

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC124214.5 - phase: 0
(266 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
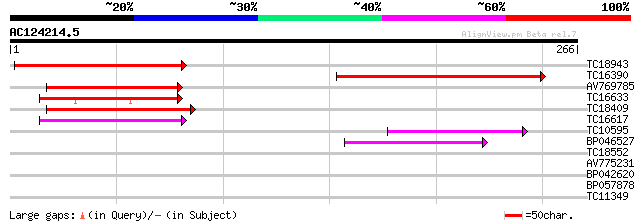
Score E
Sequences producing significant alignments: (bits) Value
TC18943 similar to GB|AAM63374.1|21554299|AY086170 apospory-asso... 104 2e-23
TC16390 similar to GB|AAM63374.1|21554299|AY086170 apospory-asso... 91 3e-19
AV769785 84 2e-17
TC16633 similar to UP|AAPC_PENCL (Q40784) Possible apospory-asso... 79 6e-16
TC18409 weakly similar to UP|Q944A5 (Q944A5) AT3g01590/F4P13_13,... 64 2e-11
TC16617 similar to UP|Q9FJY6 (Q9FJY6) Apospory-associated protei... 55 2e-08
TC10595 similar to UP|AAPC_PENCL (Q40784) Possible apospory-asso... 45 2e-05
BP046527 41 2e-04
TC18552 similar to UP|Q9FIM2 (Q9FIM2) Cell division protein FtsH... 28 2.1
AV775231 27 3.6
BP042620 27 3.6
BP057878 27 3.6
TC11349 similar to UP|Q9SIN1 (Q9SIN1) Expressed protein (At2g425... 26 8.0
>TC18943 similar to GB|AAM63374.1|21554299|AY086170 apospory-associated
protein C {Arabidopsis thaliana;}, partial (32%)
Length = 428
Score = 104 bits (259), Expect = 2e-23
Identities = 51/81 (62%), Positives = 59/81 (71%)
Frame = +2
Query: 3 HSAAVWDFRAATEFTKDWNGIDQVVLRTPQGASARVSLHGAQVTSWRNEQGEELLFTSCK 62
HS+A D R E TKD NGIDQ VLR +G+SA VSLHG QV SW+ E+GEELLF+S K
Sbjct: 86 HSSAESDNRPTVEVTKDKNGIDQFVLRNLRGSSATVSLHGGQVLSWKTERGEELLFSSSK 265
Query: 63 TLSKAPKAIRGGIPICFPQAG 83
+ PK +RGGIPICFPQ G
Sbjct: 266 AIFSPPKPVRGGIPICFPQFG 328
>TC16390 similar to GB|AAM63374.1|21554299|AY086170 apospory-associated
protein C {Arabidopsis thaliana;}, partial (34%)
Length = 532
Score = 90.5 bits (223), Expect = 3e-19
Identities = 53/98 (54%), Positives = 64/98 (65%)
Frame = +1
Query: 154 QNKEML*HLNPRWIVFILAPQT*SQF*IMRGNEHIL*ERKVSLMLPCGIHGRRNRSRWWT 213
QNKEML*HLN RWI IL P T *IM+G+EH+ *ERK ML GI GR+N S WW
Sbjct: 10 QNKEML*HLNQRWIGSILIPATW*LS*IMKGSEHL**ERKDYQMLLFGIRGRKNPSPWWI 189
Query: 214 SVMKSTNKCFV*MVQL*RNL*T*SLERSGQGGFNSQLC 251
MKST+ C+V MVQ N *S E++G+ ++ LC
Sbjct: 190 *GMKSTSTCYVLMVQQLENQSP*SQEKNGEAAWSFLLC 303
>AV769785
Length = 600
Score = 84.3 bits (207), Expect = 2e-17
Identities = 39/64 (60%), Positives = 48/64 (74%)
Frame = +1
Query: 18 KDWNGIDQVVLRTPQGASARVSLHGAQVTSWRNEQGEELLFTSCKTLSKAPKAIRGGIPI 77
K NG+D+++LR +G S V L+G Q+TSW+NE GEELLF S K K PK+IRGGIPI
Sbjct: 91 KGVNGLDKIILREVRGFSCEVYLYGGQITSWKNELGEELLFVSGKANFKPPKSIRGGIPI 270
Query: 78 CFPQ 81
CFPQ
Sbjct: 271 CFPQ 282
>TC16633 similar to UP|AAPC_PENCL (Q40784) Possible apospory-associated
protein C, partial (53%)
Length = 615
Score = 79.3 bits (194), Expect = 6e-16
Identities = 40/70 (57%), Positives = 51/70 (72%), Gaps = 3/70 (4%)
Frame = +3
Query: 15 EFTKDWNGIDQVVLR--TPQGASARVSLHGAQVTSWRNEQGEE-LLFTSCKTLSKAPKAI 71
+ K NG+D++++R T G SA V L+GA VTSW+N+ GEE LLF S K++ PKAI
Sbjct: 21 DLLKGINGLDKILIRDATATGCSAEVYLYGAHVTSWKNDHGEEELLFVSSKSIFDPPKAI 200
Query: 72 RGGIPICFPQ 81
RGGIPICFPQ
Sbjct: 201RGGIPICFPQ 230
>TC18409 weakly similar to UP|Q944A5 (Q944A5) AT3g01590/F4P13_13, partial
(51%)
Length = 609
Score = 64.3 bits (155), Expect = 2e-11
Identities = 32/70 (45%), Positives = 45/70 (63%)
Frame = +1
Query: 18 KDWNGIDQVVLRTPQGASARVSLHGAQVTSWRNEQGEELLFTSCKTLSKAPKAIRGGIPI 77
+D +G+ +++L P G+SA V L+G Q+ SW+N + EELLF S K K KAIRGGI
Sbjct: 157 QDKDGLPRILLTEPNGSSAEVLLYGGQIISWKNHRKEELLFMSSKANRKQLKAIRGGISA 336
Query: 78 CFPQAGHIVL 87
F + G + L
Sbjct: 337 YFARFGELGL 366
>TC16617 similar to UP|Q9FJY6 (Q9FJY6) Apospory-associated protein C-like
protein, partial (42%)
Length = 551
Score = 54.7 bits (130), Expect = 2e-08
Identities = 25/69 (36%), Positives = 38/69 (54%)
Frame = +2
Query: 15 EFTKDWNGIDQVVLRTPQGASARVSLHGAQVTSWRNEQGEELLFTSCKTLSKAPKAIRGG 74
+ T+ + ++VL +P G+ A + L G +TSW+ G +LLF + K I GG
Sbjct: 143 KLTEGEGSLPKLVLTSPAGSEAEIYLFGGCITSWKVPSGNDLLFVRPDAVFNKKKPISGG 322
Query: 75 IPICFPQAG 83
+P CFPQ G
Sbjct: 323 VPHCFPQFG 349
>TC10595 similar to UP|AAPC_PENCL (Q40784) Possible apospory-associated
protein C, partial (28%)
Length = 726
Score = 44.7 bits (104), Expect = 2e-05
Identities = 27/66 (40%), Positives = 38/66 (56%)
Frame = +1
Query: 178 QF*IMRGNEHIL*ERKVSLMLPCGIHGRRNRSRWWTSVMKSTNKCFV*MVQL*RNL*T*S 237
Q IM+ EH+ + LML CG G R + W VM S + CFV* + L +NL *+
Sbjct: 16 QLLIMKRREHLFCLKMAFLMLWCGTLGTRKQRLWPILVMMSISICFV*RLLLLKNLSL*N 195
Query: 238 LERSGQ 243
LE++G+
Sbjct: 196 LEKNGK 213
>BP046527
Length = 521
Score = 40.8 bits (94), Expect = 2e-04
Identities = 28/67 (41%), Positives = 36/67 (52%)
Frame = -2
Query: 158 ML*HLNPRWIVFILAPQT*SQF*IMRGNEHIL*ERKVSLMLPCGIHGRRNRSRWWTSVMK 217
ML*HL+ + +I A F IMR ++H+ + V LML CGI G R S W VM
Sbjct: 517 ML*HLHQKLTGYISALLPRFPFLIMRKSDHLCCVKMVFLMLLCGILGIRKPSLWLILVMM 338
Query: 218 STNKCFV 224
S + FV
Sbjct: 337 SISIWFV 317
>TC18552 similar to UP|Q9FIM2 (Q9FIM2) Cell division protein FtsH, partial
(10%)
Length = 643
Score = 27.7 bits (60), Expect = 2.1
Identities = 12/26 (46%), Positives = 15/26 (57%), Gaps = 6/26 (23%)
Frame = +3
Query: 205 RRNRSRWWTSV------MKSTNKCFV 224
RRN+SRWW + + TNK FV
Sbjct: 390 RRNQSRWWNDIPIF*LRFRPTNKTFV 467
>AV775231
Length = 447
Score = 26.9 bits (58), Expect = 3.6
Identities = 16/41 (39%), Positives = 20/41 (48%)
Frame = -3
Query: 78 CFPQAGHIVLSSAFECLLQQMET*P*YHE*GISMPSHLVSH 118
C P H LS F L +QM * HE* + P L++H
Sbjct: 169 CHPIFWHFCLSFTFANLHRQMNG*EHLHE*LLCSPISLITH 47
>BP042620
Length = 511
Score = 26.9 bits (58), Expect = 3.6
Identities = 22/86 (25%), Positives = 37/86 (42%)
Frame = -3
Query: 138 LRHLITWTTFPKKHELQNKEML*HLNPRWIVFILAPQT*SQF*IMRGNEHIL*ERKVSLM 197
L HL+T +F ELQ +++ P ++F+L +T + *
Sbjct: 365 LSHLVTGPSFLHPLELQEHDLI---QPMVVIFLLTQKTGQRI*----------------- 246
Query: 198 LPCGIHGRRNRSRWWTSVMKSTNKCF 223
P H R++ W+S +K T+ CF
Sbjct: 245 -PPYFH-LPGRTQHWSSSLKGTSHCF 174
>BP057878
Length = 546
Score = 26.9 bits (58), Expect = 3.6
Identities = 8/17 (47%), Positives = 14/17 (82%)
Frame = +3
Query: 209 SRWWTSVMKSTNKCFV* 225
S WW ++++S+ KCF+*
Sbjct: 384 SCWWQTILQSSTKCFI* 434
>TC11349 similar to UP|Q9SIN1 (Q9SIN1) Expressed protein
(At2g42580/F14N22.15), partial (5%)
Length = 474
Score = 25.8 bits (55), Expect = 8.0
Identities = 13/25 (52%), Positives = 15/25 (60%)
Frame = +1
Query: 104 YHE*GISMPSHLVSHLHTTHTCWFL 128
YHE I +PSH S L T H+C L
Sbjct: 109 YHE--IPIPSHPFSSLSTIHSCLVL 177
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.351 0.152 0.549
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 5,147,471
Number of Sequences: 28460
Number of extensions: 75128
Number of successful extensions: 806
Number of sequences better than 10.0: 26
Number of HSP's better than 10.0 without gapping: 797
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 804
length of query: 266
length of database: 4,897,600
effective HSP length: 89
effective length of query: 177
effective length of database: 2,364,660
effective search space: 418544820
effective search space used: 418544820
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 14 ( 7.1 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 38 (22.0 bits)
S2: 54 (25.4 bits)
Medicago: description of AC124214.5


