

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC123898.1 - phase: 0 /pseudo
(177 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
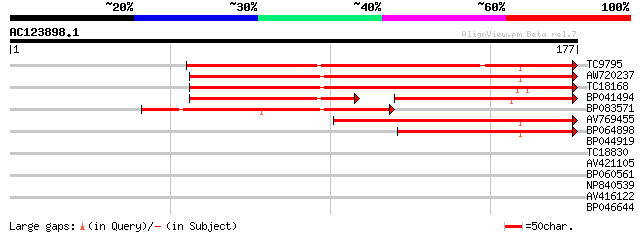
TC9795 homologue to UP|Q9ATB8 (Q9ATB8) BiP-isoform D (Fragment),... 116 3e-27
AW720237 113 2e-26
TC18168 homologue to UP|Q8GSN4 (Q8GSN4) Non-cell-autonomous heat... 103 2e-23
BP041494 55 2e-16
BP083571 69 4e-13
AV769455 61 1e-10
BP064898 48 1e-06
BP044919 35 0.006
TC18830 weakly similar to UP|O40637 (O40637) Protease, partial (5%) 28 1.2
AV421105 27 2.0
BP060561 26 3.4
NP840539 hypothetical protein [Lotus corniculatus var. japonicus] 25 5.8
AV416122 25 7.6
BP046644 25 7.6
>TC9795 homologue to UP|Q9ATB8 (Q9ATB8) BiP-isoform D (Fragment), partial
(63%)
Length = 981
Score = 116 bits (290), Expect = 3e-27
Identities = 62/127 (48%), Positives = 83/127 (64%), Gaps = 5/127 (3%)
Frame = +1
Query: 56 VIGIDLGTTNSCVSVMEGKNPKVVENSEGARTTPSVVAFTQKGELLVGTPAKRQAVTNPE 115
VIGIDLGTT SCV V + +++ N +G R TPS VAF GE L+G AK Q NPE
Sbjct: 151 VIGIDLGTTYSCVGVYRNGHVEIIANDQGNRITPSWVAFND-GERLIGEAAKNQFAVNPE 327
Query: 116 NTISGAKRLIGRRFDDPQTQKEMKMVPYKIVKAPNGDAWVEAK-----GQQYSPSQIGAF 170
TI KRLIGR+F+D + Q++MK+VPYKIV +G +++ K + +SP ++ A
Sbjct: 328 RTIFDVKRLIGRKFEDKEVQRDMKLVPYKIVN-KDGKPYIQVKIKDGETKVFSPEEVSAM 504
Query: 171 VLTKMKE 177
+LTKMKE
Sbjct: 505 ILTKMKE 525
>AW720237
Length = 459
Score = 113 bits (283), Expect = 2e-26
Identities = 59/125 (47%), Positives = 79/125 (63%), Gaps = 4/125 (3%)
Frame = +3
Query: 57 IGIDLGTTNSCVSVMEGKNPKVVENSEGARTTPSVVAFTQKGELLVGTPAKRQAVTNPEN 116
IGIDLGTT SCV V +++ N +G RTTPS V FT+ E L+G AK Q NP N
Sbjct: 45 IGIDLGTTYSCVGVWMNDRVEIIANDQGNRTTPSYVGFTES-ERLIGDAAKNQVARNPIN 221
Query: 117 TISGAKRLIGRRFDDPQTQKEMKMVPYKIVKAPNGDAWVEAK----GQQYSPSQIGAFVL 172
T+ AKRLIGR+FDDP QK++K+ P+K+ P+G + K +++ Q+ + VL
Sbjct: 222 TVFDAKRLIGRKFDDPIVQKDIKLWPFKVEAGPDGKPLIAVKYKGETKKFHAEQVSSMVL 401
Query: 173 TKMKE 177
TKMKE
Sbjct: 402 TKMKE 416
>TC18168 homologue to UP|Q8GSN4 (Q8GSN4) Non-cell-autonomous heat shock
cognate protein 70, partial (23%)
Length = 508
Score = 103 bits (256), Expect = 2e-23
Identities = 57/125 (45%), Positives = 76/125 (60%), Gaps = 4/125 (3%)
Frame = +2
Query: 57 IGIDLGTTNSCVSVMEGKNPKVVENSEGARTTPSVVAFTQKGELLVGTPAKRQAVTNPEN 116
IGIDLGTT SCV V + +++ N +G RTTPS VAFT E L+G AK Q NP N
Sbjct: 86 IGIDLGTTYSCVGVWQHDRVEIIANDQGNRTTPSYVAFTDT-ERLIGDAAKNQVAMNPIN 262
Query: 117 TISGAKRLIGRRFDDPQTQKEMKMVPYKIVKAPNGDAWVEA--KGQ--QYSPSQIGAFVL 172
T+ AKRLIGRR D Q +MK+ P+K+ P ++ KG+ Q++ +I + VL
Sbjct: 263 TVFDAKRLIGRRVSDASVQSDMKLWPFKVNSGPAEKPMIQVSYKGEDKQFAAEEISSMVL 442
Query: 173 TKMKE 177
KM+E
Sbjct: 443 MKMRE 457
>BP041494
Length = 572
Score = 55.5 bits (132), Expect(2) = 2e-16
Identities = 29/53 (54%), Positives = 35/53 (65%)
Frame = +1
Query: 57 IGIDLGTTNSCVSVMEGKNPKVVENSEGARTTPSVVAFTQKGELLVGTPAKRQ 109
IGIDLGTT SCV V + +++ N +G RTTPS VAFT E L+G AK Q
Sbjct: 88 IGIDLGTTYSCVGVWQNDRVEIIPNDQGNRTTPSYVAFTD-FERLIGDAAKNQ 243
Score = 45.1 bits (105), Expect(2) = 2e-16
Identities = 21/61 (34%), Positives = 37/61 (60%), Gaps = 4/61 (6%)
Frame = +3
Query: 121 AKRLIGRRFDDPQTQKEMKMVPYKIVKAPNGDAWV----EAKGQQYSPSQIGAFVLTKMK 176
AKRLIGR+F D + +MK+ P+K++ P + + + +Q++ +I + VL KM+
Sbjct: 246 AKRLIGRKFSDDSVESDMKLWPFKVISGPVDRPMIVVNYKNEEKQFTAEEISSMVLIKMR 425
Query: 177 E 177
E
Sbjct: 426 E 428
>BP083571
Length = 423
Score = 69.3 bits (168), Expect = 4e-13
Identities = 39/81 (48%), Positives = 53/81 (65%), Gaps = 2/81 (2%)
Frame = +1
Query: 42 SLSRPFSSRAAGNDVIGIDLGTTNSCVSVMEGKNPK--VVENSEGARTTPSVVAFTQKGE 99
SL+ +++ G IGIDLGTT SCV+V + +N + ++ N +G RTTPS VAFT +
Sbjct: 130 SLAERMATKYEG-PAIGIDLGTTYSCVAVWQEQNDRAEIIHNEQGNRTTPSFVAFTDT-Q 303
Query: 100 LLVGTPAKRQAVTNPENTISG 120
L+G AK QA TNP NT+ G
Sbjct: 304 RLIGDAAKNQAATNPTNTVLG 366
>AV769455
Length = 556
Score = 61.2 bits (147), Expect = 1e-10
Identities = 27/80 (33%), Positives = 51/80 (63%), Gaps = 4/80 (5%)
Frame = +1
Query: 102 VGTPAKRQAVTNPENTISGAKRLIGRRFDDPQTQKEMKMVPYKIVKAPNGDAWVEAK--- 158
+GT + NP+N+IS KRLIGR+F DP+ Q+++K +P+ + + P+G + A+
Sbjct: 4 LGTAGASSTMMNPKNSISQLKRLIGRQFADPELQRDLKSLPFAVTEGPDGFPLIHARYLG 183
Query: 159 -GQQYSPSQIGAFVLTKMKE 177
+ ++P+Q+ +L+ +KE
Sbjct: 184 ETRTFTPTQVFGMMLSNLKE 243
>BP064898
Length = 542
Score = 47.8 bits (112), Expect = 1e-06
Identities = 21/60 (35%), Positives = 40/60 (66%), Gaps = 4/60 (6%)
Frame = +1
Query: 122 KRLIGRRFDDPQTQKEMKMVPYKIVKAPNGDAWVEAK----GQQYSPSQIGAFVLTKMKE 177
KRLIG +F DPQ Q+++K +P+ + + P+G + A+ + ++P+Q+ A +L+ +KE
Sbjct: 337 KRLIGIQFSDPQLQRDLKSLPFLVTEGPDGYPLIHARYLGETRTFTPTQVFAMMLSNLKE 516
>BP044919
Length = 519
Score = 35.4 bits (80), Expect = 0.006
Identities = 18/56 (32%), Positives = 34/56 (60%), Gaps = 4/56 (7%)
Frame = +3
Query: 126 GRRFDDPQTQKEMKMVPYKIVKAPNGDAW--VEAKGQQYS--PSQIGAFVLTKMKE 177
GR++ D Q+++K+ P+K++ + V+ KG++ P +I + VL+KMKE
Sbjct: 27 GRKYSDSVIQEDVKLWPFKVIAGTDDKPMIVVKHKGEEKKLVPEEISSMVLSKMKE 194
>TC18830 weakly similar to UP|O40637 (O40637) Protease, partial (5%)
Length = 850
Score = 27.7 bits (60), Expect = 1.2
Identities = 14/26 (53%), Positives = 17/26 (64%)
Frame = +2
Query: 48 SSRAAGNDVIGIDLGTTNSCVSVMEG 73
SS +AG IG+D+G NSCVS G
Sbjct: 335 SSASAGT--IGVDIGLHNSCVSAHSG 406
>AV421105
Length = 415
Score = 26.9 bits (58), Expect = 2.0
Identities = 27/108 (25%), Positives = 47/108 (43%), Gaps = 5/108 (4%)
Frame = +2
Query: 68 VSVMEGKNPKVVENSEGARTTPSVVAFTQKGEL-----LVGTPAKRQAVTNPENTISGAK 122
V E K +++++S G + +VA GE V A+ + +TN + A
Sbjct: 5 VPYRESKLTRILQDSLGGTSRALMVACLNPGEYQESVHTVSLAARSRHITN---VVPSAH 175
Query: 123 RLIGRRFDDPQTQKEMKMVPYKIVKAPNGDAWVEAKGQQYSPSQIGAF 170
+ +T K M + K+ AW+E+KG+ S ++GAF
Sbjct: 176 K--------EETPKVMVDMEAKL------RAWLESKGKAKSSQKLGAF 277
>BP060561
Length = 390
Score = 26.2 bits (56), Expect = 3.4
Identities = 10/19 (52%), Positives = 12/19 (62%)
Frame = +3
Query: 147 KAPNGDAWVEAKGQQYSPS 165
+ P WVEAKG +Y PS
Sbjct: 102 ETPTYGGWVEAKGPRYCPS 158
>NP840539 hypothetical protein [Lotus corniculatus var. japonicus]
Length = 3654
Score = 25.4 bits (54), Expect = 5.8
Identities = 18/61 (29%), Positives = 25/61 (40%)
Frame = +1
Query: 55 DVIGIDLGTTNSCVSVMEGKNPKVVENSEGARTTPSVVAFTQKGELLVGTPAKRQAVTNP 114
D +G+D G + G P S+G T + E +VG K AVT+P
Sbjct: 523 DFVGLDFGDKKKTRGLPPGLVPVTDSFSDGDLT---------EVEFIVGDATKFDAVTDP 675
Query: 115 E 115
E
Sbjct: 676 E 678
>AV416122
Length = 414
Score = 25.0 bits (53), Expect = 7.6
Identities = 13/37 (35%), Positives = 23/37 (62%)
Frame = -3
Query: 16 AASSTVTAFRSLTGNTKPAYVGHNWSSLSRPFSSRAA 52
++SS+ ++ SL+ ++ + NW+SLS SS AA
Sbjct: 364 SSSSSSSSSSSLSSSSSSSSSPSNWTSLSGSLSSLAA 254
>BP046644
Length = 513
Score = 25.0 bits (53), Expect = 7.6
Identities = 16/47 (34%), Positives = 22/47 (46%)
Frame = +3
Query: 4 ATALLRSLRRRDAASSTVTAFRSLTGNTKPAYVGHNWSSLSRPFSSR 50
A+ RS D+A T F +L +P NW ++S P SSR
Sbjct: 252 ASLFSRSSTSLDSAGLGAT-FSALLEPIRPFGAEDNWDAISTPISSR 389
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.312 0.127 0.357
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 2,441,046
Number of Sequences: 28460
Number of extensions: 27822
Number of successful extensions: 97
Number of sequences better than 10.0: 28
Number of HSP's better than 10.0 without gapping: 90
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 90
length of query: 177
length of database: 4,897,600
effective HSP length: 84
effective length of query: 93
effective length of database: 2,506,960
effective search space: 233147280
effective search space used: 233147280
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (21.9 bits)
S2: 52 (24.6 bits)
Medicago: description of AC123898.1


