

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
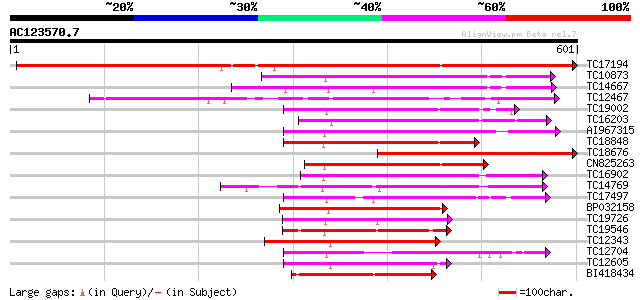
Query= AC123570.7 + phase: 0
(601 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
TC17194 UP|CAE02590 (CAE02590) Nod-factor receptor 1b, complete 667 0.0
TC10873 similar to UP|Q9LDZ5 (Q9LDZ5) F2D10.13 (F5M15.3), partia... 211 3e-55
TC14667 homologue to UP|Q84P43 (Q84P43) Protein kinase Pti1, com... 196 9e-51
TC12467 UP|CAE02597 (CAE02597) Nod-factor receptor 5, complete 193 6e-50
TC19002 weakly similar to UP|Q9XJ10 (Q9XJ10) ESTs C22458(C62866)... 184 5e-47
TC16203 UP|Q8GRU6 (Q8GRU6) LRR receptor-like kinase (Hypernodula... 182 1e-46
AI967315 182 2e-46
TC18848 weakly similar to UP|Q9SSQ6 (Q9SSQ6) F6D8.24 protein, pa... 173 6e-44
TC18676 similar to UP|Q8RYS1 (Q8RYS1) Protein kinase-like, parti... 167 6e-42
CN825263 166 1e-41
TC16902 homologue to UP|Q8VZH5 (Q8VZH5) Receptor protein kinase-... 162 2e-40
TC14769 UP|Q8LKX1 (Q8LKX1) Receptor-like kinase SYMRK, complete 160 4e-40
TC17497 UP|CAE45595 (CAE45595) S-receptor kinase-like protein 2,... 157 5e-39
BP032158 147 4e-36
TC19726 similar to UP|Q9XIS3 (Q9XIS3) Lectin-like protein kinase... 146 1e-35
TC19546 similar to UP|Q40543 (Q40543) Protein-serine/threonine k... 141 3e-34
TC12343 similar to UP|CDK9_SCHPO (Q96WV9) Serine/threonine prote... 141 3e-34
TC12704 UP|CAE45596 (CAE45596) S-receptor kinase-like protein 3,... 136 1e-32
TC12605 similar to UP|AAR99871 (AAR99871) Strubbelig receptor fa... 132 2e-31
BI418434 131 3e-31
>TC17194 UP|CAE02590 (CAE02590) Nod-factor receptor 1b, complete
Length = 2193
Score = 667 bits (1721), Expect = 0.0
Identities = 352/620 (56%), Positives = 441/620 (70%), Gaps = 26/620 (4%)
Frame = +1
Query: 8 LSFLFMLLASKSFIAESKCSKTCNIALASYYLQDDTN-LTYVSNIMQSNLVTKPEDIVSY 66
L F +LL F ES C K C++ALASYY+ L ++ MQS +V+ + I SY
Sbjct: 136 LLFFILLLGHVCFHVESNCLKGCDLALASYYILPGVFILQNITTFMQSEIVSSNDAITSY 315
Query: 67 NTDTITNKDFVQSFTRVNVPFPCDCIHDEFLGHIFQYQVATKDTYLSVASNNYSNLTTSE 126
N D I N +QSF R+N+PFPCDCI EFLGH+F+Y + DTY ++A+ Y+NLTT +
Sbjct: 316 NKDKILNDINIQSFQRLNIPFPCDCIGGEFLGHVFEYSASKGDTYETIANLYYANLTTVD 495
Query: 127 WLQNFNSYPSNDIPDTGTLNVTVNCSCGNSDVSKDYGLFITYPLRPEDSLELISNKTEID 186
L+ FNSY +IP +NVTVNCSCGNS VSKDYGLFITYP+RP D+L+ I+N++ +D
Sbjct: 496 LLKRFNSYDPKNIPVNAKVNVTVNCSCGNSQVSKDYGLFITYPIRPGDTLQDIANQSSLD 675
Query: 187 AELLQKYNPGVNFSQGSGLVYIPGKDQNRNYVPFHTR-------------------LILL 227
A L+Q +NP VNFS+ SG+ +IPG+ +N YVP + R L+LL
Sbjct: 676 AGLIQSFNPSVNFSKDSGIAFIPGRYKNGVYVPLYHRTAGLASGAAVGISIAGTFVLLLL 855
Query: 228 SFCIYVTYYRKKKIRKQEFLSEESSAIFGQVKNDEVSGNATYGTSDSASPAN-----MIG 282
+FC+YV Y +KK+ K + ++ S A+ Q N S +A Y TS S+ P +
Sbjct: 856 AFCMYVRY-QKKEEEKAKLPTDISMALSTQDGN--ASSSAEYETSGSSGPGTASATGLTS 1026
Query: 283 IRVEKSGEFSYEELANATNNFNMANKIGQGGFGEVYYAELNGEKAAIKKMDMKATKEFLA 342
I V KS EFSY+ELA ATNNF++ NKIGQGGFG VYYAEL G+K AIKKMD++A+ EFL
Sbjct: 1027IMVAKSMEFSYQELAKATNNFSLDNKIGQGGFGAVYYAELRGKKTAIKKMDVQASTEFLC 1206
Query: 343 ELKVLTRVHHVNLVRLIGYCVEGSLFLVYEYIDNGNLGQHLRSSDGEPLSWSIRVKIALD 402
ELKVLT VHH+NLVRLIGYCVEGSLFLVYE+IDNGNLGQ+L S EPL WS RV+IALD
Sbjct: 1207ELKVLTHVHHLNLVRLIGYCVEGSLFLVYEHIDNGNLGQYLHGSGKEPLPWSSRVQIALD 1386
Query: 403 SARGLEYIHEHTVPTYIHRDIKSENILLDKNFCAKVADFGLTKLIDAGISSVPTVNMAGT 462
+ARGLEYIHEHTVP YIHRD+KS NIL+DKN KVADFGLTKLI+ G S++ T + GT
Sbjct: 1387AARGLEYIHEHTVPVYIHRDVKSANILIDKNLRGKVADFGLTKLIEVGNSTLQT-RLVGT 1563
Query: 463 FGYMPPEYA-YGSVSSKIDVYAFGVVLYELISAKAAVIMGEDSGADLKGLVVLFEEVFDQ 521
FGYMPPEYA YG +S KIDVYAFGVVL+ELISAK AV+ + A+ KGLV LFEE ++
Sbjct: 1564FGYMPPEYAQYGDISPKIDVYAFGVVLFELISAKNAVLKTGELVAESKGLVALFEEALNK 1743
Query: 522 PHPIEGLKKLVDPRLGDNYPIDHVFKMAQLAKVCTNSDPQQRPNMSSVVVALTTLTSTTE 581
P + L+KLVDPRLG+NYPID V K+AQL + CT +P RP+M S+VVAL TL+S TE
Sbjct: 1744SDPCDALRKLVDPRLGENYPIDSVLKIAQLGRACTRDNPLLRPSMRSLVVALMTLSSLTE 1923
Query: 582 DWDITSIFKNPNLVNLMSGR 601
D D S +++ L+NL+S R
Sbjct: 1924DCDDESSYESQTLINLLSVR 1983
>TC10873 similar to UP|Q9LDZ5 (Q9LDZ5) F2D10.13 (F5M15.3), partial (77%)
Length = 1478
Score = 211 bits (537), Expect = 3e-55
Identities = 128/319 (40%), Positives = 181/319 (56%), Gaps = 8/319 (2%)
Frame = +1
Query: 268 TYGTSDSASPANMIGIRVEKSGEFSYEELANATNNFNMANKIGQGGFGEVYYAELN-GEK 326
T G S +N + F + ELA+AT NF AN IG+GGFG+VY L GE
Sbjct: 346 TKGKGKGVSSSNGSSNGKTAAASFGFRELADATRNFKEANLIGEGGFGKVYKGRLTTGEA 525
Query: 327 AAIKKMD---MKATKEFLAELKVLTRVHHVNLVRLIGYCVEGSL-FLVYEYIDNGNLGQH 382
A+K++ + +EF+ E+ +L+ +HH NLVRLIGYC +G LVYEY+ G+L H
Sbjct: 526 VAVKQLSHDGRQGFQEFVMEVLMLSLLHHTNLVRLIGYCTDGDQRLLVYEYMPMGSLEDH 705
Query: 383 L--RSSDGEPLSWSIRVKIALDSARGLEYIHEHTVPTYIHRDIKSENILLDKNFCAKVAD 440
L S D EPL+WS R+K+A+ +ARGLEY+H P I+RD+KS NILLD F K++D
Sbjct: 706 LFELSHDKEPLNWSTRMKVAVGAARGLEYLHCTADPPVIYRDLKSANILLDNEFNPKLSD 885
Query: 441 FGLTKLIDAGISSVPTVNMAGTFGYMPPEYAY-GSVSSKIDVYAFGVVLYELISAKAAVI 499
FGL KL G ++ + + GT+GY PEYA G ++ K D+Y+FGVVL EL++ + A+
Sbjct: 886 FGLAKLGPVGDNTHVSTRVMGTYGYCAPEYAMSGKLTLKSDIYSFGVVLLELLTGRRAID 1065
Query: 500 MGEDSGADLKGLVVLFEEVFDQPHPIEGLKKLVDPRLGDNYPIDHVFKMAQLAKVCTNSD 559
G + LV F +VDP L +P + + + +C
Sbjct: 1066TSRRPGE--QNLVSWARPYFSDR---RRFGHMVDPLLQGRFPSRCLHQAIAITAMCLQEQ 1230
Query: 560 PQQRPNMSSVVVALTTLTS 578
P+ RP ++ +VVAL L S
Sbjct: 1231PKFRPLITDIVVALEYLAS 1287
>TC14667 homologue to UP|Q84P43 (Q84P43) Protein kinase Pti1, complete
Length = 1591
Score = 196 bits (498), Expect = 9e-51
Identities = 130/368 (35%), Positives = 194/368 (52%), Gaps = 24/368 (6%)
Frame = +2
Query: 236 YRKKKIRKQEFL--SEESSAIFGQVKNDEVSGNATYGTSDSASPANMIGIRVEKSGE--- 290
Y +K+ R + +L + + + +ND + YG +S ++ E
Sbjct: 128 YLRKRTRMRRWLCCTCQVEEPYPSNENDHLKSPRNYGDGNSKGSKASAPVKHETQKAPPP 307
Query: 291 -----FSYEELANATNNFNMANKIGQGGFGEVYYAELN-GEKAAIKKMDMKA----TKEF 340
S +EL T+NF IG+G +G VYYA LN G A+KK+D+ + EF
Sbjct: 308 IEVPALSLDELKEKTDNFGSKALIGEGSYGRVYYATLNDGNAVAVKKLDVSSEPETNNEF 487
Query: 341 LAELKVLTRVHHVNLVRLIGYCVEGSL-FLVYEYIDNGNLGQHLR-------SSDGEPLS 392
L ++ +++R+ + N V L GYCVEG+L L YE+ G+L L + G L
Sbjct: 488 LTQVSMVSRLKNDNFVELHGYCVEGNLRVLAYEFATMGSLHDILHGRKGVQGAQPGPTLD 667
Query: 393 WSIRVKIALDSARGLEYIHEHTVPTYIHRDIKSENILLDKNFCAKVADFGLTKLIDAGIS 452
W RV+IA+D+ARGLEY+HE P IHRDI+S N+L+ +++ AK+ADF L+ +
Sbjct: 668 WIQRVRIAVDAARGLEYLHEKVQPAIIHRDIRSSNVLIFEDYKAKIADFNLSNQAPDMAA 847
Query: 453 SVPTVNMAGTFGYMPPEYAY-GSVSSKIDVYAFGVVLYELISAKAAVIMGEDSGADLKGL 511
+ + + GTFGY PEYA G ++ K DVY+FGVVL EL++ + V G + L
Sbjct: 848 RLHSTRVLGTFGYHAPEYAMTGQLTQKSDVYSFGVVLLELLTGRKPVDHTMPRGQ--QSL 1021
Query: 512 VVLFEEVFDQPHPIEGLKKLVDPRLGDNYPIDHVFKMAQLAKVCTNSDPQQRPNMSSVVV 571
V + + +K+ VDP+L YP V K+A +A +C + + RPNMS VV
Sbjct: 1022VTWATPRLSE----DKVKQCVDPKLKGEYPPKGVAKLAAVAALCVQYEAEFRPNMSIVVK 1189
Query: 572 ALTTLTST 579
AL L T
Sbjct: 1190ALQPLLKT 1213
>TC12467 UP|CAE02597 (CAE02597) Nod-factor receptor 5, complete
Length = 2292
Score = 193 bits (491), Expect = 6e-50
Identities = 164/531 (30%), Positives = 255/531 (47%), Gaps = 33/531 (6%)
Frame = +3
Query: 85 VPFPCDCIHDEFLGHIFQYQVATKDTYLSVASNNYSNLTTSEWLQNFN-SYPSNDIPDTG 143
VP C C + + YQ+ D+Y VA+ Y NLT +Q N +P+
Sbjct: 429 VPVTCGCAGNHSSANT-SYQIQLGDSYDFVATTLYENLTNWNIVQASNPGVNPYLLPERV 605
Query: 144 TLNVTVNCSC-GNSDVSKDYGLFITYPLRPEDSLELISNKTEID-AELLQKYNPGVNFSQ 201
+ + C C + ++K ITY +P D++ L+S K A++L + G +F+
Sbjct: 606 KVVFPLFCRCPSKNQLNKGIQYLITYVWKPNDNVSLVSAKFGASPADILTENRYGQDFTA 785
Query: 202 GSGL-VYIP--------GKDQNRNYVPFHTRLIL------------LSFCIYVTYYRKKK 240
+ L + IP N H +IL L+ + Y R+KK
Sbjct: 786 ATNLPILIPVTQLPELTQPSSNGRKSSIHLLVILGITLGCTLLTAVLTGTLVYVYCRRKK 965
Query: 241 IRKQEFLSEESSAIFGQVKNDEVSGNATYGTSDSASPANMIGIRVEKSGEFSYEELANAT 300
+ S E++ + +SG + Y V K + +E+ AT
Sbjct: 966 ALNRTASSAETA-------DKLLSGVSGY---------------VSKPNVYEIDEIMEAT 1079
Query: 301 NNFNMANKIGQGGFGEVYYAELNGEKAAIKKMDMKATKEFLAELKVLTRVHHVNLVRLIG 360
+F+ K+G+ VY A + G A+KK+ E ELK+L +V+H NLV+L+G
Sbjct: 1080KDFSDECKVGE----SVYKANIEGRVVAVKKIKEGGANE---ELKILQKVNHGNLVKLMG 1238
Query: 361 YC--VEGSLFLVYEYIDNGNLGQHLRS-SDGEP--LSWSIRVKIALDSARGLEYIHEHTV 415
+G+ FLVYEY +NG+L + L S S G P L+WS R+ IA+D A GL+Y+HEHT
Sbjct: 1239VSSGYDGNCFLVYEYAENGSLAEWLFSKSSGTPNSLTWSQRISIAVDVAVGLQYMHEHTY 1418
Query: 416 PTYIHRDIKSENILLDKNFCAKVADFGLTKLIDAGISSVPTVNMAGTFGYMPPEYAYGSV 475
P IHRDI + NILLD NF AK+A+F + + T MP
Sbjct: 1419PRIIHRDITTSNILLDSNFKAKIANFAMAR--------------TSTNPMMP-------- 1532
Query: 476 SSKIDVYAFGVVLYELISAKAAVIMGEDSGADLKGLVVLFE---EVFD-QPHPIEGLKKL 531
KIDV+AFGV+L EL++ + A+ E+ +V+L++ E+FD + + E ++K
Sbjct: 1533--KIDVFAFGVLLIELLTGRKAMTTKENG-----EVVMLWKDMWEIFDIEENREERIRKW 1691
Query: 532 VDPRLGDNYPIDHVFKMAQLAKVCTNSDPQQRPNMSSVVVALTTLTSTTED 582
+DP L Y ID+ +A LA CT RP+M+ +V++L+ LT + +
Sbjct: 1692MDPNLESFYHIDNALSLASLAVNCTADKSLSRPSMAEIVLSLSFLTQQSSN 1844
>TC19002 weakly similar to UP|Q9XJ10 (Q9XJ10) ESTs C22458(C62866), partial
(48%)
Length = 780
Score = 184 bits (466), Expect = 5e-47
Identities = 110/261 (42%), Positives = 152/261 (58%), Gaps = 11/261 (4%)
Frame = +1
Query: 291 FSYEELANATNNFNMANKIGQGGFGEVYYAEL-NGEKAAIKKMDM---KATKEFLAELKV 346
FS +EL +ATNNFN NK+G+GGFG VY+ +L +G + A+K++ + KA EF E+++
Sbjct: 28 FSLKELHSATNNFNYDNKLGEGGFGSVYWGQLWDGSQIAVKRLKVWSNKADMEFAVEVEI 207
Query: 347 LTRVHHVNLVRLIGYCVEGS-LFLVYEYIDNGNLGQHLRSSDGEP--LSWSIRVKIALDS 403
L RV H NL+ L GYC EG +VY+Y+ N +L HL L W+ R+ IA+ S
Sbjct: 208 LARVRHKNLLSLRGYCAEGQERLIVYDYMPNLSLLSHLHGQHSSECLLDWNRRMNIAIGS 387
Query: 404 ARGLEYIHEHTVPTYIHRDIKSENILLDKNFCAKVADFGLTKLIDAGISSVPTVNMAGTF 463
A G+ Y+H P IHRDIK+ N+LLD +F A+VADFG KLI G + V T + GT
Sbjct: 388 AEGIVYLHHQATPHIIHRDIKASNVLLDSDFQARVADFGFAKLIPDGATHV-TTRVKGTL 564
Query: 464 GYMPPEYA-YGSVSSKIDVYAFGVVLYELISAKAAVIMGEDSGADLKGLVVLFEEVFDQP 522
GY+ PEYA G + DV++FG++L EL S K + E + +K + D
Sbjct: 565 GYLAPEYAMLGKANECCDVFSFGILLLELASGKKPL---EKLSSTVK------RSINDWA 717
Query: 523 HPIEGLKK---LVDPRLGDNY 540
P+ KK DPRL Y
Sbjct: 718 LPLACAKKFTEFADPRLNGEY 780
>TC16203 UP|Q8GRU6 (Q8GRU6) LRR receptor-like kinase (Hypernodulation aberrant
root formation protein), complete
Length = 3308
Score = 182 bits (462), Expect = 1e-46
Identities = 116/276 (42%), Positives = 158/276 (57%), Gaps = 8/276 (2%)
Frame = +2
Query: 307 NKIGQGGFGEVYYAEL-NGEKAAIKKMDMKATKE----FLAELKVLTRVHHVNLVRLIGY 361
N IG+GG G VY + NG AIK++ + + F AE++ L ++ H N++RL+GY
Sbjct: 2204 NIIGKGGAGIVYRGSMPNGTDVAIKRLVGQGSGRNDYGFRAEIETLGKIRHRNIMRLLGY 2383
Query: 362 CV-EGSLFLVYEYIDNGNLGQHLRSSDGEPLSWSIRVKIALDSARGLEYIHEHTVPTYIH 420
+ + L+YEY+ NG+LG+ L + G L W +R KIA+++ARGL Y+H P IH
Sbjct: 2384 VSNKDTNLLLYEYMPNGSLGEWLHGAKGGHLRWEMRYKIAVEAARGLCYMHHDCSPLIIH 2563
Query: 421 RDIKSENILLDKNFCAKVADFGLTKLIDAGISSVPTVNMAGTFGYMPPEYAYG-SVSSKI 479
RD+KS NILLD +F A VADFGL K + +S ++AG++GY+ PEYAY V K
Sbjct: 2564 RDVKSNNILLDADFEAHVADFGLAKFLYDPGASQSMSSIAGSYGYIAPEYAYTLKVDEKS 2743
Query: 480 DVYAFGVVLYELISAKAAVIMGE-DSGADLKGLVVLFEEVFDQPHPIEGLKKLVDPRLGD 538
DVY+FGVVL ELI + V GE G D+ G V QP + +VDPRL
Sbjct: 2744 DVYSFGVVLLELIIGRKPV--GEFGDGVDIVGWVNKTMSELSQPSDTALVLAVVDPRL-S 2914
Query: 539 NYPIDHVFKMAQLAKVCTNSDPQQRPNMSSVVVALT 574
YP+ V M +A +C RP M VV LT
Sbjct: 2915 GYPLTSVIHMFNIAMMCVKEMGPARPTMREVVHMLT 3022
>AI967315
Length = 1308
Score = 182 bits (461), Expect = 2e-46
Identities = 109/301 (36%), Positives = 164/301 (54%), Gaps = 8/301 (2%)
Frame = +1
Query: 291 FSYEELANATNNFNMANKIGQGGFGEVYYAEL-NGEKAAIKKM-----DMKATKEFLAEL 344
FSYEEL +ATN F+ N +G+GG+ EVY L +G++ A+K++ D + KEFL E+
Sbjct: 112 FSYEELFHATNGFSSENMVGKGGYAEVYKGRLESGDEIAVKRLTRTCRDERKEKEFLTEI 291
Query: 345 KVLTRVHHVNLVRLIGYCVEGSLFLVYEYIDNGNLGQHLRSSDGEPLSWSIRVKIALDSA 404
+ V H N++ L+G C++ L+LV+E G++ + PL W R KI L +A
Sbjct: 292 GTIGHVCHSNVMPLLGCCIDNGLYLVFELSTVGSVASLIHDEKMAPLDWKTRYKIVLGTA 471
Query: 405 RGLEYIHEHTVPTYIHRDIKSENILLDKNFCAKVADFGLTKLIDAGISSVPTVNMAGTFG 464
RGL Y+H+ IHRDIK+ NILL ++F +++DFGL K + + + + GTFG
Sbjct: 472 RGLHYLHKGCQRRIIHRDIKASNILLTEDFEPQISDFGLAKWLPSQWTHHSIAPIEGTFG 651
Query: 465 YMPPE-YAYGSVSSKIDVYAFGVVLYELISAKAAVIMGEDS-GADLKGLVVLFEEVFDQP 522
++ PE Y +G V K DV+AFGV L E+IS + V S K ++ +E
Sbjct: 652 HLAPEYYMHGVVDEKTDVFAFGVFLLEVISGRKPVDGSHQSLHTWAKPILSKWE------ 813
Query: 523 HPIEGLKKLVDPRLGDNYPIDHVFKMAQLAKVCTNSDPQQRPNMSSVVVALTTLTSTTED 582
++KLVDPRL Y + ++A A +C + RP MS V+ + E
Sbjct: 814 -----IEKLVDPRLEGCYDVTQFNRVAFAASLCIRASSTWRPTMSEVLEVMEEGEMDKER 978
Query: 583 W 583
W
Sbjct: 979 W 981
>TC18848 weakly similar to UP|Q9SSQ6 (Q9SSQ6) F6D8.24 protein, partial (64%)
Length = 969
Score = 173 bits (439), Expect = 6e-44
Identities = 97/216 (44%), Positives = 136/216 (62%), Gaps = 8/216 (3%)
Frame = +2
Query: 291 FSYEELANATNNFNMANKIGQGGFGEVYYAELN-GEKAAIKK---MDMKATKEFLAELKV 346
F+Y+EL AT F+ NK+G+GGFG VY+ + G + A+KK M+ KA EF E++V
Sbjct: 266 FTYKELHAATGGFSDDNKLGEGGFGSVYWGRTSDGLQIAVKKLKAMNSKAEMEFAVEVEV 445
Query: 347 LTRVHHVNLVRLIGYCV-EGSLFLVYEYIDNGNLGQHLRSSDGEP--LSWSIRVKIALDS 403
L RV H NL+ L GYCV + +VY+Y+ N +L HL L+W R+KIA+ S
Sbjct: 446 LGRVRHKNLLGLRGYCVGDDQRLIVYDYMPNLSLLSHLHGQFAVEVQLNWQKRMKIAIGS 625
Query: 404 ARGLEYIHEHTVPTYIHRDIKSENILLDKNFCAKVADFGLTKLIDAGISSVPTVNMAGTF 463
A G+ Y+H P IHRDIK+ N+LL+ +F VADFG KLI G+S + T + GT
Sbjct: 626 AEGILYLHHEVTPHIIHRDIKASNVLLNSDFEPLVADFGFAKLIPEGVSHM-TTRVKGTL 802
Query: 464 GYMPPEYA-YGSVSSKIDVYAFGVVLYELISAKAAV 498
GY+ PEYA +G VS DVY+FG++L EL++ + +
Sbjct: 803 GYLAPEYAMWGKVSESCDVYSFGILLLELVTGRKPI 910
>TC18676 similar to UP|Q8RYS1 (Q8RYS1) Protein kinase-like, partial (15%)
Length = 814
Score = 167 bits (422), Expect = 6e-42
Identities = 95/217 (43%), Positives = 133/217 (60%), Gaps = 6/217 (2%)
Frame = +3
Query: 391 LSWSIRVKIALDSARGLEYIHEHTVPTYIHRDIKSENILLDKNFCAKVADFGLTKLIDAG 450
LSW RV+IALD+ARGLEYIHEHT Y+H+DI + NILLD +F AK++DFGL KL+
Sbjct: 36 LSWITRVQIALDAARGLEYIHEHTKTRYVHQDINTSNILLDASFRAKISDFGLAKLVSET 215
Query: 451 I-SSVPTVNMAGTFGYMPPEYAYGSV-SSKIDVYAFGVVLYELISAKAAVIMGEDS-GAD 507
I T T+GY+ PEY + +SK DVYAFGVVLYE+IS K A+I + + G +
Sbjct: 216 IEGGTTTTKGVSTYGYLAPEYLSNRIATSKSDVYAFGVVLYEIISGKKAIIQTQGTQGPE 395
Query: 508 LKGLV-VLFEEVFDQPHPIE--GLKKLVDPRLGDNYPIDHVFKMAQLAKVCTNSDPQQRP 564
+ L ++ E + P + ++ VDP + D Y D V +MA LAK C DP RP
Sbjct: 396 RRSLASIMLEVLRTVPDSLSTPSIRNHVDPIMKDLYSHDCVLQMAMLAKQCVEEDPILRP 575
Query: 565 NMSSVVVALTTLTSTTEDWDITSIFKNPNLVNLMSGR 601
+M VV++L+ + ++ +W+ T K+ L+ GR
Sbjct: 576 DMKQVVLSLSQIHLSSFEWEATLAGKSQVFSGLIQGR 686
>CN825263
Length = 663
Score = 166 bits (420), Expect = 1e-41
Identities = 95/204 (46%), Positives = 133/204 (64%), Gaps = 9/204 (4%)
Frame = +1
Query: 313 GFGEVYYAELN-GEKAAIKKM---DMKATKEFLAELKVLTRVHHVNLVRLIGYCVEGSL- 367
GFG VY LN G A+K + D + +EFLAE+++L+R+HH NLV+LIG C+E
Sbjct: 1 GFGLVYKGILNDGRDVAVKILKRDDQRGGREFLAEVEMLSRLHHRNLVKLIGICIEKQTR 180
Query: 368 FLVYEYIDNGNLGQHLRSSDGE--PLSWSIRVKIALDSARGLEYIHEHTVPTYIHRDIKS 425
L+YE + NG++ HL +D E PL W+ R+KIAL +ARGL Y+HE + P IHRD KS
Sbjct: 181 CLIYELVPNGSVESHLHGADKETGPLDWNARMKIALGAARGLAYLHEDSNPCVIHRDFKS 360
Query: 426 ENILLDKNFCAKVADFGLTK-LIDAGISSVPTVNMAGTFGYMPPEYAY-GSVSSKIDVYA 483
NILL+ +F KV+DFGL + +D G + T ++ GTFGY+ PEYA G + K DVY+
Sbjct: 361 SNILLECDFTPKVSDFGLARTALDEGNKHIST-HVMGTFGYLAPEYAMTGHLLVKSDVYS 537
Query: 484 FGVVLYELISAKAAVIMGEDSGAD 507
+GVVL EL++ V + + G +
Sbjct: 538 YGVVLLELLTGTKPVDLSQPPGQE 609
>TC16902 homologue to UP|Q8VZH5 (Q8VZH5) Receptor protein kinase-like
protein, partial (46%)
Length = 941
Score = 162 bits (409), Expect = 2e-40
Identities = 98/268 (36%), Positives = 154/268 (56%), Gaps = 6/268 (2%)
Frame = +3
Query: 309 IGQGGFGEVYYAELNG-EKAAIKK---MDMKATKEFLAELKVLTRVHHVNLVRLIGYCVE 364
+G GGFG+VYY E++G K AIK+ + + EF E+++L+++ H +LV LIGYC E
Sbjct: 15 LGVGGFGKVYYGEVDGGTKVAIKRGNPLSEQGVHEFQTEIEMLSKLRHRHLVSLIGYCEE 194
Query: 365 GS-LFLVYEYIDNGNLGQHLRSSDGEPLSWSIRVKIALDSARGLEYIHEHTVPTYIHRDI 423
+ + LVY+++ G L +HL + PL W R++I + +ARGL Y+H T IHRD+
Sbjct: 195 NTEMILVYDHMAYGTLREHLYKTQKPPLPWKQRLEICIGAARGLHYLHTGAKYTIIHRDV 374
Query: 424 KSENILLDKNFCAKVADFGLTKLIDAGISSVPTVNMAGTFGYMPPEY-AYGSVSSKIDVY 482
K+ NILLD+ + AKV+DFGL+K ++ + + G+FGY+ PEY ++ K DVY
Sbjct: 375 KTTNILLDEKWVAKVSDFGLSKTGPTLDNTHVSTVVKGSFGYLDPEYFRRQQLTDKSDVY 554
Query: 483 AFGVVLYELISAKAAVIMGEDSGADLKGLVVLFEEVFDQPHPIEGLKKLVDPRLGDNYPI 542
+FGVVL+E++ A+ A+ L V E + L +++DP L
Sbjct: 555 SFGVVLFEILCARPAL------NPSLAKEQVSLAEWASHCYNKGILDQILDPYLKGKIAP 716
Query: 543 DHVFKMAQLAKVCTNSDPQQRPNMSSVV 570
+ K A+ A C + +RP+M V+
Sbjct: 717 ECFKKFAETAMKCVSDQGIERPSMGDVL 800
>TC14769 UP|Q8LKX1 (Q8LKX1) Receptor-like kinase SYMRK, complete
Length = 3495
Score = 160 bits (406), Expect = 4e-40
Identities = 124/365 (33%), Positives = 183/365 (49%), Gaps = 18/365 (4%)
Frame = +3
Query: 224 LILLSF-CIYVTYYRKKKIRKQEFLSE----ESSAIFGQVKNDEVSGNATYGTSDSASPA 278
LI L+F ++V YR+K I + F + E++ IF D+
Sbjct: 1971 LITLAFGVLFVCRYRQKLIPWEGFAGKKYPMETNIIFSLPSKDDFF-------------- 2108
Query: 279 NMIGIRVEKSGEFSYEELANATNNFNMANKIGQGGFGEVYYAELN-GEKAAIK---KMDM 334
I+ F+ E + AT + IG+GGFG VY LN G++ A+K
Sbjct: 2109 ----IKSVSIQAFTLEYIEVATERYKTL--IGEGGFGSVYRGTLNDGQEVAVKVRSSTST 2270
Query: 335 KATKEFLAELKVLTRVHHVNLVRLIGYCVEGSL-FLVYEYIDNGNLGQHLRSSDGEP--- 390
+ T+EF EL +L+ + H NLV L+GYC E LVY ++ NG+L L GEP
Sbjct: 2271 QGTREFDNELNLLSAIQHENLVPLLGYCNESDQQILVYPFMSNGSLQDRLY---GEPAKR 2441
Query: 391 --LSWSIRVKIALDSARGLEYIHEHTVPTYIHRDIKSENILLDKNFCAKVADFGLTKLID 448
L W R+ IAL +ARGL Y+H + IHRDIKS NILLD + CAKVADFG +K
Sbjct: 2442 KILDWPTRLSIALGAARGLAYLHTFPGRSVIHRDIKSSNILLDHSMCAKVADFGFSKYAP 2621
Query: 449 AGISSVPTVNMAGTFGYMPPE-YAYGSVSSKIDVYAFGVVLYELISAKAAVIMGEDSGAD 507
S ++ + GT GY+ PE Y +S K DV++FGVVL E++S + + +
Sbjct: 2622 QEGDSYVSLEVRGTAGYLDPEYYKTQQLSEKSDVFSFGVVLLEIVSGREPL--------N 2777
Query: 508 LKGLVVLFEEVFDQPHPIEGLK--KLVDPRLGDNYPIDHVFKMAQLAKVCTNSDPQQRPN 565
+K + V I G K ++VDP + Y + ++++ ++A C RP+
Sbjct: 2778 IKRPRTEWSLVEWATPYIRGSKVDEIVDPGIKGGYHAEAMWRVVEVALQCLEPFSTYRPS 2957
Query: 566 MSSVV 570
M ++V
Sbjct: 2958 MVAIV 2972
>TC17497 UP|CAE45595 (CAE45595) S-receptor kinase-like protein 2, complete
Length = 2946
Score = 157 bits (397), Expect = 5e-39
Identities = 100/292 (34%), Positives = 164/292 (55%), Gaps = 9/292 (3%)
Frame = +1
Query: 291 FSYEELANATNNFNMANKIGQGGFGEVYYAEL-NGEKAAIKKMDM---KATKEFLAELKV 346
F + +++ATN+F+++NK+G+GGFG VY L NG++ A+K++ + +EF E+K+
Sbjct: 1636 FDFSTISSATNHFSLSNKLGEGGFGPVYKGLLANGQEIAVKRLSNTSGQGMEEFKNEIKL 1815
Query: 347 LTRVHHVNLVRLIGYCVEGSLFLVYEYIDNGNLGQHLR----SSDGEPLSWSIRVKIALD 402
+ R+ H NLV+L G V +N + + ++ S+ + + W+ R++I
Sbjct: 1816 IARLQHRNLVKLFGCSVHQD--------ENSHANKKMKILLDSTRSKLVDWNKRLQIIDG 1971
Query: 403 SARGLEYIHEHTVPTYIHRDIKSENILLDKNFCAKVADFGLTKLIDAGISSVPTVNMAGT 462
ARGL Y+H+ + IHRD+K+ NILLD K++DFGL ++ T + GT
Sbjct: 1972 IARGLLYLHQDSRLRIIHRDLKTSNILLDDEMNPKISDFGLARIFIGDQVEARTKRVMGT 2151
Query: 463 FGYMPPEYA-YGSVSSKIDVYAFGVVLYELISAKAAVIMGEDSGADLKGLVVLFEEVFDQ 521
+GYMPPEYA +GS S K DV++FGV++ E+IS K + D L L+ ++ +
Sbjct: 2152 YGYMPPEYAVHGSFSIKSDVFSFGVIVLEIISGK-KIGRFYDPHHHL-NLLSHAWRLWIE 2325
Query: 522 PHPIEGLKKLVDPRLGDNYPIDHVFKMAQLAKVCTNSDPQQRPNMSSVVVAL 573
P+E LVD L D + + +A +C P+ RP+M S+V+ L
Sbjct: 2326 ERPLE----LVDELLDDPVIPTEILRYIHVALLCVQRRPENRPDMLSIVLML 2469
>BP032158
Length = 555
Score = 147 bits (372), Expect = 4e-36
Identities = 80/185 (43%), Positives = 124/185 (66%), Gaps = 7/185 (3%)
Frame = +2
Query: 287 KSGEFSYEELANATNNFNMANKIGQGGFGEVYYAELN-GEKAAIKKMDMKA---TKEFLA 342
K+G FS ++ ATNNF+ ANKIG+GGFG VY L+ G+ A+K++ K+ +EF+
Sbjct: 2 KTGYFSLRQIKAATNNFDPANKIGEGGFGPVYKGVLSEGDVIAVKQLSSKSKQGNREFIN 181
Query: 343 ELKVLTRVHHVNLVRLIGYCVEGS-LFLVYEYIDNGNLGQHLRSSDGEPLS--WSIRVKI 399
E+ +++ + H NLV+L G C+EG+ L LVYEY++N +L + L ++ + L+ W R+KI
Sbjct: 182 EIGMISALQHPNLVKLYGCCIEGNQLLLVYEYMENNSLARALFGNEEQKLNLNWRTRMKI 361
Query: 400 ALDSARGLEYIHEHTVPTYIHRDIKSENILLDKNFCAKVADFGLTKLIDAGISSVPTVNM 459
+ A+GL Y+HE + +HRDIK+ N+LLDK+ AK++DFGL KL + + + T +
Sbjct: 362 CVGIAKGLAYLHEESRLKIVHRDIKATNVLLDKDLKAKISDFGLAKLDEEENTHIST-RI 538
Query: 460 AGTFG 464
AGT G
Sbjct: 539 AGTIG 553
>TC19726 similar to UP|Q9XIS3 (Q9XIS3) Lectin-like protein kinase, partial
(30%)
Length = 728
Score = 146 bits (368), Expect = 1e-35
Identities = 83/191 (43%), Positives = 115/191 (59%), Gaps = 11/191 (5%)
Frame = +3
Query: 290 EFSYEELANATNNFNMANKIGQGGFGEVYYAELNGEK--AAIKKMD---MKATKEFLAEL 344
EF+Y EL ATNNF+ +K+GQGG+G VY L EK A+K MK+T +FL+EL
Sbjct: 129 EFNYVELKKATNNFDEKHKLGQGGYGVVYRGMLPKEKLEVAVKMFSRDKMKSTDDFLSEL 308
Query: 345 KVLTRVHHVNLVRLIGYCVE-GSLFLVYEYIDNGNLGQHLRSSDG----EPLSWSIRVKI 399
++ R+ H +LVRL G+C + G L LVY+Y+ NG+L H+ +G PLSW +R KI
Sbjct: 309 IIINRLRHKHLVRLQGWCHKNGVLLLVYDYMPNGSLDSHIFCEEGGTITTPLSWPLRYKI 488
Query: 400 ALDSARGLEYIHEHTVPTYIHRDIKSENILLDKNFCAKVADFGLTKLID-AGISSVPTVN 458
A L Y+H +HRD+K+ NI+LD F AK+ DFGL + ++ IS
Sbjct: 489 ISGVASALNYLHNEYDQKVVHRDLKASNIMLDSEFNAKLGDFGLARALENEKISYTELEG 668
Query: 459 MAGTFGYMPPE 469
+ GT GY+ PE
Sbjct: 669 VHGTMGYIAPE 701
>TC19546 similar to UP|Q40543 (Q40543) Protein-serine/threonine kinase ,
partial (46%)
Length = 731
Score = 141 bits (355), Expect = 3e-34
Identities = 81/184 (44%), Positives = 113/184 (61%), Gaps = 5/184 (2%)
Frame = +1
Query: 290 EFSYEELANATNNFNMANKIGQGGFGEVYYAEL-NGEKAAIKKM---DMKATKEFLAELK 345
++SY+++ AT NF +GQG FG VY A + GE A+K + + EF E+
Sbjct: 196 KYSYKDIQKATQNFTTI--LGQGSFGTVYKATMPTGEVVAVKVLAPNSKQGEHEFQTEVH 369
Query: 346 VLTRVHHVNLVRLIGYCVE-GSLFLVYEYIDNGNLGQHLRSSDGEPLSWSIRVKIALDSA 404
+L R+HH NLV L+G+CV+ G LVY+++ NG+L L + E LSW R++IA+D +
Sbjct: 370 LLGRLHHRNLVNLVGFCVDKGQRILVYQFMSNGSLANLLYGEEKE-LSWDERLQIAMDIS 546
Query: 405 RGLEYIHEHTVPTYIHRDIKSENILLDKNFCAKVADFGLTKLIDAGISSVPTVNMAGTFG 464
G+EY+HE VP IHRD+KS NILLD + A VADFGL+K I + GT+G
Sbjct: 547 HGIEYLHEGAVPPVIHRDLKSANILLDDSMRAMVADFGLSK---EEIFDGRNSGLKGTYG 717
Query: 465 YMPP 468
YM P
Sbjct: 718 YMDP 729
>TC12343 similar to UP|CDK9_SCHPO (Q96WV9) Serine/threonine protein kinase
Cdk9 (Cyclin-dependent kinase Cdk9) , partial (7%)
Length = 723
Score = 141 bits (355), Expect = 3e-34
Identities = 81/192 (42%), Positives = 116/192 (60%), Gaps = 6/192 (3%)
Frame = +3
Query: 271 TSDSASPANMIGIRVEKSGEFSYEELANATNNFNMANKIGQGGFGEVYYAELN-GEKAAI 329
+S++ + ++ I ++ FSYE L AT NF+ NK+G+GGFG V+ +LN G + A+
Sbjct: 147 SSEAQNEDDIQNIAAKEHKTFSYETLVAATKNFHAVNKLGEGGFGPVFKGKLNDGREIAV 326
Query: 330 KKMDMKATK---EFLAELKVLTRVHHVNLVRLIGYCVEGS-LFLVYEYIDNGNLGQHL-R 384
KK+ ++ + +F+ E K+LTRV H N+V L GYC GS LVYEY+ +L + L R
Sbjct: 327 KKLSRRSNQGRTQFINEAKLLTRVQHRNVVSLFGYCAHGSEKLLVYEYVPRESLDKLLFR 506
Query: 385 SSDGEPLSWSIRVKIALDSARGLEYIHEHTVPTYIHRDIKSENILLDKNFCAKVADFGLT 444
S E L W R I ARGL Y+HE + IHRDIK+ NILLD+ + K+ADFGL
Sbjct: 507 SQKKEQLDWKRRFDIISGVARGLLYLHEDSHDCIIHRDIKAANILLDEKWVPKIADFGLA 686
Query: 445 KLIDAGISSVPT 456
++ + V T
Sbjct: 687 RIFPEDQTHVNT 722
>TC12704 UP|CAE45596 (CAE45596) S-receptor kinase-like protein 3, complete
Length = 2481
Score = 136 bits (342), Expect = 1e-32
Identities = 96/311 (30%), Positives = 158/311 (49%), Gaps = 28/311 (9%)
Frame = +1
Query: 291 FSYEELANATNNFNMANKIGQGGFGEVYYAEL-NGEKAAIKKMDM---KATKEFLAELKV 346
F + +++ TN+F+ +NK+G+GGFG VY L NG++ A+K++ + +EF E+K+
Sbjct: 1417 FDFSTISSTTNHFSESNKLGEGGFGPVYKGVLANGQEIAVKRLSNTSGQGMEEFKNEVKL 1596
Query: 347 LTRVHHVNLVRLIGYCV-EGSLFLVYEYIDNGNLGQHLRSSDGEPLSWSIRVKIALDSAR 405
+ R+ H NLV+L+G + + L+YE++ N R
Sbjct: 1597 IARLQHRNLVKLLGCSIHHDEMLLIYEFMHN----------------------------R 1692
Query: 406 GLEYIHEHTVPTYIHRDIKSENILLDKNFCAKVADFGLTKLIDAGISSVPTVNMAGTFGY 465
L+Y + IHRD+K+ NILLD K++DFGL ++ T + GT+GY
Sbjct: 1693 SLDYFIFDSRLRIIHRDLKTSNILLDSEMNPKISDFGLARIFTGDQVEAKTKRVMGTYGY 1872
Query: 466 MPPEYA-YGSVSSKIDVYAFGVVLYELISAKA----------AVIMGEDSGAD---LKGL 511
M PEYA +GS S K DV++FGV++ E+IS K ++ S +K L
Sbjct: 1873 MSPEYAVHGSFSVKSDVFSFGVIVLEIISGKKIGRFCDPHHHRNLLSHSSNFAVFLIKAL 2052
Query: 512 -VVLFEEV--------FDQPHPIEGLKKLVDPRLGDNYPIDHVFKMAQLAKVCTNSDPQQ 562
+ +FE V + + P+E + +L+D G P + + + +A +C P+
Sbjct: 2053 RICMFENVKNRKAWRLWIEERPLELVDELLD---GLAIPTE-ILRYIHIALLCVQQRPEY 2220
Query: 563 RPNMSSVVVAL 573
RP+M SVV+ L
Sbjct: 2221 RPDMLSVVLML 2253
>TC12605 similar to UP|AAR99871 (AAR99871) Strubbelig receptor family 3,
partial (22%)
Length = 680
Score = 132 bits (332), Expect = 2e-31
Identities = 83/190 (43%), Positives = 110/190 (57%), Gaps = 12/190 (6%)
Frame = +3
Query: 291 FSYEELANATNNFNMANKIGQGGFGEVYYAEL-NGEKAAIKKMDMKATK-----EFLAEL 344
F+ L TN+F+ N IG G G V AEL NG+ A+ K+D +A+ EFL +
Sbjct: 117 FTIASLQQYTNSFSQENLIGGGMLGSVXKAELPNGKLLAVXKLDKRASAHQKDDEFLELV 296
Query: 345 KVLTRVHHVNLVRLIGYCVE-GSLFLVYEYIDNGNLGQHLRSSDG--EPLSWSIRVKIAL 401
+ R+ H N+V LIGYC E G L+YEY NG+L L S D LSW+ R+++AL
Sbjct: 297 NSIDRIRHANIVELIGYCSEHGQRLLIYEYCSNGSLHDALHSDDEIKTKLSWNARIRMAL 476
Query: 402 DSARGLEYIHEHTVPTYIHRDIKSENILLDKNFCAKVADFGLTKLI---DAGISSVPTVN 458
+AR LEY+HE P IHRD+KS NILLD++ V+D GLT LI D +SS N
Sbjct: 477 GAARALEYLHELCHPPVIHRDLKSANILLDEDLSVHVSDCGLTPLITFRDL*VSS--RGN 650
Query: 459 MAGTFGYMPP 468
+ +GY P
Sbjct: 651 LLTAYGYGXP 680
>BI418434
Length = 478
Score = 131 bits (330), Expect = 3e-31
Identities = 74/158 (46%), Positives = 99/158 (61%), Gaps = 4/158 (2%)
Frame = +2
Query: 299 ATNNFNMANKIGQGGFGEVYYAELNGEK-AAIKKMDMKA--TKEFLAELKVLTRVHHVNL 355
AT F+ K+G GGFG V+ EL+ A+K+++ KEF AE+ + + HVNL
Sbjct: 11 ATRGFS--EKVGHGGFGTVFQGELSDSSLVAVKRLERPGGGEKEFRAEVSTIGNIQHVNL 184
Query: 356 VRLIGYCVEGS-LFLVYEYIDNGNLGQHLRSSDGEPLSWSIRVKIALDSARGLEYIHEHT 414
VRL G+C E S LVYEY+ NG L +LR DG LSW +R ++A+ +A+G+ Y+HE
Sbjct: 185 VRLRGFCSENSHRLLVYEYMHNGALSAYLRK-DGPCLSWDVRFQVAIGTAKGIAYLHEEL 361
Query: 415 VPTYIHRDIKSENILLDKNFCAKVADFGLTKLIDAGIS 452
+H DIK ENILLD +F AKV+DFGL KLI S
Sbjct: 362 RCCILHCDIKPENILLDSDFTAKVSDFGLAKLIGRDFS 475
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.317 0.135 0.390
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 9,441,117
Number of Sequences: 28460
Number of extensions: 121728
Number of successful extensions: 1167
Number of sequences better than 10.0: 322
Number of HSP's better than 10.0 without gapping: 909
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 920
length of query: 601
length of database: 4,897,600
effective HSP length: 95
effective length of query: 506
effective length of database: 2,193,900
effective search space: 1110113400
effective search space used: 1110113400
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 58 (26.9 bits)
Medicago: description of AC123570.7


