

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC123976.3 - phase: 0
(447 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
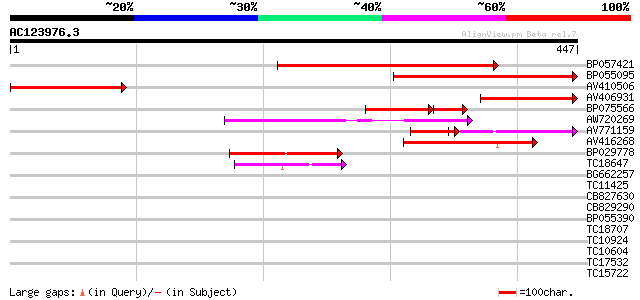
Score E
Sequences producing significant alignments: (bits) Value
BP057421 329 6e-91
BP055095 233 6e-62
AV410506 167 3e-42
AV406931 122 1e-28
BP075566 67 2e-18
AW720269 80 5e-16
AV771159 59 5e-15
AV416268 73 1e-13
BP029778 66 1e-11
TC18647 similar to UP|Q9ASR9 (Q9ASR9) At2g01320/F10A8.20, partia... 62 3e-10
BG662257 40 0.001
TC11425 similar to UP|Q9M1Q9 (Q9M1Q9) P-glycoprotein-like proeti... 40 0.001
CB827630 37 0.009
CB829290 35 0.019
BP055390 35 0.019
TC18707 homologue to PIR|T02901|T02901 MSP1 protein homolog T13J... 35 0.033
TC10924 homologue to UP|Q8H9D4 (Q8H9D4) 26S proteasome AAA-ATPas... 34 0.056
TC10604 similar to AAQ89671 (AAQ89671) At4g33460, partial (36%) 33 0.095
TC17532 similar to UP|Q9LZ98 (Q9LZ98) ABC transporter-like prote... 33 0.12
TC15722 similar to UP|Q9FMU0 (Q9FMU0) Similarity to ABC transpor... 31 0.47
>BP057421
Length = 547
Score = 329 bits (843), Expect = 6e-91
Identities = 164/174 (94%), Positives = 170/174 (97%)
Frame = +3
Query: 212 KLDSELRVQGDITYNGHRLNEFVPRKTSAYISQNDVHVGEMTVKETLDFSARCQGVGTRY 271
K+D ELRV G+ITYNGH+LNEFVPRKT+AYISQNDVHVGEMTVKETLDFSARCQGVGTRY
Sbjct: 24 KMDHELRVTGEITYNGHKLNEFVPRKTAAYISQNDVHVGEMTVKETLDFSARCQGVGTRY 203
Query: 272 DLLSELARREKEAGIFPEAELDLFMKATAVKGTESSLITDYTLKILGLDICKDTIVGDEM 331
D LSEL RREKEAGIFPEAELDLFMKATA+KGTESSLITDYTLKILGLDICKDTIVGDEM
Sbjct: 204 DXLSELGRREKEAGIFPEAELDLFMKATALKGTESSLITDYTLKILGLDICKDTIVGDEM 383
Query: 332 NRGVSGGQKKRVTTGEMIVGPTKTLFMDEISTGLDSSTTFQIVKCLQQIVHLTE 385
+RGVSGGQKKRVTTGEMIVGPTKTLFMDEISTGLDSSTTFQIVKCLQQIVHLTE
Sbjct: 384 HRGVSGGQKKRVTTGEMIVGPTKTLFMDEISTGLDSSTTFQIVKCLQQIVHLTE 545
>BP055095
Length = 538
Score = 233 bits (593), Expect = 6e-62
Identities = 108/145 (74%), Positives = 129/145 (88%)
Frame = +3
Query: 303 GTESSLITDYTLKILGLDICKDTIVGDEMNRGVSGGQKKRVTTGEMIVGPTKTLFMDEIS 362
G S L+TDY LK+LGLDIC D ++GDEM RG+SGGQKKRVTTGEM+VGP K LFMDEIS
Sbjct: 6 GQRSQLVTDYALKVLGLDICADVMIGDEMRRGISGGQKKRVTTGEMLVGPAKALFMDEIS 185
Query: 363 TGLDSSTTFQIVKCLQQIVHLTEGTILMSLLQPAPETFDLFDDIILISEGQVVYQGPREH 422
TGLDSSTTFQI K ++Q+VH+ + T+++SLLQPAPETF+LFDDIIL+SEGQ+VYQGPRE+
Sbjct: 186 TGLDSSTTFQICKFMRQMVHIMDVTMVISLLQPAPETFELFDDIILLSEGQIVYQGPREN 365
Query: 423 IVEFFESCGFRCPERKGTADFLQEV 447
++EFFE GF+CPERKG ADFLQEV
Sbjct: 366 VLEFFEYMGFKCPERKGAADFLQEV 440
>AV410506
Length = 433
Score = 167 bits (423), Expect = 3e-42
Identities = 82/92 (89%), Positives = 86/92 (93%)
Frame = +1
Query: 1 MDSNLSRSISRSLSRSSWKMEEVFASGRYSRRTSQVDEDEEALKWAAIEKLPTYDRLRTS 60
MD+NLSRSIS+SLSRSSWKMEEVFASGRYSRR S VDEDEEALKWAAIEKLPTYDRLRTS
Sbjct: 157 MDTNLSRSISKSLSRSSWKMEEVFASGRYSRRASHVDEDEEALKWAAIEKLPTYDRLRTS 336
Query: 61 IMQTFTEGDQPQPGNRQQHKEVDVTKLDMNER 92
I+QT EGDQPQ GNR QHKEVDVTKLDMN+R
Sbjct: 337 IIQTIAEGDQPQGGNRMQHKEVDVTKLDMNDR 432
>AV406931
Length = 438
Score = 122 bits (306), Expect = 1e-28
Identities = 56/76 (73%), Positives = 67/76 (87%)
Frame = +3
Query: 372 QIVKCLQQIVHLTEGTILMSLLQPAPETFDLFDDIILISEGQVVYQGPREHIVEFFESCG 431
QIV L+Q VH+ GT ++SLLQPAPET+DLFDDIILIS+GQVVY GPRE++++FFES G
Sbjct: 3 QIVSSLRQYVHILNGTAVISLLQPAPETYDLFDDIILISDGQVVYHGPREYVLDFFESMG 182
Query: 432 FRCPERKGTADFLQEV 447
F+CPERKG ADFLQEV
Sbjct: 183 FKCPERKGAADFLQEV 230
>BP075566
Length = 253
Score = 66.6 bits (161), Expect(2) = 2e-18
Identities = 32/55 (58%), Positives = 43/55 (78%), Gaps = 1/55 (1%)
Frame = -1
Query: 281 EKEAGIFPEAELDLFM-KATAVKGTESSLITDYTLKILGLDICKDTIVGDEMNRG 334
+++ GIF + F + T VKGTE+SLITDYTL +LGLDICKDT++GD+M+RG
Sbjct: 247 KRKTGIFTRTQNLTFS*RLTXVKGTETSLITDYTLTLLGLDICKDTLLGDDMHRG 83
Score = 42.4 bits (98), Expect(2) = 2e-18
Identities = 20/27 (74%), Positives = 23/27 (85%)
Frame = -2
Query: 335 VSGGQKKRVTTGEMIVGPTKTLFMDEI 361
+SGGQKKRVTTGEMIVGPT+ L +I
Sbjct: 81 LSGGQKKRVTTGEMIVGPTQHLLWMDI 1
>AW720269
Length = 524
Score = 80.5 bits (197), Expect = 5e-16
Identities = 58/197 (29%), Positives = 96/197 (48%), Gaps = 1/197 (0%)
Frame = +3
Query: 170 TTKRTKLTILKNASGIVKPSRMALLLGPPSSGKTTLLLALAGKLDSELRVQGDITYNGHR 229
T K + ILK+ S + + S + ++GP +GK+TLL +AG++ I+ N H
Sbjct: 30 TQKPKPVNILKSVSFVARSSEIVAVVGPSGTGKSTLLRIIAGRVKDRDFDPKTISINDHP 209
Query: 230 LNEFVP-RKTSAYISQNDVHVGEMTVKETLDFSARCQGVGTRYDLLSELARREKEAGIFP 288
+ RK +++Q D + +TVKETL FSA+ + L E+ ++E +
Sbjct: 210 MTSPAQLRKICGFVAQEDNLLPLLTVKETLLFSAKFR--------LKEMTPNDREMRV-- 359
Query: 289 EAELDLFMKATAVKGTESSLITDYTLKILGLDICKDTIVGDEMNRGVSGGQKKRVTTGEM 348
+ ++ LGL D+ VGDE NRG+SGG++KRV+ G
Sbjct: 360 ----------------------ESLMQELGLFHVSDSFVGDEENRGISGGERKRVSIGVD 473
Query: 349 IVGPTKTLFMDEISTGL 365
++ L +DE ++GL
Sbjct: 474 MIHNPPILLLDEPTSGL 524
>AV771159
Length = 486
Score = 59.3 bits (142), Expect(2) = 5e-15
Identities = 32/102 (31%), Positives = 61/102 (59%), Gaps = 1/102 (0%)
Frame = -3
Query: 347 EMIVGPTKTLFMDEISTGLDSSTTFQIVKCLQQIVHLTEG-TILMSLLQPAPETFDLFDD 405
++++ P+ L +DE ++GLDS+T +I+ + ++ T G T++ ++ QP+ + +FD
Sbjct: 343 KLLIHPS-LLLLDEPTSGLDSTTALRILNTIPRLA--TGGRTVVTTIHQPSSRLYYMFDK 173
Query: 406 IILISEGQVVYQGPREHIVEFFESCGFRCPERKGTADFLQEV 447
++L+SEG +Y GP +E+F S GF AD L ++
Sbjct: 172 VVLLSEGCPIYYGPASTALEYFSSVGFSTCVTVNPADLLLDL 47
Score = 38.1 bits (87), Expect(2) = 5e-15
Identities = 15/38 (39%), Positives = 26/38 (67%)
Frame = -2
Query: 317 LGLDICKDTIVGDEMNRGVSGGQKKRVTTGEMIVGPTK 354
LGL C+ +++G + RG+SG +K+RV+ G+ P+K
Sbjct: 437 LGLSGCRSSMIGGPLLRGISGAEKRRVSIGQETAHPSK 324
>AV416268
Length = 424
Score = 72.8 bits (177), Expect = 1e-13
Identities = 39/108 (36%), Positives = 67/108 (61%), Gaps = 2/108 (1%)
Frame = +3
Query: 311 DYTLKILGLDICKDTIVGDEMNRGVSGGQKKRVTTGEMIVGPTKTLFMDEISTGLDSSTT 370
D+T+K +GL +T +G ++GVSGGQK+RV+ I+ + LF+DE ++GLDS+ +
Sbjct: 96 DFTIKEMGLQDAINTRIGGWGSKGVSGGQKRRVSICIEILTHPRLLFLDEPTSGLDSAAS 275
Query: 371 FQIVKCLQQIVHL--TEGTILMSLLQPAPETFDLFDDIILISEGQVVY 416
+ ++ + + + TI+ S+ QP+ E F LF + L+S G+ VY
Sbjct: 276 YHVISRISSLNKKDGIQRTIVASIHQPSNEIFQLFHSLCLLSSGKTVY 419
>BP029778
Length = 438
Score = 66.2 bits (160), Expect = 1e-11
Identities = 34/89 (38%), Positives = 54/89 (60%)
Frame = -2
Query: 174 TKLTILKNASGIVKPSRMALLLGPPSSGKTTLLLALAGKLDSELRVQGDITYNGHRLNEF 233
TKL +L N SG+ P + L+G +GKTTL+ LAG+ ++GDI +G+ +
Sbjct: 290 TKLKLLSNVSGVFAPGVLTALMGSSGAGKTTLMDVLAGRKTGGY-IEGDIKISGYPKVQH 114
Query: 234 VPRKTSAYISQNDVHVGEMTVKETLDFSA 262
+ + Y+ QND+H ++TV+E+L FSA
Sbjct: 113 TFARVAGYVEQNDIHSPQVTVEESLLFSA 27
>TC18647 similar to UP|Q9ASR9 (Q9ASR9) At2g01320/F10A8.20, partial (17%)
Length = 430
Score = 61.6 bits (148), Expect = 3e-10
Identities = 35/90 (38%), Positives = 52/90 (56%), Gaps = 2/90 (2%)
Frame = +1
Query: 178 ILKNASGIVKPSRMALLLGPPSSGKTTLLLALAGKL--DSELRVQGDITYNGHRLNEFVP 235
+LKN SG KP R+ ++GP SGKTTLL LAG+L L + G + +NG ++
Sbjct: 106 LLKNVSGEAKPGRLLAIMGPSGSGKTTLLNVLAGQLAASPRLHLSGLLEFNGKPGSKNAY 285
Query: 236 RKTSAYISQNDVHVGEMTVKETLDFSARCQ 265
+ AY+ Q D+ ++TV+ETL + Q
Sbjct: 286 K--FAYVRQEDLFFSQLTVRETLSLATELQ 369
>BG662257
Length = 336
Score = 39.7 bits (91), Expect = 0.001
Identities = 23/59 (38%), Positives = 34/59 (56%), Gaps = 2/59 (3%)
Frame = +1
Query: 324 DTIVGDEMNRGV--SGGQKKRVTTGEMIVGPTKTLFMDEISTGLDSSTTFQIVKCLQQI 380
DT+VGD RG+ SGGQK+RV IV K L +DE ++ LD+ + + L ++
Sbjct: 109 DTVVGD---RGIQLSGGQKQRVAIARAIVKSPKILLLDEATSALDAESEKVVQDALDRV 276
>TC11425 similar to UP|Q9M1Q9 (Q9M1Q9) P-glycoprotein-like proetin, partial
(9%)
Length = 712
Score = 39.7 bits (91), Expect = 0.001
Identities = 28/101 (27%), Positives = 48/101 (46%)
Frame = +2
Query: 324 DTIVGDEMNRGVSGGQKKRVTTGEMIVGPTKTLFMDEISTGLDSSTTFQIVKCLQQIVHL 383
DT+VG E +SGGQK+RV I+ L +DE ++ LD+ + + L +++
Sbjct: 77 DTVVG-ERGTLLSGGQKQRVAIARAIIKTPNILLLDEATSALDAESERVVQDALDKVMVN 253
Query: 384 TEGTILMSLLQPAPETFDLFDDIILISEGQVVYQGPREHIV 424
I+ L T D I ++ G +V +G E ++
Sbjct: 254 RTTVIVAHRL----STIKNADVITVLKNGVIVEKGRHETLI 364
>CB827630
Length = 405
Score = 36.6 bits (83), Expect = 0.009
Identities = 20/59 (33%), Positives = 32/59 (53%)
Frame = +2
Query: 327 VGDEMNRGVSGGQKKRVTTGEMIVGPTKTLFMDEISTGLDSSTTFQIVKCLQQIVHLTE 385
V D+ SGG K+R+ ++G + ++MDE STGLD ++ K L +V L +
Sbjct: 113 VADKQAGKYSGGMKRRLIVAISLIGDPRVVYMDEPSTGLDPASR----KSLWNVVKLAK 277
>CB829290
Length = 546
Score = 35.4 bits (80), Expect = 0.019
Identities = 22/79 (27%), Positives = 45/79 (56%)
Frame = -3
Query: 336 SGGQKKRVTTGEMIVGPTKTLFMDEISTGLDSSTTFQIVKCLQQIVHLTEGTILMSLLQP 395
S GQ++ V G +++ +K L +DE + +D++T I + L+Q H +E T++ ++
Sbjct: 496 SMGQRQLVCLGRVLLKKSKVLVLDEATASVDTATDNLIQQTLRQ--HFSESTVI-TIAHR 326
Query: 396 APETFDLFDDIILISEGQV 414
D D ++L+S+G +
Sbjct: 325 ITSVID-SDMVLLLSQGLI 272
>BP055390
Length = 488
Score = 35.4 bits (80), Expect = 0.019
Identities = 25/88 (28%), Positives = 41/88 (46%), Gaps = 2/88 (2%)
Frame = +3
Query: 333 RGV--SGGQKKRVTTGEMIVGPTKTLFMDEISTGLDSSTTFQIVKCLQQIVHLTEGTILM 390
RGV SGGQK+R+ I+ L +DE ++ LD+ + + + L+ +H ++
Sbjct: 219 RGVQLSGGQKQRIAIARAILKNPPILLLDEATSALDTESEKLVQEALETAMHGRTVILIA 398
Query: 391 SLLQPAPETFDLFDDIILISEGQVVYQG 418
L D I ++ GQVV G
Sbjct: 399 HRLSTVVNA----DVIAVVENGQVVETG 470
>TC18707 homologue to PIR|T02901|T02901 MSP1 protein homolog T13J8.110 -
Arabidopsis thaliana {Arabidopsis thaliana;}, partial
(29%)
Length = 771
Score = 34.7 bits (78), Expect = 0.033
Identities = 15/29 (51%), Positives = 20/29 (68%)
Frame = +3
Query: 184 GIVKPSRMALLLGPPSSGKTTLLLALAGK 212
G++KP R LL GPP +GKT L A+A +
Sbjct: 288 GLLKPCRGILLFGPPGTGKTMLAXAIANE 374
>TC10924 homologue to UP|Q8H9D4 (Q8H9D4) 26S proteasome AAA-ATPase subunit
RPT4a, partial (49%)
Length = 648
Score = 33.9 bits (76), Expect = 0.056
Identities = 14/30 (46%), Positives = 20/30 (66%)
Frame = +2
Query: 186 VKPSRMALLLGPPSSGKTTLLLALAGKLDS 215
+KP + LL GPP +GKT L A+A +D+
Sbjct: 551 IKPPKGVLLYGPPGTGKTLLARAIASNIDA 640
>TC10604 similar to AAQ89671 (AAQ89671) At4g33460, partial (36%)
Length = 498
Score = 33.1 bits (74), Expect = 0.095
Identities = 17/48 (35%), Positives = 26/48 (53%)
Frame = +2
Query: 166 FGFNTTKRTKLTILKNASGIVKPSRMALLLGPPSSGKTTLLLALAGKL 213
F F + + +L++ S + + +LLGP GK+TLL LAG L
Sbjct: 164 FSFTARQTNDVRVLRDCSLRIPSGQFWMLLGPNGCGKSTLLKILAGLL 307
>TC17532 similar to UP|Q9LZ98 (Q9LZ98) ABC transporter-like protein
(At5g02270), partial (48%)
Length = 476
Score = 32.7 bits (73), Expect = 0.12
Identities = 28/116 (24%), Positives = 52/116 (44%), Gaps = 4/116 (3%)
Frame = +2
Query: 314 LKILGLDICKDTIVGDEMNRGVSGGQKKRVTTGEMIVGPTKTLFMDEISTGLDSSTTFQI 373
+K+LG+D+ VS GQ++RV ++ P K L +DEI+ LD +
Sbjct: 128 IKVLGIDLSWRL-------HKVSDGQRRRVQICMGLLKPFKVLLLDEITVDLDVLARADL 286
Query: 374 VKCLQQIVHLTEGTILMSLLQPAPETFDLFDD----IILISEGQVVYQGPREHIVE 425
+ L++ TI+ A FD +D ++ ++ G++ P + + E
Sbjct: 287 LSFLKKECDERGATIIY-----ATHIFDGLEDWPTHMVYVAHGKLQLAMPMDKVKE 439
>TC15722 similar to UP|Q9FMU0 (Q9FMU0) Similarity to ABC transporter,
partial (37%)
Length = 624
Score = 30.8 bits (68), Expect = 0.47
Identities = 16/52 (30%), Positives = 26/52 (49%)
Frame = +3
Query: 329 DEMNRGVSGGQKKRVTTGEMIVGPTKTLFMDEISTGLDSSTTFQIVKCLQQI 380
D+ + +SGG K+R+ +V L +DE GLD +VK L+ +
Sbjct: 51 DKNPQSLSGGYKRRLALAIQLVQVPDLLILDEPLAGLDWKARADVVKLLKHL 206
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.317 0.135 0.377
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 5,937,163
Number of Sequences: 28460
Number of extensions: 66830
Number of successful extensions: 401
Number of sequences better than 10.0: 66
Number of HSP's better than 10.0 without gapping: 393
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 396
length of query: 447
length of database: 4,897,600
effective HSP length: 93
effective length of query: 354
effective length of database: 2,250,820
effective search space: 796790280
effective search space used: 796790280
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 57 (26.6 bits)
Medicago: description of AC123976.3


