)
Length = 659
Score = 28.9 bits (63), Expect = 1.3
Identities = 18/66 (27%), Positives = 35/66 (52%), Gaps = 6/66 (9%)
Frame = -3
Query: 162 PRRAKRP------KESKSDSPNKKTPKSRKVKKEGDDLNKTMFANNKELEWKSSEEIING 215
PRR P K + SD+P + + K ++EG++ K +N + + W+ SEE++
Sbjct: 243 PRRFPFPVPALDIKSNPSDNPESE--EEGKEREEGEE-RKEQRSNGERVLWRDSEEVVGI 73
Query: 216 DDDLNK 221
+++ K
Sbjct: 72 EEEAVK 55
>TC11276 weakly similar to UP|HDA1_CHICK (P56517) Histone deacetylase 1
(HD1), partial (5%)
Length = 636
Score = 28.9 bits (63), Expect = 1.3
Identities = 18/64 (28%), Positives = 26/64 (40%), Gaps = 7/64 (10%)
Frame = +3
Query: 154 VSLEVGKPPRRAKRPK-------ESKSDSPNKKTPKSRKVKKEGDDLNKTMFANNKELEW 206
VS+ K P + + PK E K D P + P+ + KKE + KE E
Sbjct: 3 VSVGPAKEPEKKEEPKKEEPQKEEEKKDEPKSEEPQKEEAKKEEPKKEEEKKDEKKEEEK 182
Query: 207 KSSE 210
K +
Sbjct: 183 KKEQ 194
Score = 26.6 bits (57), Expect = 6.4
Identities = 11/45 (24%), Positives = 24/45 (52%)
Frame = +1
Query: 160 KPPRRAKRPKESKSDSPNKKTPKSRKVKKEGDDLNKTMFANNKEL 204
K PR+ +R ++S +K + ++ K++ + L T+F + L
Sbjct: 100 KNPRKKRRRRKSPRKKKRRKMKRKKRRKRKSNQLKLTLF*SGSRL 234
>AV418552
Length = 386
Score = 28.5 bits (62), Expect = 1.7
Identities = 18/82 (21%), Positives = 35/82 (41%)
Frame = +2
Query: 16 SAQGQWLMQQHQHQHPSMKQIMSIMAERDAAIQERNLALSEKKAALAERDMAFLQRDTAI 75
S+Q Q LMQQHQ + ++ Q + L++ + L++ + LQ+
Sbjct: 47 SSQEQQLMQQHQQMLSYFQPQQQLLPHLQLQQQLQQQQLAQLQQQLSQLQLTLLQQQQLT 226
Query: 76 AERNNALMERDNAIATLQFREN 97
+ L + +A LQ ++
Sbjct: 227 QLQQQQLAQLQQQLAQLQLTQH 292
>TC17967 similar to UP|Q7XA66 (Q7XA66) At1g49590, partial (17%)
Length = 662
Score = 28.5 bits (62), Expect = 1.7
Identities = 18/54 (33%), Positives = 28/54 (51%), Gaps = 4/54 (7%)
Frame = +2
Query: 140 TRELHTTDALPAAPVSLEVGKPPRRAKRPK----ESKSDSPNKKTPKSRKVKKE 189
T L+ T + AAP SL +GK R ++PK E KS ++ + R ++E
Sbjct: 149 TTSLNPTRNVKAAPSSLSLGKRKRPGEKPKVVSEEEKSALKAREAARKRVQERE 310
>TC13508
Length = 522
Score = 28.1 bits (61), Expect = 2.2
Identities = 21/70 (30%), Positives = 29/70 (41%), Gaps = 2/70 (2%)
Frame = -1
Query: 121 HHLPQQVNHLPNMGDSSYGTRELHTTDA--LPAAPVSLEVGKPPRRAKRPKESKSDSPNK 178
H LPQ + P + +TD+ L + P S PPR + + +S P K
Sbjct: 240 HPLPQPTRNSPCVQVPESSPPPTSSTDSKQLQSPPFS-SPPSPPRTSSSSRSCQSPHPTK 64
Query: 179 KTPKSRKVKK 188
KT K KK
Sbjct: 63 KTLKKTTAKK 34
>BP039095
Length = 472
Score = 28.1 bits (61), Expect = 2.2
Identities = 12/40 (30%), Positives = 21/40 (52%)
Frame = -3
Query: 1 MDDRENGRHKADQYKSAQGQWLMQQHQHQHPSMKQIMSIM 40
+DD E+G + + G W +Q H PSMK ++ ++
Sbjct: 362 VDDDEDGDFGIAKKLAIVGLWCIQWHPMHRPSMKTVLQML 243
>BG662281
Length = 399
Score = 28.1 bits (61), Expect = 2.2
Identities = 31/119 (26%), Positives = 46/119 (38%), Gaps = 2/119 (1%)
Frame = +2
Query: 94 FRENALANGGMSSCPPGCQISRGVKHIHHLPQQVNHLPNMGDSSYGTRELHTTDALPAAP 153
FRE A A P Q R + +LP + + PN + L +T A AAP
Sbjct: 62 FREPARATFQKQRTPVHIQPRRAL----NLPTKTSRKPNQRTTQQWILLLPSTAARNAAP 229
Query: 154 VSLEVGKPPRRAKRPKESKSDSPNKKTPKSRKVKKEGDDLNKTMFA--NNKELEWKSSE 210
S + PP + R K ++ N +P + +T F N +L S+E
Sbjct: 230 TS--ISTPPTPSHRTSTLKPET-NTPSPSPPSIPPSSPSRKRTKFVLLRNPQLLGDSAE 397
>TC10598 weakly similar to GB|AAM78036.1|21928015|AY125526
At2g01100/F23H14.7 {Arabidopsis thaliana;}, partial
(33%)
Length = 902
Score = 28.1 bits (61), Expect = 2.2
Identities = 25/100 (25%), Positives = 42/100 (42%), Gaps = 9/100 (9%)
Frame = +3
Query: 156 LEVGKPPRRAKRPKESK--SDSPNKKTPKS--RKVKKEGD-----DLNKTMFANNKELEW 206
L+ K +RA ES SD +K+T + RK KK + K + K +W
Sbjct: 372 LKAAKHKKRADSDSESDYDSDDESKRTSRRSHRKHKKRSHYDSDHEKRKEKLSKRKTKKW 551
Query: 207 KSSEEIINGDDDLNKQLAISKADWKPQDLALNQVAYDDST 246
S +GD+ ++ + K + +QV+ DS+
Sbjct: 552 SSESSDFSGDESESRSEEERRRKRKQKKKLRDQVSRSDSS 671
>TC11156
Length = 553
Score = 27.7 bits (60), Expect = 2.9
Identities = 11/22 (50%), Positives = 14/22 (63%)
Frame = +1
Query: 113 ISRGVKHIHHLPQQVNHLPNMG 134
I+R + H+HHLP N L N G
Sbjct: 79 IARNMPHLHHLPLMGNWLTNYG 144
>AV764807
Length = 529
Score = 27.3 bits (59), Expect = 3.8
Identities = 16/59 (27%), Positives = 21/59 (35%)
Frame = +3
Query: 111 CQISRGVKHIHHLPQQVNHLPNMGDSSYGTRELHTTDALPAAPVSLEVGKPPRRAKRPK 169
C S V H HH + N R H+ D LP + P+R RP+
Sbjct: 315 CGASGSVTHQHHRGPPLRRAENRSHLQPSKRGAHSQDKLPQLSQREALHGHPQRLPRPR 491
>TC14416 similar to UP|Q8SMH8 (Q8SMH8) RNA-binding protein, partial (28%)
Length = 695
Score = 26.9 bits (58), Expect = 4.9
Identities = 15/47 (31%), Positives = 23/47 (48%), Gaps = 6/47 (12%)
Frame = +2
Query: 149 LPAAPVSLEVGKPPRRAKRPKESKSDSPNK------KTPKSRKVKKE 189
LP P + + P + K+ ++ S P + KT KSR +KKE
Sbjct: 320 LPTGPSKKKTTRSPSKTKKKEQKVSAKPKRVGNPTVKTLKSRVLKKE 460
>TC13235 similar to UP|Q9LQB6 (Q9LQB6) F4N2.1, partial (38%)
Length = 467
Score = 26.9 bits (58), Expect = 4.9
Identities = 13/34 (38%), Positives = 17/34 (49%)
Frame = +3
Query: 150 PAAPVSLEVGKPPRRAKRPKESKSDSPNKKTPKS 183
P P KPP AKR + S+S +P+ P S
Sbjct: 210 PRPPNMPTSSKPPPSAKRIRSSRSPTPSPSLPNS 311
>TC18996 similar to UP|O24323 (O24323) Cysteine proteinase precursor ,
partial (29%)
Length = 549
Score = 26.6 bits (57), Expect = 6.4
Identities = 17/52 (32%), Positives = 25/52 (47%)
Frame = +3
Query: 265 GNGGWQSACCTTTLSVYPLPAVPNKRHARVGGRKMSGSAFNKLLSRLAAEGH 316
G GGW+S TT+ + V N + +GGRK+ +K+ A GH
Sbjct: 360 GGGGWRSPPPATTVMLRVSVTVRNCLNLWIGGRKVLW*E-SKIKENAGAPGH 512
>TC16355 similar to UP|Q9M4U6 (Q9M4U6) Histone H1 variant, partial (50%)
Length = 584
Score = 26.6 bits (57), Expect = 6.4
Identities = 15/36 (41%), Positives = 19/36 (52%), Gaps = 1/36 (2%)
Frame = +1
Query: 154 VSLEVGKPPRRAKRPKESKSDSPNK-KTPKSRKVKK 188
V+ V KP K PKE+K K K PK +K K+
Sbjct: 85 VTPAVEKPVEEVKAPKETKPAKEKKPKAPKEKKPKQ 192
>TC18874 similar to UP|Q9FJJ0 (Q9FJJ0) Genomic DNA, chromosome 5, TAC
clone:K19B1, partial (33%)
Length = 543
Score = 26.6 bits (57), Expect = 6.4
Identities = 8/21 (38%), Positives = 15/21 (71%), Gaps = 1/21 (4%)
Frame = +3
Query: 255 TGVLRQCYKWGNGG-WQSACC 274
+G+ R+C++W G W++ CC
Sbjct: 408 SGLSRRCFQWWRTGIWRAGCC 470
>TC15784 similar to UP|PE31_ARATH (Q9LHA7) Peroxidase 31 precursor (Atperox
P31) (ATP41) , partial (38%)
Length = 475
Score = 26.6 bits (57), Expect = 6.4
Identities = 20/78 (25%), Positives = 30/78 (37%), Gaps = 8/78 (10%)
Frame = +2
Query: 119 HIHHLPQQVNHLPNMGDSSYGTRELHTTDALPAAPVSLEVGK--------PPRRAKRPKE 170
H HH P+ LP R LH P L+ + PPRR RP+
Sbjct: 188 HPHHRPRHAPPLPPRLPPPQRLRRLH-----PPLLNFLQQSRTRRRHQPLPPRRRLRPRR 352
Query: 171 SKSDSPNKKTPKSRKVKK 188
+ P + P+ R +++
Sbjct: 353 PS*NRPRTRLPQHRLLRR 406
>TC17186 homologue to UP|AAD03372 (AAD03372) Expressed protein, partial (7%)
Length = 595
Score = 26.6 bits (57), Expect = 6.4
Identities = 21/65 (32%), Positives = 29/65 (44%), Gaps = 8/65 (12%)
Frame = -1
Query: 139 GTRELHTTDALPAAPVSLEVGKPPRRAKRPKE-----SKSDSPNK---KTPKSRKVKKEG 190
GT ELHT+ P+ + G RAK P + DS K + P+ KVK+ G
Sbjct: 469 GTHELHTSRQQPSTSPTQRQGDKTNRAKTPTQ*SLHGLYQDSSLKQLLQIPRVPKVKQLG 290
Query: 191 DDLNK 195
L +
Sbjct: 289 TCLQR 275
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.314 0.128 0.385
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 6,037,177
Number of Sequences: 28460
Number of extensions: 84842
Number of successful extensions: 546
Number of sequences better than 10.0: 44
Number of HSP's better than 10.0 without gapping: 537
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 542
length of query: 340
length of database: 4,897,600
effective HSP length: 91
effective length of query: 249
effective length of database: 2,307,740
effective search space: 574627260
effective search space used: 574627260
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (22.0 bits)
S2: 55 (25.8 bits)
Medicago: description of AC124216.5


