

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC124216.10 - phase: 0
(543 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
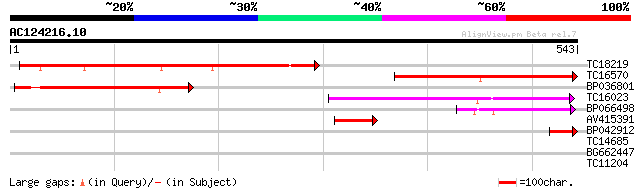
TC18219 242 1e-64
TC16570 similar to UP|Q9LR03 (Q9LR03) F10A5.18, partial (18%) 233 6e-62
BP036801 199 9e-52
TC16023 182 2e-46
BP066498 88 3e-18
AV415391 65 2e-11
BP042912 42 3e-04
TC14685 weakly similar to UP|Q43383 (Q43383) 2A6 protein, partia... 28 3.8
BG662447 28 3.8
TC11204 similar to GB|AAO64909.1|29029060|BT005974 At1g77410 {Ar... 28 4.9
>TC18219
Length = 936
Score = 242 bits (618), Expect = 1e-64
Identities = 137/311 (44%), Positives = 188/311 (60%), Gaps = 24/311 (7%)
Frame = +2
Query: 10 LLPILITTITTTPYLVAAG--------------NGQWQVLQKSIGIVAMHMQLLHNDRIV 55
+LP+ IT +L+ AG GQWQ+L + G+V MH L + I+
Sbjct: 5 VLPLQQIMITILLFLLCAGFSHGAKNEGSLEGRKGQWQMLLNNTGVVGMHTALTSKNTII 184
Query: 56 IFDRTDFGLSKLPLP---NG-KCRHDPRETTVKTDCTAHSVEYNIKSNTFRPLFVQTDVW 111
+FD+T+ G S PL NG +C V + C AHSVEY+I +N RPL + +D W
Sbjct: 185 MFDQTEAGPSGYPLHKRFNGSRCNTRSHNDVVDSTCYAHSVEYDISANRVRPLRLDSDPW 364
Query: 112 CSSGSVNPKGTLVQTGGYNDGDRTIRMFDTCNN--CDWQEFDGGLAARRWYATNHILPD- 168
CSSGS GTL+QTGGY G + IR + C N C W + L+ RWYA++ ILPD
Sbjct: 365 CSSGSFLSNGTLLQTGGYGRGAKRIRFYRPCGNHQCHWIQSRKSLSDERWYASSQILPDH 544
Query: 169 GRQIIIGGRKQFNYEFYPKNNIGV--YRLPFLEQTNDAGAE-NNLYPFVILNVDGNLFIF 225
R +I+GGR+ F YEF PK + + LPFL +TN+ A+ NNLYPF+ L+ DGNLFIF
Sbjct: 545 DRVVIVGGRRTFTYEFIPKISPTEKSFDLPFLHKTNERNAKGNNLYPFLHLSSDGNLFIF 724
Query: 226 ANNRAILFDYTKNVVVRTFPQIPGGDPRSYPSSGSGVLLPLKNLQSKFIEAEVLICGGAP 285
AN +IL + +N V++TFP+IPG R+YPSSGS V+LPL + F + EV++CGG+
Sbjct: 725 ANRDSILLNPRRNRVIKTFPRIPGDGSRNYPSSGSSVMLPLDH-SDNFHKVEVMVCGGSA 901
Query: 286 KGSYQKASKRE 296
G+ + AS R+
Sbjct: 902 TGALRAASNRK 934
>TC16570 similar to UP|Q9LR03 (Q9LR03) F10A5.18, partial (18%)
Length = 794
Score = 233 bits (594), Expect = 6e-62
Identities = 120/179 (67%), Positives = 144/179 (80%), Gaps = 4/179 (2%)
Frame = +3
Query: 369 PIGARFELQNPSHTPRMYHSTAILVRDGRVLVGGSNPHIGYNFNNVLFPTELSIEAFSPS 428
P ++FELQNP+ TP MYHSTA+L+RDGRVLV GSNPH YNF++VLFPTELS+EAFSP
Sbjct: 21 PGRSQFELQNPTKTPWMYHSTAVLLRDGRVLVAGSNPHEYYNFSSVLFPTELSLEAFSPP 200
Query: 429 YLEPRFANVRPRIVASTSELQ---KHGQKLGLRFQVKAAL-DKNLVYVTMLAPPFNTHSF 484
YLEP++ NVRPRIV+ T + Q K+GQ L +RFQV A L + V +TMLAPPFNTHSF
Sbjct: 201 YLEPQYTNVRPRIVSPTPQAQTKLKYGQGLRIRFQVTALLAAPDSVSMTMLAPPFNTHSF 380
Query: 485 SMNQRLLVLESNKVNIVEGTTYDVQVTMPGSPILAPPGFYLLFVVHKEIPSEGIWIQIL 543
SMNQRLLVLE N + V + ++V+VT PGS LAPPGFYL+FVVH+ IPSEGIW+QIL
Sbjct: 381 SMNQRLLVLEPNGLKDVGESIHEVEVTAPGSATLAPPGFYLVFVVHQGIPSEGIWVQIL 557
>BP036801
Length = 540
Score = 199 bits (506), Expect = 9e-52
Identities = 94/175 (53%), Positives = 124/175 (70%), Gaps = 3/175 (1%)
Frame = +1
Query: 5 TYLLFLLPILITTITTTPYLVAAGNGQWQVLQKSIGIVAMHMQLLHNDRIVIFDRTDFGL 64
T + FLL + TT ++T G W LQ+SIGI AMHMQ+++++++VIFDRTDFG
Sbjct: 37 TMITFLLFCITTTCSST-------GGHWVQLQRSIGISAMHMQVMNDNKVVIFDRTDFGP 195
Query: 65 SKLPLPNGKCRHDPRETTVKTDCTAHSVEYNIKSNTFRPLFVQTDVWCSSGSVNPKGTLV 124
S + L NG+CR +PR+ +K DCTAHSV Y++ +NTFRPL +QTD WCSSG++ P GTL+
Sbjct: 196 SNISLSNGRCRFNPRDMALKLDCTAHSVLYDLTTNTFRPLTIQTDAWCSSGALTPDGTLL 375
Query: 125 QTGGYNDGDRTIRMFDTC---NNCDWQEFDGGLAARRWYATNHILPDGRQIIIGG 176
QTGG+NDG +R F C N CDW E L + RWYA+N ILPDGR I++GG
Sbjct: 376 QTGGFNDGYHKLRTFTPCPQHNLCDWLELPQNLTSPRWYASNQILPDGRIIVVGG 540
>TC16023
Length = 894
Score = 182 bits (461), Expect = 2e-46
Identities = 102/242 (42%), Positives = 141/242 (58%), Gaps = 6/242 (2%)
Frame = +2
Query: 306 RIKITDPNPTWVVETMPRARVMGDMVMLPNGDVLIINGAGSGTAGWEYGRDPVLNPVLYK 365
R+ IT + W +E MP R++ DM++LPNG++LIINGA G AG++ R+ L P LY
Sbjct: 5 RMVITXNDNKWEMEYMPEPRLLHDMLILPNGNILIINGAKHGCAGYDNARNASLEPYLYS 184
Query: 366 TNNPIGARFELQNPSHTPRMYHSTAILVRDGRVLVGGSNPHIGYNFNNVLFPTELSIEAF 425
+G RF + + RMYHS+A L+ DGRVLV G NPH Y+F+NV +PTEL ++AF
Sbjct: 185 PKKRLGKRFRVLKSTKIARMYHSSATLLPDGRVLVAGGNPHGRYSFHNVAYPTELRLQAF 364
Query: 426 SPSYLEPRFANVRPRIVASTS------ELQKHGQKLGLRFQVKAALDKNLVYVTMLAPPF 479
P Y+ R+ + RP + S + +G + +RF V N V ++ APPF
Sbjct: 365 VPHYMHRRYHSWRPSNLTVKSGGGGERGVIGYGAEFRVRFLV-GTRPSNEVAFSVYAPPF 541
Query: 480 NTHSFSMNQRLLVLESNKVNIVEGTTYDVQVTMPGSPILAPPGFYLLFVVHKEIPSEGIW 539
THS +MNQR+L L + G D + P SP +AP G+YLL VV+ IPS W
Sbjct: 542 TTHSVAMNQRMLKLRCRSMVRSSGGWVDAVLEAPPSPRVAPSGYYLLTVVNGGIPSVSQW 721
Query: 540 IQ 541
IQ
Sbjct: 722 IQ 727
Score = 29.6 bits (65), Expect = 1.3
Identities = 11/30 (36%), Positives = 19/30 (62%)
Frame = +2
Query: 156 ARRWYATNHILPDGRQIIIGGRKQFNYEFY 185
AR ++++ +LPDGR ++ GG Y F+
Sbjct: 236 ARMYHSSATLLPDGRVLVAGGNPHGRYSFH 325
>BP066498
Length = 512
Score = 88.2 bits (217), Expect = 3e-18
Identities = 51/118 (43%), Positives = 71/118 (59%), Gaps = 4/118 (3%)
Frame = -1
Query: 429 YLEPRFANVRPRIVA--STSELQKHGQKLGLRFQVK--AALDKNLVYVTMLAPPFNTHSF 484
YL+ F P I ST EL ++GQ F V+ A L N + VTM PPF TH F
Sbjct: 512 YLDVDFDIYXPMINEDESTKEL-RYGQFFETVFAVQDGAELTMNDIKVTMYFPPFTTHGF 336
Query: 485 SMNQRLLVLESNKVNIVEGTTYDVQVTMPGSPILAPPGFYLLFVVHKEIPSEGIWIQI 542
SM QRLLVL+ +K+ + Y V++ P + ++APPG+YLLFV H+ +P +G+W+ I
Sbjct: 335 SMGQRLLVLKLDKLMVDSPGLYRVRIAAPPNSVIAPPGYYLLFVNHRGVPGKGMWVHI 162
>AV415391
Length = 138
Score = 65.5 bits (158), Expect = 2e-11
Identities = 27/41 (65%), Positives = 34/41 (82%)
Frame = +2
Query: 312 PNPTWVVETMPRARVMGDMVMLPNGDVLIINGAGSGTAGWE 352
P P WV+ETMP RVM DM++LP G+V+I+NGA +GTAGWE
Sbjct: 2 PKPGWVMETMPIPRVMPDMILLPTGNVMILNGAANGTAGWE 124
>BP042912
Length = 223
Score = 41.6 bits (96), Expect = 3e-04
Identities = 15/26 (57%), Positives = 21/26 (80%)
Frame = -3
Query: 518 LAPPGFYLLFVVHKEIPSEGIWIQIL 543
+AP G+YLLFVVH +PS G W+Q++
Sbjct: 209 IAPAGYYLLFVVHAGVPSTGSWVQVM 132
>TC14685 weakly similar to UP|Q43383 (Q43383) 2A6 protein, partial (38%)
Length = 1323
Score = 28.1 bits (61), Expect = 3.8
Identities = 13/30 (43%), Positives = 18/30 (59%)
Frame = +3
Query: 355 RDPVLNPVLYKTNNPIGARFELQNPSHTPR 384
R P NP+ +T P RF+ +NP+H PR
Sbjct: 234 RSPARNPLRPQT--PTRIRFQTRNPNHRPR 317
>BG662447
Length = 367
Score = 28.1 bits (61), Expect = 3.8
Identities = 13/29 (44%), Positives = 16/29 (54%)
Frame = -3
Query: 371 GARFELQNPSHTPRMYHSTAILVRDGRVL 399
G+R E Q+P P YH VRDG +L
Sbjct: 161 GSRLEAQSPYLQPHHYHLHPYCVRDGSML 75
>TC11204 similar to GB|AAO64909.1|29029060|BT005974 At1g77410 {Arabidopsis
thaliana;}, partial (13%)
Length = 568
Score = 27.7 bits (60), Expect = 4.9
Identities = 20/80 (25%), Positives = 27/80 (33%), Gaps = 2/80 (2%)
Frame = +2
Query: 147 WQEFDGGLAARRWYATNHILPDGRQIIIGGRKQFNYEFYPKNNIGVYRLPFLEQTNDAGA 206
W+ F WY T P+G ++ N G+ R T D +
Sbjct: 146 WESFQSST*PLTWYQTTFDAPEGNDPVVLNLGSMGKGITWVNGQGIGRYWVSFHTPDGTS 325
Query: 207 ENNLY--PFVILNVDGNLFI 224
N Y P IL GNL +
Sbjct: 326 SQNWYHIPRSILKSTGNLLV 385
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.321 0.139 0.427
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 9,804,359
Number of Sequences: 28460
Number of extensions: 141537
Number of successful extensions: 551
Number of sequences better than 10.0: 20
Number of HSP's better than 10.0 without gapping: 537
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 541
length of query: 543
length of database: 4,897,600
effective HSP length: 95
effective length of query: 448
effective length of database: 2,193,900
effective search space: 982867200
effective search space used: 982867200
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 57 (26.6 bits)
Medicago: description of AC124216.10


