

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC124214.3 - phase: 0
(529 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
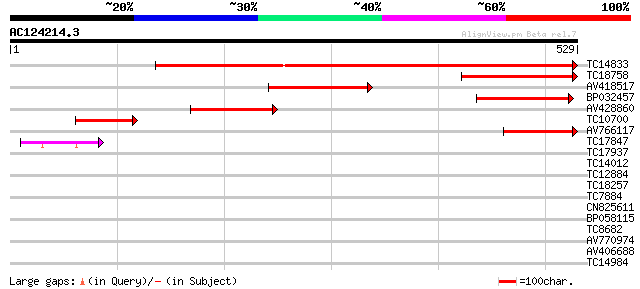
Score E
Sequences producing significant alignments: (bits) Value
TC14833 similar to UP|Q9ZPK4 (Q9ZPK4) Glutamyl-tRNA reductase (... 540 e-154
TC18758 similar to UP|HMA2_CUCSA (P49295) Glutamyl-tRNA reductas... 182 9e-47
AV418517 138 2e-33
BP032457 129 1e-30
AV428860 126 1e-29
TC10700 similar to UP|Q9ZPK4 (Q9ZPK4) Glutamyl-tRNA reductase (... 98 4e-21
AV766117 65 2e-11
TC17847 homologue to UP|Q9ZPK4 (Q9ZPK4) Glutamyl-tRNA reductase ... 42 3e-04
TC17937 similar to GB|AAM52235.1|21464561|AY120692 AT5g19180/T24... 33 0.089
TC14012 similar to UP|Q8GSM9 (Q8GSM9) Squalene monooxygenase 2 ... 31 0.57
TC12884 similar to UP|Q9SJM8 (Q9SJM8) Expressed protein, partial... 30 0.98
TC18257 weakly similar to UP|Q9LFY5 (Q9LFY5) T7N9.6, partial (46%) 29 1.7
TC7884 similar to UP|RL4_PRUAR (Q9XF97) 60S ribosomal protein L4... 29 1.7
CN825611 28 4.9
BP058115 27 6.3
TC8682 similar to UP|HPPD_SOLSC (Q9ARF9) 4-hydroxyphenylpyruvate... 27 6.3
AV770974 27 8.3
AV406688 27 8.3
TC14984 similar to GB|AAM62447.1|21553354|AY085214 glycine-rich ... 27 8.3
>TC14833 similar to UP|Q9ZPK4 (Q9ZPK4) Glutamyl-tRNA reductase (GluTR) ,
partial (72%)
Length = 1538
Score = 540 bits (1390), Expect = e-154
Identities = 277/394 (70%), Positives = 333/394 (84%), Gaps = 1/394 (0%)
Frame = +3
Query: 137 VALSQHRGVREVTDWISKKSGVSVPEISKHQILLYNKDATQHLFEVAAGLDSLVLGEGQI 196
VALSQHRGV+EV +W+SK S V V E+ KH+ LLYNKDATQH+FEV+AGLDSLVLGEGQI
Sbjct: 3 VALSQHRGVKEVMEWMSKTSSVPVSELCKHRFLLYNKDATQHIFEVSAGLDSLVLGEGQI 182
Query: 197 LSQVKQVVKSGQGVPGFDRKISGLFKQAISVGKRVRTETNISSGSVSVSSAAVELALMKR 256
L+QVKQVVK GQGV GF R ISGLFK AI+VGKRVR ETNI++G+VSVSSAAVELA MK
Sbjct: 183 LAQVKQVVKVGQGVTGFGRNISGLFKHAITVGKRVRAETNIAAGAVSVSSAAVELAFMK- 359
Query: 257 LESSFGDAKVLVIGAGKMGKLVIKHLVAKGCRKMVVVNRSEEKVNAIRKELKDVDIVYRP 316
L + DA++LVIGAGKMGKLVIKHLVAKGC KMVVVNR+EE+V AIR+ELKDV+I+Y+P
Sbjct: 360 LPGTSHDARMLVIGAGKMGKLVIKHLVAKGCTKMVVVNRTEERVAAIREELKDVEIIYKP 539
Query: 317 LSEMMECAAEADVIFTGTASESLLFSKENVEILPPVGQGVR-RRLFVDISIPRNVDPGVS 375
LSEM+ C EADV+FT TASE+ LF K++V+ LPP Q V +R+F+DIS+PRNVD VS
Sbjct: 540 LSEMLTCVGEADVVFTSTASETPLFMKDHVKDLPPASQDVGGQRIFIDISVPRNVDSCVS 719
Query: 376 ELENALVYNVDDLREVVDANKEDRQRKAMEARGIIKEELNTFEAWKDSLETVPTIKKFRA 435
+L++ VYNVDDL+EVV ANKEDR RKAMEA+ II EE FEAW+DSLETVPTIKK RA
Sbjct: 720 DLDSVRVYNVDDLKEVVAANKEDRLRKAMEAQTIIAEESKQFEAWRDSLETVPTIKKLRA 899
Query: 436 YVERIRASELEKCLSKMRGDVSKEQKEAMYALSMGIVNKLLHGPMQHLRCDGNDTKCLDE 495
Y ERIR +EL+KCL KM D+SK+ + A+ LS G+VN+LLHGPMQHLRCDG+D++ L+E
Sbjct: 900 YAERIRVAELDKCLGKMGEDISKKMRRAVDDLSRGMVNRLLHGPMQHLRCDGSDSRTLNE 1079
Query: 496 VLENMRALNRMYDLETEISLMEEKIRVKMEKAKK 529
LENM ALNRM+ LETEIS++E+KIR K+E+ +K
Sbjct: 1080TLENMHALNRMFSLETEISVLEQKIRAKVEQNQK 1181
>TC18758 similar to UP|HMA2_CUCSA (P49295) Glutamyl-tRNA reductase 2,
chloroplast precursor (GluTR) , partial (20%)
Length = 635
Score = 182 bits (463), Expect = 9e-47
Identities = 87/108 (80%), Positives = 98/108 (90%)
Frame = +1
Query: 422 DSLETVPTIKKFRAYVERIRASELEKCLSKMRGDVSKEQKEAMYALSMGIVNKLLHGPMQ 481
DSLETVPTIKKFRAY ERI +SE+EKCLSKM D+SKEQK+A+Y+L MGIVNKLLHGPMQ
Sbjct: 1 DSLETVPTIKKFRAYAERISSSEVEKCLSKMGNDISKEQKDAIYSLGMGIVNKLLHGPMQ 180
Query: 482 HLRCDGNDTKCLDEVLENMRALNRMYDLETEISLMEEKIRVKMEKAKK 529
HLRCDGND +CL EVLENMRALNRMYDLETE SL+EEKI+ KME+ +K
Sbjct: 181 HLRCDGNDNRCLSEVLENMRALNRMYDLETETSLIEEKIKAKMERVQK 324
>AV418517
Length = 298
Score = 138 bits (348), Expect = 2e-33
Identities = 69/97 (71%), Positives = 86/97 (88%)
Frame = +3
Query: 242 VSVSSAAVELALMKRLESSFGDAKVLVIGAGKMGKLVIKHLVAKGCRKMVVVNRSEEKVN 301
VSVSSAAVELALMK ++S + ++L+IGAGKMGKLVIKHLVAKG +KMVVVNR+EEKV
Sbjct: 6 VSVSSAAVELALMKLPQASHANVRMLIIGAGKMGKLVIKHLVAKGSKKMVVVNRTEEKVA 185
Query: 302 AIRKELKDVDIVYRPLSEMMECAAEADVIFTGTASES 338
AIR+ELKD++I+Y+PLSEM+ C EADV+FT TAS++
Sbjct: 186 AIREELKDIEIIYKPLSEMLTCVGEADVVFTCTASDT 296
>BP032457
Length = 546
Score = 129 bits (324), Expect = 1e-30
Identities = 61/91 (67%), Positives = 76/91 (83%)
Frame = -1
Query: 436 YVERIRASELEKCLSKMRGDVSKEQKEAMYALSMGIVNKLLHGPMQHLRCDGNDTKCLDE 495
Y ERIRA+ELEKCL KM D+SK+ + A+ LS GIVNK+LHGPMQHLRCDG+D++ L E
Sbjct: 546 YAERIRAAELEKCLGKMGDDISKKTQRAVDDLSRGIVNKMLHGPMQHLRCDGSDSRTLSE 367
Query: 496 VLENMRALNRMYDLETEISLMEEKIRVKMEK 526
LENM ALNRM+ LETEIS++E+KIR K+E+
Sbjct: 366 TLENMHALNRMFSLETEISVLEQKIRAKVEQ 274
>AV428860
Length = 248
Score = 126 bits (316), Expect = 1e-29
Identities = 65/82 (79%), Positives = 73/82 (88%)
Frame = +1
Query: 169 LLYNKDATQHLFEVAAGLDSLVLGEGQILSQVKQVVKSGQGVPGFDRKISGLFKQAISVG 228
LLYNKDATQH+FEV+ GLDSLV+GE QIL+QVKQVVK+GQGV GF R ISGLFK AI+V
Sbjct: 1 LLYNKDATQHIFEVSTGLDSLVMGESQILAQVKQVVKAGQGVNGFGRNISGLFKHAITV* 180
Query: 229 KRVRTETNISSGSVSVSSAAVE 250
KRVRTETNI+ G+VSVSS AVE
Sbjct: 181 KRVRTETNIAVGAVSVSSTAVE 246
>TC10700 similar to UP|Q9ZPK4 (Q9ZPK4) Glutamyl-tRNA reductase (GluTR) ,
partial (20%)
Length = 455
Score = 97.8 bits (242), Expect = 4e-21
Identities = 46/58 (79%), Positives = 53/58 (91%)
Frame = +2
Query: 62 SPLEILKTSSADRYTKEKSSIIVIGLNVHTAPVEMREKLAIPEAQWPQVIQELCALNH 119
S LE LKTS+ADRYTKE+SSI+VIGL+VHT PVEMREKLAIPEA+WP+ I ELC+LNH
Sbjct: 281 SALEQLKTSAADRYTKERSSIVVIGLSVHTTPVEMREKLAIPEAEWPRAIGELCSLNH 454
>AV766117
Length = 427
Score = 65.5 bits (158), Expect = 2e-11
Identities = 30/69 (43%), Positives = 48/69 (69%)
Frame = -1
Query: 461 KEAMYALSMGIVNKLLHGPMQHLRCDGNDTKCLDEVLENMRALNRMYDLETEISLMEEKI 520
++ + + GIV K+LHGP+ LRCDG+ T+ L + LEN +ALN+M+ LE +I +++KI
Sbjct: 424 QKPVVVFTRGIVTKMLHGPIHPLRCDGSATRPLSKTLENRQALNKMFTLEPKIFFLDQKI 245
Query: 521 RVKMEKAKK 529
R K+ + K
Sbjct: 244 RPKVGQNPK 218
>TC17847 homologue to UP|Q9ZPK4 (Q9ZPK4) Glutamyl-tRNA reductase (GluTR) ,
partial (6%)
Length = 418
Score = 41.6 bits (96), Expect = 3e-04
Identities = 29/82 (35%), Positives = 44/82 (53%), Gaps = 5/82 (6%)
Frame = +1
Query: 11 HTLRSSVKFSPAQFPKSTF--APQRLSFTTPKIPKCTLRSDNPLPQNFIVSKP---SPLE 65
H SS + SP+ + +F R + + +C ++ + + + S S LE
Sbjct: 172 HNCSSSSRASPSTVTQLSFPGGKPRRNLIHKGVVRCDAQASDVVVKTDGSSNATALSALE 351
Query: 66 ILKTSSADRYTKEKSSIIVIGL 87
LK S+ADRYTKE+SSI+VIGL
Sbjct: 352 QLKASAADRYTKERSSIVVIGL 417
>TC17937 similar to GB|AAM52235.1|21464561|AY120692 AT5g19180/T24G5_80
{Arabidopsis thaliana;}, partial (26%)
Length = 566
Score = 33.5 bits (75), Expect = 0.089
Identities = 21/58 (36%), Positives = 34/58 (58%), Gaps = 1/58 (1%)
Frame = +2
Query: 264 AKVLVIGAGKMGKLVIKHLVAKGCRKMVVVNRSE-EKVNAIRKELKDVDIVYRPLSEM 320
AKVLV+GAG +G ++K L G R + V++ E N R+ L ++ V +P +E+
Sbjct: 275 AKVLVVGAGGLGCELLKDLALTGFRNLEVIDMDRIEVTNLNRQFLFRLEDVGKPKAEV 448
>TC14012 similar to UP|Q8GSM9 (Q8GSM9) Squalene monooxygenase 2 , partial
(13%)
Length = 480
Score = 30.8 bits (68), Expect = 0.57
Identities = 24/88 (27%), Positives = 42/88 (47%)
Frame = +2
Query: 220 LFKQAISVGKRVRTETNISSGSVSVSSAAVELALMKRLESSFGDAKVLVIGAGKMGKLVI 279
L+ + R ++ + S +V++AA E E GDA V+++GAG G +
Sbjct: 188 LYSVVFAGNSRGSSQGKAAPRSENVTTAAGECGS----EKLDGDADVIIVGAGVAGS-AL 352
Query: 280 KHLVAKGCRKMVVVNRSEEKVNAIRKEL 307
H + K R++ V+ R + + I EL
Sbjct: 353 AHTLGKDGRRVHVIERDLSEPDRIVGEL 436
>TC12884 similar to UP|Q9SJM8 (Q9SJM8) Expressed protein, partial (29%)
Length = 475
Score = 30.0 bits (66), Expect = 0.98
Identities = 20/64 (31%), Positives = 28/64 (43%), Gaps = 1/64 (1%)
Frame = +3
Query: 20 SPAQFPKSTFAPQRLSFTTPKIPKC-TLRSDNPLPQNFIVSKPSPLEILKTSSADRYTKE 78
SPAQ S P+R S TP P C + + P P + P+P ++S R
Sbjct: 165 SPAQTSSSAAYPRRRSNPTPASPSCPST*APPPRPPPPVKFTPAPASAAPSTSTTRSLSS 344
Query: 79 KSSI 82
SS+
Sbjct: 345 TSSV 356
>TC18257 weakly similar to UP|Q9LFY5 (Q9LFY5) T7N9.6, partial (46%)
Length = 583
Score = 29.3 bits (64), Expect = 1.7
Identities = 19/48 (39%), Positives = 30/48 (61%), Gaps = 1/48 (2%)
Frame = +3
Query: 235 TNISSGSVSVSSAAVELALMKRLESSFGDAKVLVI-GAGKMGKLVIKH 281
++ +S S+S+S A A+ ++S G K+LVI GAG G ++IKH
Sbjct: 9 SSFNSVSLSLSLAP---AMAMAMQSGIGITKILVIAGAGYTGTVLIKH 143
>TC7884 similar to UP|RL4_PRUAR (Q9XF97) 60S ribosomal protein L4 (L1),
partial (97%)
Length = 1476
Score = 29.3 bits (64), Expect = 1.7
Identities = 24/87 (27%), Positives = 37/87 (41%)
Frame = +1
Query: 380 ALVYNVDDLREVVDANKEDRQRKAMEARGIIKEELNTFEAWKDSLETVPTIKKFRAYVER 439
A + N D+++ VV K+D +R ++ K L T K+ E
Sbjct: 919 ARIINSDEVQSVVRPIKKDVKRATLK-----KNPLKNLNVMLRLNPYAKTAKRMALLAEA 1083
Query: 440 IRASELEKCLSKMRGDVSKEQKEAMYA 466
R ++ L KMR VSKE+ A+ A
Sbjct: 1084 QRLQAKKEKLDKMRKIVSKEEASAIKA 1164
>CN825611
Length = 623
Score = 27.7 bits (60), Expect = 4.9
Identities = 11/25 (44%), Positives = 16/25 (64%)
Frame = +2
Query: 262 GDAKVLVIGAGKMGKLVIKHLVAKG 286
G +VLV+G +GK + HL+ KG
Sbjct: 443 GQVRVLVVGDSGVGKTSLVHLIVKG 517
>BP058115
Length = 590
Score = 27.3 bits (59), Expect = 6.3
Identities = 17/54 (31%), Positives = 27/54 (49%)
Frame = +1
Query: 359 RLFVDISIPRNVDPGVSELENALVYNVDDLREVVDANKEDRQRKAMEARGIIKE 412
RLF + +P PG+S + + LR + D E R+ A+E GI+K+
Sbjct: 286 RLFTNQLLPTL--PGISMADALSLIPAAVLRNLADKLYEKRKNAALEIEGIVKQ 441
>TC8682 similar to UP|HPPD_SOLSC (Q9ARF9) 4-hydroxyphenylpyruvate
dioxygenase (4HPPD) (HPD) (HPPDase) , partial (29%)
Length = 576
Score = 27.3 bits (59), Expect = 6.3
Identities = 22/60 (36%), Positives = 24/60 (39%)
Frame = +1
Query: 4 AASSSTDHTLRSSVKFSPAQFPKSTFAPQRLSFTTPKIPKCTLRSDNPLPQNFIVSKPSP 63
+ASSS HTL S P P S P S P S +P P SKPSP
Sbjct: 397 SASSSPPHTLPKSPSPPPPPSPLSPPPPASPSPPPTASPSAPSPSKSPTP-----SKPSP 561
>AV770974
Length = 487
Score = 26.9 bits (58), Expect = 8.3
Identities = 23/84 (27%), Positives = 41/84 (48%), Gaps = 4/84 (4%)
Frame = -2
Query: 434 RAYVERIRASELEKCLSKMRGDVSKEQKEAMYA-LSMGIVNKLLHGPMQHLRCDG---ND 489
+AY++R+ AS+ +K + + KEAM+ L+ G+VN + G DG N
Sbjct: 423 QAYIKRVTASKESTRETKKKPIHKMKMKEAMHTKLNEGVVNGNVDGIASGSVMDGGIVNG 244
Query: 490 TKCLDEVLENMRALNRMYDLETEI 513
+ +LE + + DL T++
Sbjct: 243 STIYGYILEGLI*EMKEMDLHTQM 172
>AV406688
Length = 423
Score = 26.9 bits (58), Expect = 8.3
Identities = 16/53 (30%), Positives = 23/53 (43%)
Frame = +1
Query: 1 MSMAASSSTDHTLRSSVKFSPAQFPKSTFAPQRLSFTTPKIPKCTLRSDNPLP 53
+S+ +SS S+V F P+ F P T K+P L + PLP
Sbjct: 232 LSLNPTSSPPMNSSSTVSFFPSTFYPPQLNPTHFPNLTQKLPPQHLSQNPPLP 390
>TC14984 similar to GB|AAM62447.1|21553354|AY085214 glycine-rich RNA binding
protein 7 {Arabidopsis thaliana;} , partial (19%)
Length = 483
Score = 26.9 bits (58), Expect = 8.3
Identities = 23/67 (34%), Positives = 31/67 (45%)
Frame = +2
Query: 15 SSVKFSPAQFPKSTFAPQRLSFTTPKIPKCTLRSDNPLPQNFIVSKPSPLEILKTSSADR 74
+++K SP+ +T AP S T+P P +S P P + PSP TSSA
Sbjct: 197 ATLKSSPSS-ATATAAPSTKSTTSPPPPSTL*KSSTPAPTPPRAAVPSP---KSTSSAAP 364
Query: 75 YTKEKSS 81
T SS
Sbjct: 365 PTAPTSS 385
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.316 0.132 0.361
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 7,054,589
Number of Sequences: 28460
Number of extensions: 84723
Number of successful extensions: 407
Number of sequences better than 10.0: 38
Number of HSP's better than 10.0 without gapping: 405
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 405
length of query: 529
length of database: 4,897,600
effective HSP length: 94
effective length of query: 435
effective length of database: 2,222,360
effective search space: 966726600
effective search space used: 966726600
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.6 bits)
S2: 57 (26.6 bits)
Medicago: description of AC124214.3


