

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC122729.6 + phase: 0 /pseudo
(247 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
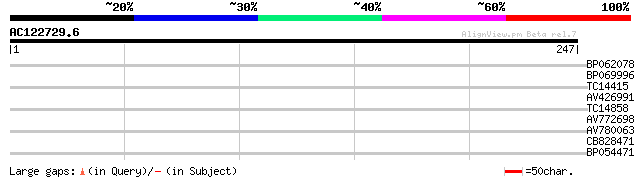
Score E
Sequences producing significant alignments: (bits) Value
BP062078 29 0.66
BP069996 28 1.5
TC14415 similar to UP|O04237 (O04237) Transcription factor, part... 27 4.3
AV426991 26 5.6
TC14858 homologue to UP|NUIM_SOLTU (P80269) NADH-ubiquinone oxid... 26 5.6
AV772698 26 5.6
AV780063 26 7.3
CB828471 25 9.5
BP054471 25 9.5
>BP062078
Length = 419
Score = 29.3 bits (64), Expect = 0.66
Identities = 28/104 (26%), Positives = 45/104 (42%), Gaps = 2/104 (1%)
Frame = -1
Query: 32 EEEREDIYESFEDETLEKPIYDEEYVGADICEVFDEEGNIDPIYDENGLDDIHEAFEKEE 91
++E ++ E FE E ++E V D E NID + G D E +E+
Sbjct: 380 DDENDEFEEDFETELNGGKHEEDEEVEEDA-----ENENIDNSWSIGG-DIRMEKYERSR 219
Query: 92 HN--EPIYDEEYVLAEYGEYLETKKRFQTSTNKDMFLMSSSITK 133
E ++ +L +GE L TS + D+ +S S+TK
Sbjct: 218 GRCQEGDHENXELLVNWGEPLRVALALATS*SSDLDSLSVSVTK 87
>BP069996
Length = 511
Score = 28.1 bits (61), Expect = 1.5
Identities = 12/31 (38%), Positives = 19/31 (60%)
Frame = -2
Query: 86 AFEKEEHNEPIYDEEYVLAEYGEYLETKKRF 116
+F E H PIY E + + +G YL+ KK++
Sbjct: 246 SFWAEVHYNPIYPEGELPSSHGSYLKKKKKW 154
>TC14415 similar to UP|O04237 (O04237) Transcription factor, partial (83%)
Length = 1666
Score = 26.6 bits (57), Expect = 4.3
Identities = 16/50 (32%), Positives = 25/50 (50%)
Frame = +2
Query: 66 DEEGNIDPIYDENGLDDIHEAFEKEEHNEPIYDEEYVLAEYGEYLETKKR 115
DE+ ++ + D N +E EKE D+E L EY + LE K++
Sbjct: 845 DEKPAVEDVADGNKESSANENEEKEPE-----DKEMTLEEYEKVLEEKRK 979
>AV426991
Length = 429
Score = 26.2 bits (56), Expect = 5.6
Identities = 14/52 (26%), Positives = 24/52 (45%)
Frame = -1
Query: 4 CQGYGHIAIDCVNYKAVTIVNREINNIFEEEREDIYESFEDETLEKPIYDEE 55
C + H D V+ K V + + E+NNI + + + TLE+ D +
Sbjct: 378 CSDFWHSGSDTVSQKMVFLEHSELNNIGRDYGVVVPVAVASSTLEREREDND 223
>TC14858 homologue to UP|NUIM_SOLTU (P80269) NADH-ubiquinone oxidoreductase
23 kDa subunit, mitochondrial precursor (Complex
I-23KD) (CI-23KD) (Complex I-28.5KD) (CI-28.5KD) ,
partial (93%)
Length = 1106
Score = 26.2 bits (56), Expect = 5.6
Identities = 21/74 (28%), Positives = 31/74 (41%)
Frame = +2
Query: 40 ESFEDETLEKPIYDEEYVGADICEVFDEEGNIDPIYDENGLDDIHEAFEKEEHNEPIYDE 99
E ED + YD + C E +D I + + F E H E +YD+
Sbjct: 494 EEREDGSRRTTRYDIDMTKCIYCGFCQEACPVDAIVEGPNFE-----FVTETHEELLYDQ 658
Query: 100 EYVLAEYGEYLETK 113
E +L E G+ ET+
Sbjct: 659 EKLL-ENGDRWETE 697
>AV772698
Length = 521
Score = 26.2 bits (56), Expect = 5.6
Identities = 15/35 (42%), Positives = 20/35 (56%), Gaps = 1/35 (2%)
Frame = +3
Query: 74 IYDEN-GLDDIHEAFEKEEHNEPIYDEEYVLAEYG 107
+YD N G+ + ++ K E NEP Y EEY YG
Sbjct: 96 VYDPNYGMSQLLISYSKLEFNEPEY-EEYDPTPYG 197
>AV780063
Length = 506
Score = 25.8 bits (55), Expect = 7.3
Identities = 10/17 (58%), Positives = 14/17 (81%)
Frame = -1
Query: 163 STRITR*KYLNTPLSHF 179
S RI+R +YL+TP+ HF
Sbjct: 428 SCRISRTRYLHTPIPHF 378
>CB828471
Length = 525
Score = 25.4 bits (54), Expect = 9.5
Identities = 14/53 (26%), Positives = 26/53 (48%), Gaps = 3/53 (5%)
Frame = +2
Query: 186 KKMYGVILFQWILATLILGCHGNMIVVHYMMVMPTST---LL*NMVLKSSWHV 235
KKM G L +L ++ CH + IV ++V+ T ++ M++ W +
Sbjct: 308 KKMKGHDLLSNVLLLIVTTCHPSPIVAALLLVLTMMTCP*MMFTMLILIRWRI 466
>BP054471
Length = 493
Score = 25.4 bits (54), Expect = 9.5
Identities = 11/16 (68%), Positives = 13/16 (80%)
Frame = +2
Query: 33 EEREDIYESFEDETLE 48
EEREDI+E+F DE E
Sbjct: 182 EEREDIWETF*DENRE 229
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.331 0.144 0.442
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 4,052,003
Number of Sequences: 28460
Number of extensions: 51135
Number of successful extensions: 279
Number of sequences better than 10.0: 18
Number of HSP's better than 10.0 without gapping: 272
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 277
length of query: 247
length of database: 4,897,600
effective HSP length: 88
effective length of query: 159
effective length of database: 2,393,120
effective search space: 380506080
effective search space used: 380506080
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.9 bits)
S2: 54 (25.4 bits)
Medicago: description of AC122729.6


