

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
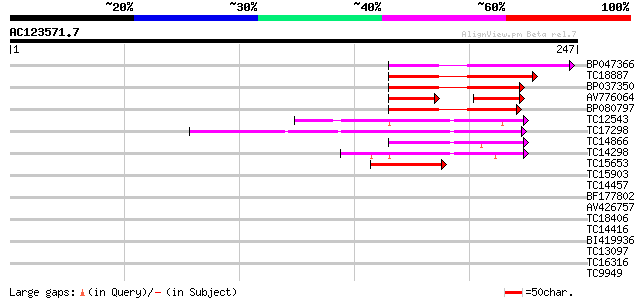
Query= AC123571.7 - phase: 0
(247 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
BP047366 68 1e-12
TC18887 similar to UP|Q94KA6 (Q94KA6) bZIP transcription factor ... 65 1e-11
BP037350 61 2e-10
AV776064 43 1e-07
BP080797 51 2e-07
TC12543 similar to UP|Q9AT29 (Q9AT29) bZIP transcription factor,... 42 1e-04
TC17298 similar to UP|Q9AT29 (Q9AT29) bZIP transcription factor,... 40 5e-04
TC14866 similar to UP|Q8L5W2 (Q8L5W2) bZIP transcription factor ... 40 5e-04
TC14298 homologue to UP|Q8L5W2 (Q8L5W2) bZIP transcription facto... 39 6e-04
TC15653 similar to UP|Q941L5 (Q941L5) bZIP transcription factor ... 39 6e-04
TC15903 similar to UP|Q40724 (Q40724) Transcriptional activator ... 38 0.001
TC14457 similar to UP|Q8GTZ1 (Q8GTZ1) bZIP transcription factor,... 37 0.003
BF177802 36 0.005
AV426757 33 0.035
TC18406 similar to UP|O23726 (O23726) B-Zip DNA binding protein,... 31 0.17
TC14416 similar to UP|Q8SMH8 (Q8SMH8) RNA-binding protein, parti... 30 0.29
BI419936 30 0.50
TC13097 similar to UP|Q9ZVB8 (Q9ZVB8) At2g41330 protein (At2g413... 29 0.86
TC16316 homologue to UP|Q9LTA2 (Q9LTA2) Similarity to AT-hook DN... 28 1.9
TC9949 weakly similar to UP|O82047 (O82047) PRIB5 protein (Fragm... 28 1.9
>BP047366
Length = 535
Score = 68.2 bits (165), Expect = 1e-12
Identities = 45/81 (55%), Positives = 47/81 (57%)
Frame = -3
Query: 166 ERRNKRMIKNRESAARSRARKQEKITSFLFSKFSAYTNELEQKVQLLQEENARLRRQQQE 225
ERR KRMIKNRESAARSRARKQ AYT ELE KV L+EEN RLRRQ +
Sbjct: 488 ERRQKRMIKNRESAARSRARKQ------------AYTQELEIKVSRLEEENERLRRQHEI 345
Query: 226 LWEAESGGQQKKKSSLYRTSS 246
S K L RTSS
Sbjct: 344 EQVLPSVPPPDPKHQLRRTSS 282
>TC18887 similar to UP|Q94KA6 (Q94KA6) bZIP transcription factor 6, partial
(64%)
Length = 944
Score = 64.7 bits (156), Expect = 1e-11
Identities = 36/65 (55%), Positives = 45/65 (68%)
Frame = +3
Query: 166 ERRNKRMIKNRESAARSRARKQEKITSFLFSKFSAYTNELEQKVQLLQEENARLRRQQQE 225
ERR +RMIKNRESAARSRARKQ AYT ELEQ+V L+EEN L+++Q+E
Sbjct: 780 ERRQRRMIKNRESAARSRARKQ------------AYTMELEQEVAKLKEENEELQKKQEE 923
Query: 226 LWEAE 230
+ E +
Sbjct: 924 IMEMQ 938
>BP037350
Length = 495
Score = 61.2 bits (147), Expect = 2e-10
Identities = 35/59 (59%), Positives = 40/59 (67%)
Frame = -2
Query: 166 ERRNKRMIKNRESAARSRARKQEKITSFLFSKFSAYTNELEQKVQLLQEENARLRRQQQ 224
ERR KRMIKNRESAARSRARK AYTNEL KV L+EEN RLR++++
Sbjct: 377 ERRQKRMIKNRESAARSRARKH------------AYTNEL*NKVSRLEEENERLRKRKE 237
>AV776064
Length = 418
Score = 43.1 bits (100), Expect(2) = 1e-07
Identities = 21/22 (95%), Positives = 21/22 (95%)
Frame = -1
Query: 166 ERRNKRMIKNRESAARSRARKQ 187
ERR KRMIKNRESAARSRARKQ
Sbjct: 355 ERRQKRMIKNRESAARSRARKQ 290
Score = 28.1 bits (61), Expect(2) = 1e-07
Identities = 12/22 (54%), Positives = 17/22 (76%)
Frame = -2
Query: 203 NELEQKVQLLQEENARLRRQQQ 224
NELE KV L+EEN R R++++
Sbjct: 279 NELENKVSRLEEENERPRKRKE 214
>BP080797
Length = 498
Score = 51.2 bits (121), Expect = 2e-07
Identities = 29/58 (50%), Positives = 38/58 (65%)
Frame = -2
Query: 166 ERRNKRMIKNRESAARSRARKQEKITSFLFSKFSAYTNELEQKVQLLQEENARLRRQQ 223
ERR +R IKNRESAARSRARKQ AY NEL KV L+++N +L++++
Sbjct: 449 ERRLRRKIKNRESAARSRARKQ------------AYHNELVSKVTRLEQDNIKLKKEK 312
>TC12543 similar to UP|Q9AT29 (Q9AT29) bZIP transcription factor, partial
(52%)
Length = 510
Score = 41.6 bits (96), Expect = 1e-04
Identities = 39/134 (29%), Positives = 55/134 (40%), Gaps = 32/134 (23%)
Frame = +3
Query: 125 SNVSPCFVNASPCDQNVGVPASSSFTCFGKRFGEAPDISP-------------------- 164
SN P N + N+ + F F + + PD SP
Sbjct: 54 SNPPPYPANYTMIQNNI---PTFQFQRFSNQLFQVPDFSPQSSCISSNSTSDEADEQQQG 224
Query: 165 --GERRNKRMIKNRESAARSRARKQEKITSFLFSKFSAYTNELEQKVQLLQ--------- 213
ER+++RMI NRESA RSR RKQ + L+S+ NE Q + L
Sbjct: 225 LINERKHRRMISNRESARRSRMRKQRHLDE-LWSQVVWLRNENHQLLDKLSHASESHDQV 401
Query: 214 -EENARLRRQQQEL 226
+ENA+L+ + EL
Sbjct: 402 VQENAQLKEEAIEL 443
>TC17298 similar to UP|Q9AT29 (Q9AT29) bZIP transcription factor, partial
(39%)
Length = 484
Score = 39.7 bits (91), Expect = 5e-04
Identities = 43/147 (29%), Positives = 66/147 (44%)
Frame = +1
Query: 79 STTPPPLVTALSLNSRPDFLYDPLIRHNKHNNSQLLLQQQQHNIGVSNVSPCFVNASPCD 138
ST +T LS D +I++N QL Q G N F + SP +
Sbjct: 49 STMQASEITGLSYFLPSDPSPFSMIQNNNIPTFQLQKFSNQF-FGYQNNLQAFQDFSPQN 225
Query: 139 QNVGVPASSSFTCFGKRFGEAPDISPGERRNKRMIKNRESAARSRARKQEKITSFLFSKF 198
+ + +S+S + + + + ER+++RMI NRESA RSR RKQ+ + L+S
Sbjct: 226 SSSCI-SSNSTSDEAEDQQQQQSLIINERKHRRMISNRESARRSRMRKQKHLDE-LWSMV 399
Query: 199 SAYTNELEQKVQLLQEENARLRRQQQE 225
NE Q ++ L + R QE
Sbjct: 400 VCLRNENHQLLEKLNHLSESHDRVVQE 480
>TC14866 similar to UP|Q8L5W2 (Q8L5W2) bZIP transcription factor ATB2,
partial (60%)
Length = 948
Score = 39.7 bits (91), Expect = 5e-04
Identities = 28/71 (39%), Positives = 37/71 (51%), Gaps = 10/71 (14%)
Frame = +3
Query: 166 ERRNKRMIKNRESAARSRARKQEKITSFLFSKFSAYTNE----------LEQKVQLLQEE 215
+R+NKR NRESA RSR RKQ + L S+ + T E Q+ Q ++ E
Sbjct: 54 QRKNKRKQSNRESARRSRMRKQSHLED-LTSQVTQLTKENGEILTNINITSQQYQNVETE 230
Query: 216 NARLRRQQQEL 226
N+ LR Q EL
Sbjct: 231 NSILRAQMGEL 263
>TC14298 homologue to UP|Q8L5W2 (Q8L5W2) bZIP transcription factor ATB2,
partial (62%)
Length = 1585
Score = 39.3 bits (90), Expect = 6e-04
Identities = 35/98 (35%), Positives = 47/98 (47%), Gaps = 16/98 (16%)
Frame = +3
Query: 145 ASSSFTCFGKRF----GEAPDISP--GERRNKRMIKNRESAARSRARKQEKITSFLFSKF 198
ASSS T G G D+ +R+ KRMI NRESA RSR RKQ+ + L S+
Sbjct: 615 ASSSGTSSGSSLLQNSGSEEDLQAVMDQRKRKRMISNRESARRSRMRKQKHLDD-LASQV 791
Query: 199 SAYTNELEQKVQ----------LLQEENARLRRQQQEL 226
+ E +Q + ++ EN+ LR Q EL
Sbjct: 792 TQLRKENQQIITSVNITTQQYLSVEAENSVLRAQMGEL 905
>TC15653 similar to UP|Q941L5 (Q941L5) bZIP transcription factor BZI-4,
partial (25%)
Length = 511
Score = 39.3 bits (90), Expect = 6e-04
Identities = 17/33 (51%), Positives = 25/33 (75%)
Frame = +2
Query: 158 EAPDISPGERRNKRMIKNRESAARSRARKQEKI 190
E D++ +R+ KRM+ NRESA RSR RKQ+++
Sbjct: 377 EGGDLAIDDRKRKRMVSNRESARRSRMRKQKQL 475
>TC15903 similar to UP|Q40724 (Q40724) Transcriptional activator protein
(RITA-1 protein), partial (21%)
Length = 603
Score = 38.1 bits (87), Expect = 0.001
Identities = 24/56 (42%), Positives = 32/56 (56%)
Frame = +1
Query: 167 RRNKRMIKNRESAARSRARKQEKITSFLFSKFSAYTNELEQKVQLLQEENARLRRQ 222
+R +R + NRESA RSR RKQ A+ ELE +V+ L+ ENA L +Q
Sbjct: 466 KRLRRKVSNRESARRSRRRKQ------------AHLAELETQVEKLKLENATLYKQ 597
>TC14457 similar to UP|Q8GTZ1 (Q8GTZ1) bZIP transcription factor, partial
(50%)
Length = 645
Score = 37.0 bits (84), Expect = 0.003
Identities = 25/70 (35%), Positives = 36/70 (50%), Gaps = 9/70 (12%)
Frame = +1
Query: 166 ERRNKRMIKNRESAARSRARKQEKI--TSFLFSKFSAYTNELEQKVQL-------LQEEN 216
+++ KRM NRESA RSR RKQE + S + N++ + + ++ EN
Sbjct: 16 QKKRKRMQSNRESARRSRMRKQEHLEGMSAQVEQLKKENNQISTNIGVTTQMYLNVEAEN 195
Query: 217 ARLRRQQQEL 226
A LR Q EL
Sbjct: 196 AILRVQMAEL 225
>BF177802
Length = 481
Score = 36.2 bits (82), Expect = 0.005
Identities = 16/22 (72%), Positives = 20/22 (90%)
Frame = +1
Query: 166 ERRNKRMIKNRESAARSRARKQ 187
++R KR++KNRESA RSRARKQ
Sbjct: 406 DQRRKRILKNRESALRSRARKQ 471
Score = 25.4 bits (54), Expect = 9.5
Identities = 16/66 (24%), Positives = 31/66 (46%)
Frame = +1
Query: 3 EVWKDINLSSLNDQNTRPMIMSTRNSTFGGGVILQDFLTRPLTLDPTKSLDYSSNNSSSS 62
EVWKD+ LSSL D + S +S + Q + ++ + + + +++SS
Sbjct: 190 EVWKDLKLSSLCDSSMEMEASSPWHSLCPASSVFQKQSHQHHSMLSSFNTSFDASSSSGE 369
Query: 63 VASDQN 68
+ Q+
Sbjct: 370 KIAQQD 387
>AV426757
Length = 415
Score = 33.5 bits (75), Expect = 0.035
Identities = 21/53 (39%), Positives = 29/53 (54%)
Frame = +1
Query: 167 RRNKRMIKNRESAARSRARKQEKITSFLFSKFSAYTNELEQKVQLLQEENARL 219
+R KR++ NR+SA RSR RK + Y +ELE+ V LQ E + L
Sbjct: 46 KRVKRILANRQSAQRSRVRKLQ------------YISELERSVTTLQTEVSAL 168
>TC18406 similar to UP|O23726 (O23726) B-Zip DNA binding protein, partial
(14%)
Length = 491
Score = 31.2 bits (69), Expect = 0.17
Identities = 18/46 (39%), Positives = 26/46 (56%)
Frame = +3
Query: 167 RRNKRMIKNRESAARSRARKQEKITSFLFSKFSAYTNELEQKVQLL 212
+R KR++ NR+SAARS+ RK + Y ELE+K+ L
Sbjct: 390 KRAKRILANRQSAARSKERK------------ACYVLELERKIHSL 491
>TC14416 similar to UP|Q8SMH8 (Q8SMH8) RNA-binding protein, partial (28%)
Length = 695
Score = 30.4 bits (67), Expect = 0.29
Identities = 19/55 (34%), Positives = 27/55 (48%), Gaps = 2/55 (3%)
Frame = +2
Query: 37 QDFLTRPLTLDPTKSLDYSSNNSSSSVASDQNNNNA--SFYCPISTTPPPLVTAL 89
Q + +L P S N SSS + D +N+N+ S P+S +PP TAL
Sbjct: 131 QQWRNHVFSLHPPHSSTTKPNTSSSQLPPDPSNSNSHPSTLHPLSPSPPQPTTAL 295
>BI419936
Length = 503
Score = 29.6 bits (65), Expect = 0.50
Identities = 22/72 (30%), Positives = 34/72 (46%), Gaps = 1/72 (1%)
Frame = +3
Query: 51 SLDYSSNNSSSSVASDQNNNNASFYCPISTTPPPLVTALSLNSRPDFLYD-PLIRHNKHN 109
S D S + S A +NN+A+F P S+ LVT + N + +F P +K
Sbjct: 90 SKDLSPWSGLHSRAESSDNNSANFEAPRSSVRTNLVTRIEGNKKWEFANSIPANFKSKGK 269
Query: 110 NSQLLLQQQQHN 121
+S+LL + N
Sbjct: 270 SSELLSTESPEN 305
>TC13097 similar to UP|Q9ZVB8 (Q9ZVB8) At2g41330 protein
(At2g41330/F13H10.12), partial (7%)
Length = 660
Score = 28.9 bits (63), Expect = 0.86
Identities = 15/59 (25%), Positives = 31/59 (52%)
Frame = -2
Query: 47 DPTKSLDYSSNNSSSSVASDQNNNNASFYCPISTTPPPLVTALSLNSRPDFLYDPLIRH 105
+P+ S +S++SSSS ++ +++++ C S+ PP +L + PL+ H
Sbjct: 347 NPSPSASLASSSSSSSFSASSSSSSSKSQCSNSSPKPPRALSLPM---------PLVHH 198
>TC16316 homologue to UP|Q9LTA2 (Q9LTA2) Similarity to AT-hook DNA-binding
protein, partial (7%)
Length = 488
Score = 27.7 bits (60), Expect = 1.9
Identities = 15/48 (31%), Positives = 26/48 (53%), Gaps = 2/48 (4%)
Frame = +3
Query: 49 TKSLDYSSNNSSSSVASDQNNNNASFYCPIST--TPPPLVTALSLNSR 94
T S +++NN++++ + ++N S S TPPP TA L S+
Sbjct: 273 TCSQSFTTNNNNTTTTNHHTHHNTSSLTTPSNSPTPPPWTTAAHLRSQ 416
>TC9949 weakly similar to UP|O82047 (O82047) PRIB5 protein (Fragment),
partial (37%)
Length = 587
Score = 27.7 bits (60), Expect = 1.9
Identities = 18/57 (31%), Positives = 27/57 (46%), Gaps = 4/57 (7%)
Frame = -2
Query: 97 FLYDPLI--RHNKHNNSQLLLQQQQ--HNIGVSNVSPCFVNASPCDQNVGVPASSSF 149
F+Y ++ R H+NS + +QQ I + + CFV SPC +N P F
Sbjct: 247 FIYCSILFERSKSHSNSFSMCNKQQVLSRIFICKLLDCFVIPSPCFKNWLFPLGQLF 77
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.313 0.127 0.361
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 4,404,746
Number of Sequences: 28460
Number of extensions: 63827
Number of successful extensions: 450
Number of sequences better than 10.0: 68
Number of HSP's better than 10.0 without gapping: 435
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 440
length of query: 247
length of database: 4,897,600
effective HSP length: 88
effective length of query: 159
effective length of database: 2,393,120
effective search space: 380506080
effective search space used: 380506080
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (21.9 bits)
S2: 54 (25.4 bits)
Medicago: description of AC123571.7


