

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC124214.11 + phase: 0 /pseudo
(240 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
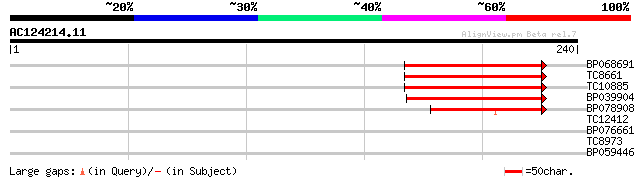
Score E
Sequences producing significant alignments: (bits) Value
BP068691 65 8e-12
TC8661 weakly similar to UP|O81348 (O81348) Symbiotic ammonium t... 65 8e-12
TC10885 59 6e-10
BP039904 56 6e-09
BP078908 39 6e-04
TC12412 weakly similar to UP|O81348 (O81348) Symbiotic ammonium ... 37 0.004
BP076661 30 0.48
TC8973 similar to PIR|A86477|A86477 protein F15O4.33 [imported] ... 26 5.3
BP059446 26 6.9
>BP068691
Length = 500
Score = 65.5 bits (158), Expect = 8e-12
Identities = 30/60 (50%), Positives = 45/60 (75%)
Frame = -1
Query: 168 LPEIEARFCDRNVLIRVHCEKIKGVVEKTIHKIEKLNLKVTNSSFMTFGSCALDITIIAQ 227
LP +EAR +++VL+R+HC+K KG++ K + +I+ L+L V NSS + FG LDITI+AQ
Sbjct: 344 LPHLEARVSEKDVLLRLHCQKQKGLLLKILFEIQNLHLFVVNSSVLPFGDSILDITIVAQ 165
>TC8661 weakly similar to UP|O81348 (O81348) Symbiotic ammonium
transporter, partial (34%)
Length = 1007
Score = 65.5 bits (158), Expect = 8e-12
Identities = 30/60 (50%), Positives = 45/60 (75%)
Frame = +1
Query: 168 LPEIEARFCDRNVLIRVHCEKIKGVVEKTIHKIEKLNLKVTNSSFMTFGSCALDITIIAQ 227
LP +EAR +++VL+R+HC+K KG++ K + +I+ L+L V NSS + FG LDITI+AQ
Sbjct: 310 LPHVEARVSEKDVLLRLHCKKQKGLLLKILFEIQNLHLFVVNSSVLPFGDSILDITIVAQ 489
>TC10885
Length = 573
Score = 59.3 bits (142), Expect = 6e-10
Identities = 29/60 (48%), Positives = 38/60 (63%)
Frame = +2
Query: 168 LPEIEARFCDRNVLIRVHCEKIKGVVEKTIHKIEKLNLKVTNSSFMTFGSCALDITIIAQ 227
LPEI AR + VLI +HCEK G K + +E L+L V SS + FG+ AL +T+IAQ
Sbjct: 137 LPEIAARALGKEVLIEIHCEKENGTELKILDHLENLHLSVNGSSVLPFGNSALSVTVIAQ 316
>BP039904
Length = 544
Score = 55.8 bits (133), Expect = 6e-09
Identities = 27/59 (45%), Positives = 38/59 (63%)
Frame = -2
Query: 169 PEIEARFCDRNVLIRVHCEKIKGVVEKTIHKIEKLNLKVTNSSFMTFGSCALDITIIAQ 227
P+++AR + VLI +HCE+ G+ K + +E L L VT SS + FG+ L ITIIAQ
Sbjct: 462 PDVKARVLENEVLIVIHCERENGIELKILDLLENLRLCVTGSSVLPFGNSTLSITIIAQ 286
>BP078908
Length = 443
Score = 39.3 bits (90), Expect = 6e-04
Identities = 19/50 (38%), Positives = 33/50 (66%), Gaps = 1/50 (2%)
Frame = -1
Query: 179 NVLIRVHCEKIKGVVEKTIHKIEKLN-LKVTNSSFMTFGSCALDITIIAQ 227
++LI++HC+K G + ++EK + L V +SS + FG+ D+TIIA+
Sbjct: 440 DLLIKIHCDKQSGCAATILRELEKHDYLTVQSSSILPFGNNITDVTIIAK 291
>TC12412 weakly similar to UP|O81348 (O81348) Symbiotic ammonium
transporter, partial (7%)
Length = 654
Score = 36.6 bits (83), Expect = 0.004
Identities = 17/40 (42%), Positives = 25/40 (62%)
Frame = +2
Query: 181 LIRVHCEKIKGVVEKTIHKIEKLNLKVTNSSFMTFGSCAL 220
LIR+ CEK +GV+ + +I LNL V N+ + FG+ L
Sbjct: 2 LIRIPCEKQEGVLMNILKEIHNLNLSVINAGTLLFGTSKL 121
>BP076661
Length = 532
Score = 29.6 bits (65), Expect = 0.48
Identities = 13/29 (44%), Positives = 21/29 (71%)
Frame = -3
Query: 199 KIEKLNLKVTNSSFMTFGSCALDITIIAQ 227
++EK +L V ++S + FG+ LDITI+ Q
Sbjct: 515 ELEKHHLTVQSTSILPFGNNTLDITIVTQ 429
>TC8973 similar to PIR|A86477|A86477 protein F15O4.33 [imported] -
Arabidopsis thaliana {Arabidopsis thaliana;}, partial
(66%)
Length = 1358
Score = 26.2 bits (56), Expect = 5.3
Identities = 10/18 (55%), Positives = 13/18 (71%)
Frame = -2
Query: 49 TLHNTIHTFIQTSTLELQ 66
TLHNT+HTF+ S + Q
Sbjct: 1087 TLHNTLHTFLSFSIILFQ 1034
>BP059446
Length = 542
Score = 25.8 bits (55), Expect = 6.9
Identities = 17/65 (26%), Positives = 30/65 (46%)
Frame = +2
Query: 176 CDRNVLIRVHCEKIKGVVEKTIHKIEKLNLKVTNSSFMTFGSCALDITIIAQPTLSFYVF 235
C R++ E + ++ K + +I+ + MTF LD T+ QP SF
Sbjct: 230 CINFSFFRINSESQEELLTKLLPRIDSAFRR------MTFKHALLDFTMC*QPRNSFQWV 391
Query: 236 HLRTF 240
HL+++
Sbjct: 392 HLQSY 406
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.364 0.163 0.551
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 4,095,538
Number of Sequences: 28460
Number of extensions: 54909
Number of successful extensions: 784
Number of sequences better than 10.0: 18
Number of HSP's better than 10.0 without gapping: 781
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 782
length of query: 240
length of database: 4,897,600
effective HSP length: 88
effective length of query: 152
effective length of database: 2,393,120
effective search space: 363754240
effective search space used: 363754240
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 14 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 36 (21.5 bits)
S2: 54 (25.4 bits)
Medicago: description of AC124214.11


